Mouse lines
All experiments were conducted in compliance with the Association for Assessment of Laboratory Animal Care policies and approved by the Institutional Animal Care and Use Committees of the University of Virginia, University of Washington and University of California, Irvine. Mice were housed on a 12-h:12-h light–dark (LD) cycle with food (PicoLab Rodent Diet 5053) and water provided ad libitum unless otherwise indicated. For experiments, we used 8-week or older male and female C57BL/6J mice, Glp1r-IRES-Cre mice (Glp1rtm1.1(cre)Lbrl/RcngJ, strain 029283, RRID: IMSR_JAX:029283), Glp1r-IRES-Cre mice crossed to the Ai14 tdTomato reporter line (B6.Cg-Gt(ROSA)26Sortm14(CAG-tdTomato)Hze/J, strain 007914, RRID: IMSR_JAX:007914), Glp1rflox/flox mice (B6(SJL)-Glp1rtm1.1Stof/J, strain 035238, RRID: IMSR_JAX:035238), Dat-Cre mice (B6.SJL-Slc6a3tm1.1(cre)Bkmn/J, strain 006660, RRID: IMSR_JAX:006660) Gcg-Cre mice (C57BL/6J-Tg(Gcg-cre)-1Mmsc/Mmmh, stock 051056-MU, RRID: MMRRC_051056-MU) and Glp1rS33W mice (described below). Gcg-Cre mice were rederived by in vitro fertilization from frozen sperm (MMRRC, 051056-MU). Approximately equal numbers of males and females were used per group unless otherwise specified.
Generation of Glp1r
S33W mice
The Glp1rS33W mouse line was generated by CRISPR–Cas9 homologous repair at the University of Virginia Genetically Engineered Murine Model Core. In brief, Cas9 (Alt-R S.p. Cas9 Nuclease V3, 100 µg, 1081058), Alt-R HDR Donor Oligo repair template (see subsection below), tracrRNA (Alt-R CRISPR–Cas9 tracrRNA, 5 nmol, 1072532) and CRISPR–Cas9 crRNA XT (ATTTCTGCACCGTCTCTGAG) were microinjected into a fertilized B6SJL zygote and implanted into a pseudopregnant female. Founder pups were genotyped as described below and backcrossed to C57BL/6J mice for at least four generations before experimentation. This strain will be available at The Jackson Laboratory Repository with the JAX 040551 Glp1rS33W mouse line.
Repair template
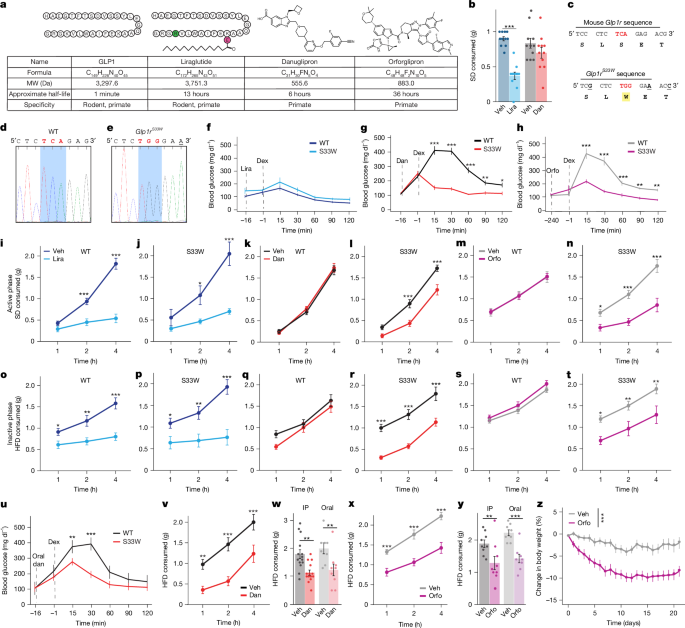
aagagggtgggagtccagtgggaccagaggggctgctggagccacggggcttctgcttttatttctgctttcccttgtagGGTACCACGGTGTCGCTCTGGGAAACCGTCCAAAAGTGGAGAGAATACCGGCGGCAGTGCCAGCGTTTCCTCACGGAAGCGCCACTCCTGGCCACAGgtgcgtccagatgaggcctcacg. Lowercase: non-coding regions; uppercase: coding regions; bold: mutated region; underline: codon coding for S33W mutation.
