Study sites and sampling design
We used a pan-European network of 64 mature, uneven-aged forest plots (30 × 30 m2) consisting of three-species mixture stands (34 plots) and corresponding monospecific stands (30 plots; Extended Data Table 1). These plots are part of the FunDivEUROPE (Functional Significance of Forest Biodiversity in Europe) exploratory platform59, and were established across European forests over 2011–2012 to investigate the role of the diversity and composition of regionally common and economically important tree species on ecosystem functioning. The studied plots were distributed across four locations featuring different European forest types and spanning a large biogeographic gradient: North Karelia (Finland), Białowieża (Poland), Râşca (Romania) and Colline Metallifere (Italy), corresponding to typical boreal, hemi-boreal, mountainous beech and thermophilous deciduous (Mediterranean) forests, respectively (Supplementary Table 4 and Supplementary Fig. 5a). In each location, plots were carefully selected based on tree species richness and composition while minimizing as much as possible covariation with potentially confounding environmental factors such as topography and soil conditions59 (Supplementary Fig. 2a). Plot selection was performed so as to include monospecific stands of all tree species from the local species pool and replicate the three-species mixture treatment with different tree species combinations while maximizing community evenness (Extended Data Table 1). This allowed strict avoidance of a dilution gradient, such as would occur in a design with monospecific stands of only one species combined with mixture stands including this species, along with a clear distinction between the effects of species mixing and composition. Our stratified plot selection procedure enabled us to mimic formal biodiversity experiments, given that such manipulative approaches are virtually impossible to undertake in mature forests owing to the high longevity of tree species. Tree species diversity and composition in the studied plots were predominantly the result of natural community assembly from the regional species pool, combined with local forest management practices. The investigated levels of species richness, that is, one versus three tree species, are typical for European forest ecosystems (https://forest.eea.europa.eu/topics/forest-biodiversity-and-ecosystems/forest-ecosystems), boreal forests in Asia and North-America, as well as managed forests and plantations worldwide, although clearly much less diversified than many (sub)tropical forests and some mid-latitude temperate forests47. Overall, our sampling design encompassed a total pool of 13 tree species, including 12 ectomycorrhizal and one arbuscular mycorrhizal tree species, and the local species pool ranged from three to five tree species per location (Extended Data Table 1 and Supplementary Table 4).
Soil organism sampling and analysis
In each plot, we assessed energy fluxes through the soil food web by measuring the biomass of major groups of organisms within this food web22,23,43,60, including both microbial (bacteria and fungi) and faunal (nematodes, microarthropods and macroinvertebrates) groups. Biomass data for all soil organism groups were expressed per unit surface area (g dry weight m−2) at the plot level. Further details on biomass calculation and the trophic classification of soil organisms are provided in the Supplementary Methods.
Sampling
Soil organisms were sampled in all plots during the phenological spring of 2017 (Supplementary Table 4), a period of high soil biological activity. We sampled both the litter layer (unfragmented aboveground litter, OL horizon) and the soil layer (including both fragmented/humified organic matter and mineral soil, OF/OH/A horizons). In each plot, we selected five 10 × 10-m2 subplots, with samples taken equidistantly from three trees of either the same species in monospecific stands or of different species for three-species mixture stands (Supplementary Fig. 5b). For each subplot, soil samples for microbial analyses and nematode extraction were collected by taking five soil cores (10-cm depth, 5.3-cm diameter), spaced approximately 35 cm apart around the equidistant point between the three trees, weighted by tree individual size, that is, individual diameter at breast height. The five cores were gently sieved through 6-mm mesh (to avoid damaging nematodes), homogenized and pooled at the subplot level for nematode extraction. Pooled soil was then sieved through 2-mm mesh for microbial analyses. All subsamples were stored at 4 °C until further processing. For microarthropod extraction, an intact core (10-cm depth, 10-cm diameter), including both the litter and soil layers, was collected from each of three subplots along a southwest–northeast transect and stored at 4 °C until further processing. For the hand-sorting of soil macroinvertebrates, an intact monolith (25-cm depth, 25 × 25-cm2 surface), including both the litter and soil layers, was collected from each of the same three subplots. To express all data per unit surface area, an extra core was sampled, sieved through 2-mm mesh, dried at 105 °C for 48 h and weighted to measure soil bulk density.