Validation of Glp1rS33W mice
Tail snips were obtained from pups at 3 weeks of age. DNA was extracted with an extraction buffer (Sigma, E7526) and tissue prep solution (Sigma, T3073), heated for 10 min and 3 min at 55 °C and 100 °C, respectively, then neutralized with a neutralization solution (Sigma, N3910). PrimeSTAR High Fidelity PCR (Takara, R050A) was performed with 1 µl cDNA and 10 µM 5′–3′ F (GATCCCCAAAGTGGCAGTCA) and 5′–3′ R (AGCTATGGACTGGGGATCGT) primers. After amplification, the PCR product was run on a 1.2% agarose gel and bands were cut out at 330 bp. DNA was gel-extracted and purified (QIAGEN, 28704), mixed with 5 µM right primer, H2O, and subsequently sent to be analysed by Sanger sequencing (Azenta). Chromatogram results were analysed to assign WT, heterozygous or homozygous genotypes for each mouse.
Generation of GLP1R viruses
The full-length human GLP1R gene was obtained by PCR, amplifying the human fragment from GLP1R-Tango (plasmid from Addgene, 66295, RRID: Addgene_66295), including the leader sequence present in the GLP1R-Tango. The primers used were: 5′–3′ F (AAAGCTAGCGCCACCATGAAGACGATCATCGCCCTGAGC) and 5′–3′ R (TTTGGCGCGCCCTAA-GAGCAGGACGCCTGACAAGT), ligating the product into pAAV-hSyn-DIO-EGFP (plasmid from Addgene, 50457, RRID: Addgene_50457) in place of the EGFP in NheI and AscI sites to produce the human GLP1R virus construct (AAV-hSyn-DIO-hGLP1R). The full-length mouse Glp1r WT gene was synthesized by Twist Biosciences, generating an NheI and an AscI fragment. This construct included the same leader sequence present in the human construct, as well as an HA tag encoded at the C terminus of the full-length mouse protein-coding region. The fragment was inserted into pAAV-hSyn-DIO-EGFP (plasmid from Addgene, 50457, RRID: Addgene_50457) in place of EGFP to produce the plasmid construct (AAV-hSyn-DIO-mGLP1R-HA). The full-length mouse Glp1r gene bearing a Ser-to-Trp mutation at position 33 (S33W) was made by inserting a synthetic NheI and StuI fragment prepared by Twist Biosciences, containing the single mutation within this fragment. This was cloned into the sites present in the WT construct to produce the S33W mouse mutant, followed by an HA tag encoded at the C terminus (AAV-hSyn-DIO-mGLP1RS33W-HA). Viral plasmid constructs were confirmed by Sanger sequencing. Virus plasmid constructs were prepared and sent to the University of North Carolina Viral Core for preparation of the AAV (serotype 8). In experiments using AAV-DIO-hGLP1R, AAV-DIO-mGlp1r-HA and AAV-DIO-mGLP1RS33W-HA to drive receptor expression, viral titres were carefully calibrated to approximate endogenous receptor levels (Supplementary Fig. 5b); nonetheless, this overexpression approach might alter receptor distribution or signalling, and constitutes a limitation of the model.