Microorganisms
The biomass of bacteria (gram-positive and gram-negative), arbuscular mycorrhizal fungi and non-arbuscular mycorrhizal fungi was quantified at the plot level using phospholipid fatty acid data61. Fungal community data based on metagenomic amplicon sequencing and bioinformatics62 were used to partition non-arbuscular mycorrhizal fungal biomass into five trophic guilds: ericoid mycorrhizal fungi, ectomycorrhizal fungi, general saprotrophic fungi, wood saprotrophic fungi and plant pathogenic fungi (Supplementary Methods). The biomass of each fungal trophic guild was calculated by multiplying its relative abundance, that is, number of reads divided by the total number of reads for the five trophic guilds, by the total non-arbuscular mycorrhizal fungal biomass.
Nematodes
Nematodes were extracted for each subplot within 72 h after sampling from approximately 100 g of fresh soil using a modified sugar flotation method63, before being heat-killed and fixed in 4% formaldehyde. Nematodes were then pooled and counted at the plot level, and a subsample of approximately 160 randomly selected individuals were identified to family level. The biomass of nematode families was calculated based on body mass data retrieved from the Nemaplex database (http://nemaplex.ucdavis.edu). Nematode families were assigned to five trophic guilds64: herbivores, bacterivores, fungivores, omnivores and carnivores.
Microarthropods
Microarthropods were extracted from the intact core (including both the litter and soil layers) within 72 h after sampling for each subplot using the Berlese–Tullgren funnel method65, and were fixed in 70% ethanol. Microarthropods were then counted and identified to species level for collembola and to order level for mites. Microarthropod biomass was estimated based on an allometric model using body length data retrieved from the BETSI database (https://portail.betsi.cnrs.fr) for collembola, and data from the literature for mites60. Microarthropod taxa were assigned to seven trophic guilds64 belonging to the following broad trophic groups: microbi-detritivores, fungivores, omnivores and carnivores.
Macroinvertebrates
Macroinvertebrates were hand sorted in the field for each subplot and fixed in 70% ethanol. Macroinvertebrates were then counted and identified to species level for Lumbricidae, Isopoda, Diplopoda, Chilopoda and Araneae; and order to family level for other taxa66. All macroinvertebrate individuals were weighed for body mass. Macroinvertebrates were assigned to 23 trophic guilds64 belonging to the following broad trophic groups: herbivores, detritivores, humi-detritivores, omnivores and carnivores.
Metabolic rates
We used soil microbial respiration and biomass data67 to calculate the metabolic rates of microbial groups, whereas we used a model based on individual body mass, environmental temperature (mean soil temperature during the growing season; see ‘Microclimate’ section below) and phylogenetic grouping68 to calculate the metabolic rates of faunal groups (Supplementary Methods).
Assimilation efficiencies
Assimilation efficiencies (that is, the proportion of consumed food assimilated by digestion) specific to each food type (plant-derived resource or prey) for faunal consumers were calculated using a model based on food N content69, and we also applied a temperature correction of assimilation efficiency70 (Supplementary Methods). Assimilation efficiencies of all food resources were set to 1 for microbial consumers, given their external digestion system.
Plant-derived resources
In each plot, we quantified the biomass of the following plant-derived (basal) resources of the soil food web: living fine roots38 (absorptive roots belonging the first three root orders) and associated photosynthates/rhizodeposits34, litter (including dead leaves67, dead roots71 and dead wood) and SOM72 (Supplementary Methods). Photosynthates/rhizodeposits refer to organic compounds provided by roots to mycorrhizal fungi, or released directly by roots into the soil. To estimate the biomass of rhizodeposits, we used a mass ratio of 0.4 between net rhizodeposition (the portion of rhizodeposited C remaining in the soil after microbial utilization and respiration) and root biomass, based on the results of a meta-analysis of fixed C partitioning in plant–soil systems34. The biomass of rhizodeposits was calculated by multiplying the biomass of living fine roots by this factor.
Food web reconstruction
We constructed our soil food web from 47 network nodes, including six plant-derived (basal) resources and 41 trophic guilds of soil organisms (consumers) that were differentiated based on multiple traits64 (Supplementary Table 5). To establish the topology of the soil food web and quantify trophic interaction strengths (Supplementary Fig. 1), a food web interaction matrix was constructed based on basic food web principles, and a priori knowledge of soil organism biology and key traits of consumers following the approach in ref. 7, except that microbes were considered here as consumers rather than basal resources (Supplementary Methods). Briefly, the food web interaction matrix was calculated by multiplying five matrices representing different trait dimensions: phylogenetically defined feeding preferences, density dependence, predator–prey interactions related to body mass ratio and hunting strategy, prey defence mechanisms and spatial niche overlap related to vertical stratification. The five matrices relied on the following assumptions, respectively7: (1) there are phylogenetically conserved differences in feeding preferences of consumers; (2) food consumption is density (biomass) dependent, that is, consumers will preferentially feed on food resources that are locally abundant owing to a higher encounter rate; (3) the strength of predator–prey interactions is primarily defined by the optimum predator–prey mass ratio, that is, a predator is typically larger than its prey, but certain predator hunting traits can modify the optimum predator–prey mass ratio; (4) the strength of predator–prey interactions can be weakened by prey defence traits, that is, prey with efficient physical, chemical or behavioural protection are consumed less; (5) the strength of trophic interactions between a consumer and its food resource is modulated by the overlap in their spatial niches related to vertical stratification, with greater overlap leading to a stronger interaction. Food web reconstruction was carried out separately for each plot to account for plot-specific density dependence.
Food web energy fluxes and functions
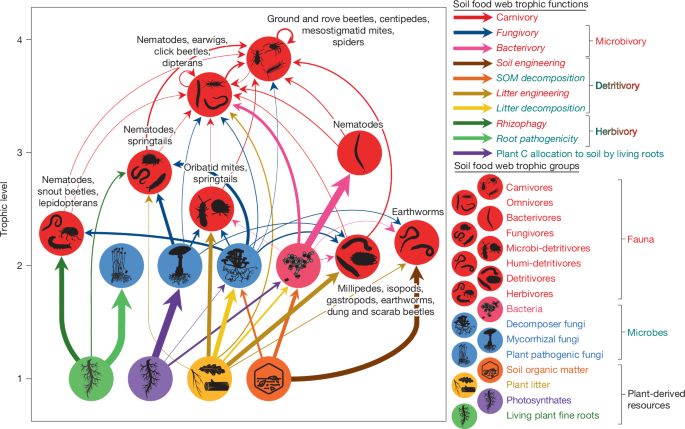
For the calculation of energy fluxes, we assumed a steady state of the soil food web6. This means that the energy flowing into a given feeding guild of the food web through food consumption balances the energy lost by excretion, metabolism and predation of that feeding guild. Energy fluxes to each feeding guild within the food web (kJ m−2 d−1) were then calculated based on the trophic interaction matrix using the following equation6: \(F=\frac1e_\rma\times (X+L)\), where F is the total flux of energy into the feeding guild, ea is the diet-specific assimilation efficiency, X is the community metabolism of the feeding guild and L is the energy loss to predation (see the Supplementary Methods for further details). To simplify the representation of the food web, we aggregated biomass and energy flux matrices at broader trophic group levels by summing the rows and columns of trophic nodes belonging to the same trophic group (Fig. 1 and Supplementary Table 5).