Stereotaxic surgery
Mice were anaesthetized with isoflurane (5% induction and 2–2.5% maintenance; Isothesia) and placed in a stereotaxic apparatus (AWD). A heating pad was used throughout the surgery to maintain body temperature, and ocular lubricant was applied to the eyes to prevent desiccation. A total of 200–400 nl virus (rAAV8/AAV2-hSyn-DIO-hGLP1R, plasmid from Addgene, virus packaged at UNC GTC Vector Core, AV9862 (100 µl at titre ≥ 1.5 × 1013 vg ml−1); AAV8-hSyn-DIO-mGLP1RS33W-HA, synthesized by Twist Biosciences, virus packaged at UNC GTC Vector Core, AV10104 (100 µl at titre ≥ 8.2 × 1012 vg ml−1); AAV8-hSyn-DIO-mGLP1R-HA, synthesized by Twist Biosciences, virus packaged at UNC GTC Vector Core, AV10103 (100 µl at titre ≥ 4.5 × 1012 vg ml−1); pAAV9-syn-dLight1.3b, plasmid from Addgene, 135762, RRID: Addgene_135762, virus packaged at UNC GTC Vector Core (100 µl at titre ≥ 1.5 × 1013 vg ml−1); pAAV1-EF1a-DIO- hChR2(H134R)-EYFP-WPRE-HGHpA, plasmid from Addgene, 20298, RRID: Addgene_20298, virus packaged at UNC GTC Vector Core (100 µl at titre ≥ 7 × 1012 vg ml−1); pGP-AAV1-syn-DIO-jGCaMP7s-WPRE, plasmid from Addgene, 104491, RRID: Addgene_104491, virus packaged at UNC GTC Vector Core (100 µl at titre ≥ 1 × 1013 vg ml−1); pAAV1-Ef1a-DIO-EYFP, plasmid from Addgene, 27056, RRID: Addgene_27056, virus packaged at UNC GTC Vector Core (100 µl at titre ≥ 1 × 1013 vg ml−1); AAV8-hSyn-DIO-mCherry, plasmid from Addgene 50459, RRID: Addgene_50459, virus packaged at UNC Vector Core (100 µl at titre ≥ 7 × 1012 vg ml−1); pAAV-hSyn-FLEx-mGFP-2A-Synaptophysin-mRuby, plasmid from Addgene, 71760, RRID: Addgene_71760, virus packaged at UNC Vector Core; pENN.AAV.hSyn.HI.eGFP-Cre.WPRE.SV40, plasmid from Addgene, 105540-AAV8, RRID: Addgene_105540-AAV8, virus packaged at UNC Vector Core (100 µl at titre ≥ 1 × 1013 vg ml−1); pAAV-hSyn-DIO-ChrimsonR-mRuby2-ST, plasmid from Addgene, 105448, RRID: Addgene_105448-AAV9, virus packaged at UNC Vector Core (100 µl at titre ≥ 1 × 1013 vg ml−1); pAAV-hSyn-EGFP, plasmid from Addgene, 50465-AAV8, RRID: Addgene_50465-AAV8, virus packaged at UNC Vector Core (100 µl at titre ≥ 7 × 1012 vg ml−1), retroAAV2-EF1a-oNLS-oScarlet52 (100 µl at titre ≥ 3 × 1013 vg ml−1)), was delivered using a glass pipette at a flow rate of 50 nl per min driven by a microsyringe pump controller (World Precision Instruments, model Micro 4) per targeted site. The syringe needle was left in place for 10 min and was completely withdrawn 17 min after viral delivery. For in vivo calcium and dopamine imaging and optogenetics, a unilateral fibre-optic cannula (RWD, Ceramic Ferrule, Ø 400 μm, 0.5 numerical aperture) was implanted 0.2 mm dorsal to the viral injection coordinates after viral delivery and stabilized on the skull with dental cement (C&B Metabond, Parkell). A minimum of 3 weeks was allowed for recovery and transgene expression after surgery. Stereotaxic coordinates relative to Bregma (G. Paxinos and K. B. J. Franklin): BMH (encompassing the DMH, arcuate nucleus, median eminence and ventromedial hypothalamus), mediolateral (ML): ±0.3 mm, anterior–posterior (AP): −1.4 mm, dorsoventral (DV): −5.9 mm; DMH, ML: ±0.3 mm, AP: −1.8 mm, DV: −5.4 mm; CeA, ML: ±2.7 mm, AP: −1.3 mm, DV: −4.6 mm; VTA, ML: ±0.5 mm, AP: −3.6 mm, DV: −4.5 mm; NAc, ML: ±1.25 mm, AP: +1.0 mm, DV: −4.7 mm from Bregma; and NTS–AP, ML: ±0.15 mm, AP: −0.3 mm, DV: −0.1, −0.4 mm from the zero point of the calamus scriptorius.