We then calculated five broad trophic functions of the soil food web (Fig. 1): carnivory, the sum of energy fluxes outgoing from fauna to their faunal consumers; microbivory, the sum of energy fluxes outgoing from microbes to their faunal consumers; herbivory, the sum of energy fluxes outgoing from living fine roots to their consumers; plant C allocation to soil by living roots, the sum of energy fluxes outgoing from photosynthates and rhizodeposits (via living fine roots) to their microbial consumers; and detritivory, the sum of energy fluxes outgoing from detritus (dead organic matter, including plant litter and SOM) to their consumers. Additionally, we calculated eight more specific trophic functions (Fig. 1): bacterivory and fungivory, the sum of energy fluxes outgoing from bacteria and fungi to their faunal consumers, respectively; root pathogenicity and rhizophagy, the sum of energy fluxes outgoing from living fine roots to their microbial and faunal consumers, respectively; litter decomposition and litter engineering, the sum of energy fluxes outgoing from plant litter to their microbial and faunal consumers, respectively; and SOM decomposition and soil engineering, the sum of energy fluxes outgoing from SOM to their microbial and faunal consumers, respectively. Decomposition refers to the assimilation and mineralization of dead organic matter through respiration, a process that is mediated mainly by microbes in soil60. Engineering refers to the physical modification, maintenance or creation of habitats1, which is a major way through which soil faunal detritivory affects the decomposition, transformation and translocation of dead organic matter7,73. Such inferences from energy fluxes about effects that are not purely trophic are especially justifiable in soil where habitat and food resources are tightly interlinked7. As such, the faunal consumption of litter results in the conversion of litter into faeces which in turn accelerates its decomposition through chemical and physical changes73, as a form of litter engineering. Similarly, the faunal consumption of SOM is linked to biopedturbation and soil structure maintenance73 in a manner that represents soil engineering.
We quantified soil food web multifunctionality based on ten trophic functions: plant C allocation to soil by living roots, root pathogenicity, rhizophagy, litter decomposition, litter engineering, SOM decomposition, soil engineering, bacterivory, fungivory and carnivory. Defining high multifunctionality by fast rates of multiple functions simultaneously, we calculated soil food web multifunctionality using the ‘averaging’ approach74, that is, the main range-standardized values for the ten trophic functions (Supplementary Methods). Consistent results were found when using the alternative ‘threshold’ approach74 (see ‘Sensitivity analyses’ in the Supplementary Methods). Additionally, we adopted the ‘single functions’ approach to help illuminate which individual functions drive trends in the effects of tree communities on soil food web multifunctionality74.
Tree functional (trait-based) properties
Functional diversity and composition of tree communities were characterized using a set of nine plant functional traits known to be directly involved in resource economics9,32. These traits included three leaf traits: leaf nitrogen content, specific leaf area and leaf dry matter content; and six fine (absorptive) root traits71: root nitrogen content, root tissue density, specific root length, mean root diameter, root length density and ectomycorrhizal colonisation intensity. Trait data of each tree species were mostly derived from plot-specific measurement in the field, but specific leaf area and leaf dry matter content data for Poland and Italy were retrieved from the TRY plant trait database75 (https://www.try-db.org). Using trait values from databases has limitations because of intraspecific trait variability. However, the sensitivity analysis that we performed revealed that the associated uncertainty in trait values had little bearing on our overall inferences (see ‘Sensitivity analyses’ in the Supplementary Methods). See the Supplementary Methods for further details and Supplementary Table 6 for trait values of each tree species and geographic location.
To quantify functional diversity, we calculated the functional dispersion (FDis) index corresponding to the mean distance of each species to the centroid of all species within the multidimensional trait space76. For functional composition, we calculated CWMs of each trait. Both FDis and CWM values were computed based on the relative basal area of tree species. We performed a principal component analysis (PCA) on all CWM traits, which simplified functional composition into two dimensions71 (Extended Data Fig. 4a and Supplementary Table 7): (1) an LES (45.2% of variation), which ranged from slow/conservative leaf attributes (N-poor and dry-matter-rich leaves with low leaf area per unit mass) to fast/acquisitive leaf attributes (N-rich and dry-matter-poor leaves with high leaf area per unit mass), and also aligned here with a fine-root gradient of belowground resource foraging strategies ranging from low foraging efficiency (thick fine roots with low length per unit mass) to high foraging efficiency (thin fine roots with high length per unit mass); (2) an RES (31.5% of variation), which ranged from slow/conservative fine-root attributes (N-poor fine roots with low tissue density) to fast/acquisitive fine-root attributes (N-rich fine roots with high tissue density), and also aligned here with a fine-root gradient of soil exploration strategies ranging from ‘do-it-yourself’ attributes (high root length density and low ectomycorrhizal colonisation intensity) to ‘outsourcing’ attributes (low root length density and high ectomycorrhizal colonisation intensity). The mean values of FDis and the scores of the two first PCA axes of CWM trait ordination for each location are shown in Extended Data Table 1.
Tree and understorey vegetation
We characterized forest vegetation using a set of eight variables measured in each plot: tree aboveground biomass, tree aboveground litterfall, total root biomass, aboveground biomass of both woody and herbaceous understorey plants, and aboveground C:N ratio and species richness of understorey plant communities (Supplementary Methods).
Environmental drivers
Abiotic conditions
We characterized abiotic conditions in each plot using a set of six variables related to soil texture, macroclimate and topography: the sand, silt and clay content of mineral soil (A horizon, 10-cm depth), mean annual temperature and precipitation, and altitude (Supplementary Methods). We performed a PCA to reduce the dimensionality of abiotic properties into two dimensions (Supplementary Fig. 2a): (1) a soil texture and macroclimate gradient ranging from coarse soil texture, and dry and cold macroclimate to fine soil texture, and wet and hot macroclimate (70.8% of variation); (2) a temperature gradient ranging from cold macroclimate with high elevation to warm macroclimate with low elevation (19.1% of variation).
Microclimate
We characterized microclimate using a set of six variables: soil temperature and moisture (measured at 8-cm soil depth) and air temperature (measured 50 cm above the ground) for both annual and growing season (daily mean temperatures > 5 °C) periods (Supplementary Methods). We performed a PCA to simplify microclimatic properties into a single dimension ranging from cold and wet to hot and dry (Extended Data Fig. 4b; 63.1% of variation).
Leaf litter quality
We characterized freshly fallen tree leaf litter quality using a set of 16 chemical and physical variables mainly related to elemental composition and C quality77: the concentrations of N, P, Ca, Mg and K, and the C:N and C:P ratios, the proportion of C fractions (lignin, cellulose, hemicellulose and water-soluble compounds), the concentrations of secondary metabolites (condensed tannins, total phenolics and soluble phenolics), as well as the litter pH and water holding capacity. Leaf litter quality data of each tree species were derived from location-specific measurement (Supplementary Methods).
We then quantified the functional diversity and composition of tree leaf litter by calculating the FDis index and CWMs of each litter property based on the relative leaf litter mass of the component tree species. We performed a PCA to simplify tree leaf litter properties into one dimension ranging from low to high nutritional quality (Extended Data Fig. 4c; 42.5% of variation).
Soil fertility
We characterized soil fertility using a set of six variables measured in each plot72: the organic C content of mineral soil (A horizon, 10-cm depth), the mass of the forest floor (including unfragmented aboveground litter and fragmented/humified organic matter, OL/OF/OH horizons), as well as the pH and C:N ratios of both the forest floor and mineral soil (Supplementary Methods). We performed a PCA to simplify soil properties into a single dimension ranging from low to high fertility (Supplementary Fig. 2b; 40.1% of variation).
Measured ecosystem processes
To characterize in situ patterns of litter decomposition, we used field-based data of naturally occurring tree leaf litter decomposition measured in each plot78 (Supplementary Methods). To characterize SOM decomposition, we further used soil microbial respiration and biomass data67 (Supplementary Methods). To characterize soil and litter engineering, we used humus type and forest floor mass data72 as these two variables reflect the transformation and translocation of dead organic matter by faunal detritivory73.
Statistical analyses
All analyses were performed using R v.4.3.0 (ref. 79). To test how tree community properties affect soil food web functioning, we used Bayesian multi-level models that accounted for the hierarchical design of the study, with plots nested within four geographic locations across Europe. We used random intercept models (with varying intercepts but a common slope across locations) to assess general patterns across European forests. Similar results were found when using random slope models (with varying intercepts but a common slope across locations), indicating that our findings are robust to model specification (see ‘Sensitivity analyses’ in the Supplementary Methods). To ensure that the effects of tree communities were not biased by confounding factors, we also explicitly included abiotic covariates into the models, that is, the first two PCA axes representing abiotic conditions (Supplementary Fig. 2a). This statistical adjustment allowed us to control for variation in abiotic conditions both among and within locations. Following the principle of the ‘structural causal model’ framework, controlling for confounding factors allows to satisfy the ‘backdoor criterion’ required for quantifying unbiased causal relationships from observational data80. We also checked whether omitted variable bias generated by potential unmeasured confounders at the location level could affect our inference, and found that it was not an issue in our study (see ‘Sensitivity analyses’ in the Supplementary Methods). We adopted both taxonomic and functional approaches. For the taxonomic approach, we used linear mixed-effects models (LMMs) including tree species richness (comparing three-species mixture stands to corresponding monospecific stands) and abiotic covariates (see above) as fixed factors, and tree species composition (a factor listing all tree species present; Extended Data Table 1) and location as random factors, as in the following equation:
$$\beginarraycy_ij=\alpha +\beta _1\rmr\rmi\rmc\rmh\rmn\rme\rms\rms_ij+\beta _2\rma\rmb\rmi\rmo\rmt\rmi\rmc\; PC1_ij+\beta _3\rma\rmb\rmi\rmo\rmt\rmi\rmc\; PC2_ij+\delta _ij^\rmc\rmo\rmm\rmp\rmo\rms\rmi\rmt\rmi\rmo\rmn+\delta _i^\rml\rmo\rmc\rma\rmt\rmi\rmo\rmn+\epsilon _ij\\ \rmw\rmi\rmt\rmh\,\delta _ij^\rmc\rmo\rmm\rmp\rmo\rms\rmi\rmt\rmi\rmo\rmn \sim N(0,\sigma _\rmc\rmo\rmm\rmp\rmo\rms\rmi\rmt\rmi\rmo\rmn^2),\delta _i^\rml\rmo\rmc\rma\rmt\rmi\rmo\rmn\, \sim N(0,\sigma _\rml\rmo\rmc\rma\rmt\rmi\rmo\rmn^2)\\ \,\rma\rmn\rmd\,\epsilon _ij \sim N(0,\sigma ^2)\endarray$$
(1)
where i refers to location and j refers to plot, y is the dependent variable, α is the general intercept, βx terms are partial slopes, δ terms are random effects, that is, factor-specific deviation from the common intercept, ϵ is the model error, that is, residuals, and σ2 is the variance. For the functional approach, we used LMMs including tree functional diversity (FDis), tree functional composition (first two PCA axes of tree CWM traits corresponding, respectively, to the LES and RES; Extended Data Fig. 4a) and abiotic covariates (see above) as fixed factors, and location as a random factor, as in the following equation:
$$\beginarraycy_ij=\alpha +\beta _1\mathrmFDis_ij+\beta _2\mathrmCWM\; trait\; PC1_ij+\beta _3\mathrmCWM\; trait\; PC2_ij\\ \,+\beta _4\mathrmabiotic\; PC1_ij+\beta _5\mathrmabiotic\; PC2_ij+\delta _i^\mathrmlocation+\varepsilon _ij\\ \,\mathrmwith\,\delta _i^\mathrmlocation \sim N(0,\sigma _\mathrmlocation^2)\,\mathrmand\,\epsilon _ij \sim N(0,\sigma ^2)\endarray$$
(2)
To further investigate the effects of functional composition, we also used LMMs including individual tree CWM trait and abiotic covariates as fixed factors, and location as a random factor, as in the following equation:
$$\beginarraycy_ij=\beta _+\beta _1\mathrmCWM\; trait_ij+\beta _2\mathrmabiotic\; PC1_ij\\ \,+\beta _3\mathrmabiotic\; PC2_ij+\delta _i^\mathrmlocation+\varepsilon _ij\\ \mathrmwith\,\delta _i^\mathrmlocation \sim N(0,\sigma _\mathrmlocation^2)\,\mathrmand\,\epsilon _ij \sim N(0,\sigma ^2)\,\endarray$$
(3)
Bayesian LMMs were fitted with Markov chain Monte Carlo methods with non-informative priors for all parameters. Each model was fitted based on 10,000 iterations of both warm-up and sampling phases and was checked for their convergence and compliance with statistical assumptions, including normality and independence of residuals, normality of random effects, homogeneity of variance, linearity of relationships and absence of multicollinearity (Supplementary Methods).
For each model, we performed variance partitioning following a model-based approach based on variance components81 to quantify the proportion of variance in the dependent variable explained by three groups of predictors: diversity (tree species richness or FDis), composition (tree species composition or CWM traits PC1 and PC2) and biogeography (abiotic conditions PC1 and PC2, and location). To minimize the influence of multicollinearity within each group (especially between abiotic covariates and location in the biogeography group; Supplementary Fig. 2a), we performed variance partitioning at the level of variable groups. This was done by summing the variance terms of all predictors within each group81. This allowed us to incorporate the covariances between group predictors to the variance component of that group, thereby reducing the covariance between variance components at the variable group level. Variance components of each group were standardized by the sum of all group-level variance components, including the residuals. To assess how covariance between group-level variance components might affect total variance accounting81, we computed further variance partitioning metrics: (1) basic variance partition, which expresses the variance of each group standardized by the total variance of the dependent variable; (2) marginal variance partition, which quantifies the contribution of each group while accounting for covariances with all other groups; and (3) partial variance partition, which quantifies the contribution of each group that cannot be explained by a linear combination of the remaining groups. Further methodological details are provided in the Supplementary Methods.
We calculated the effect size of fixed factors as the slope coefficient standardized by the standard deviation (Supplementary Methods). We quantified the uncertainty of effect size based on credible intervals. Standardized slope (βst) values of |βst| < 0.12, 0.12 ≤ |βst| < 0.24, 0.24 ≤ |βst| < 0.41 and |βst| ≥ 0.41 were interpreted as neutral, weak, moderate and strong effects, respectively. We used P values of slope coefficients and Bayes factors as measures of evidence for fixed and random effects, respectively. The P values were derived from posterior distributions using a two-tailed test82. Bayes factors were used to quantify the support for models including the random factor tested, compared with null models without the random factor. Bayes factors were calculated as the ratio of marginal likelihoods of the two models (Supplementary Methods).
To test a priori multivariate causal hypotheses regarding how tree community effects on soil food web multifunctionality are mediated by changes in tree and understorey plant properties and microenvironment83,84, we performed piecewise multi-level structural equation modelling (SEM)85. We first constructed a causal diagram based on ecological theory and our previous knowledge of the system85 (Extended Data Fig. 1c and Supplementary Fig. 3) and built an initial SEM model (Supplementary Table 8). Following a local estimation approach85, we tested hypothesized causal relationships by fitting component LMMs with random intercepts across locations using Bayesian modelling, along with the use of variance partitioning and effect size computation methods in a manner similar to those described above. For each component LMM, abiotic covariates (see above) were included to ensure that mediation effects were not biased by confounding factors84, thereby satisfying the ‘backdoor criterion’. The initial SEM model was then simplified by removing unsupported linkages85 using information-theoretic methods to select the most parsimonious SEM model. For each component LMM within the initial SEM model, stepwise backward selection was performed by removing the predictor with the highest P value, verifying at each step that model simplification improved goodness-of-fit using Pareto smoothed importance-sampling leave-one-out cross-validation86 (Supplementary Methods). To test the goodness-of-fit of the selected SEM model, we used Shipley’s test of directional separation87, which assesses whether any important causal pathways may be missing (Supplementary Methods). To quantify and compare the magnitude of tree community effects on soil food web multifunctionality, indirect effects were calculated as the product of the standardized coefficients along each path. Total effects were calculated as the sum of indirect effects.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.