For rabies tracing, a total of 500 nl, containing a 1:1 mixture of AAV5-FLExloxP-TC (UNC Vector Core, titre ≥ 2.4×1012 gc ml−1) and AAV8-FLExloxP-RABV-G (UNC Vector Core, titre ≥ 1.0 × 1012 gc ml−1), was injected into the VTA of DAT-Cre mice. Fourteen days later, 500 nl of G-deleted, GFP-expressing, EnvA-pseudotyped rabies virus (RABVΔG-H2B-GFP-EnvA (generated at UC Irvine, K.T.B. laboratory, titre ≥ 5 × 108 colony forming units per ml)) was injected into the same site. Five days after the rabies virus injection, brains were collected for further processing. All surgical procedures were performed under sterile conditions and in accordance with the guidelines of the University of Virginia Institutional Animal Care and Use Committee. Histological analysis was performed to validate the success of intracranial surgeries. Mice with unsuccessful viral or implant targeting were excluded from the analysis.
GLP1R agonists
Liraglutide powder (Selleck, S8256) was dissolved in 0.9% NaCl sterile saline, lightly sonicated and further diluted in 0.9% NaCl sterile saline to 0.03 mg ml−1. Danuglipron powder (Selleck, S9851) was dissolved to 30 mg ml−1 in 100% ethanol with gentle sonication, then diluted to 3 mg ml−1 (food intake) or 0.3 mg ml−1 (GTT) in vehicle (1 M NaOH, 2% Tween 80, 5% polyethylene glycol (PEG) 400, 5% dextrose)7. Orforglipron powder (MedChemExpress, HY-112185) was dissolved to 10 mg ml−1 in dimethyl sulfoxide and further diluted in 0.9% NaCl sterile saline to 0.1 mg ml−1. The dosage for danuglipron and orforglipron was decided on the basis of previous literature.
Histological analysis and imaging
For fixed tissue collection, mice were deeply anaesthetized (ketamine:xylazine, 280:80 mg kg−1, intraperitoneally) and perfused intracardially with ice-cold 0.01 M phosphate buffer solution (PBS), followed by fixative solution (4% paraformaldehyde in PBS at a pH of 7.4). For testing brain region FOS activation (Fig. 3), vehicle (control), danuglipron (30 mg kg−1), orforglipron (1 mg kg−1) or liraglutide (0.3 mg kg−1) was delivered by intraperitoneal injection or oral gavage 2 hours (or 6 hours for orforglipron) before perfusion and brain collection. After perfusion, brains were collected and post-fixed overnight at 4 °C in paraformaldehyde. Fixed brains were then transferred into 30% sucrose in PBS for 24 h and then frozen on dry ice. Frozen brains were sectioned immediately or stored at −80 °C for future processing. Coronal sections (30 μm) were collected with a cryostat (Microm HM 505 E). Sections were permeabilized with 0.3% Triton X-100 in PBS (PBS-T) and blocked with 3% normal donkey serum (Jackson ImmunoResearch, RRID: AB_2337258) in PBS-T (PBS-T DS) for 30 min at room temperature. Sections were then incubated overnight at room temperature in primary antibodies diluted in PBS-T DS. For visualization, sections were washed with PBS-T and incubated with appropriate secondary antibodies diluted in PBS-T DS for 2 hours at room temperature. Sections were washed three times with PBS and mounted using DAPI Fluoromount-G (Southern Biotech, 0100-20). Images were captured on a Zeiss Axioplan 2 Imaging microscope equipped with an AxioCam MRm camera using AxioVision 4.6 software (Zeiss), or confocal microscope imaging was performed on a Zeiss LSM 800 microscope. The following primary antibodies were used for fluorescent labelling: anti-c-FOS (rabbit, 1:1,000; Synaptic Systems, 226003, RRID: AB_2231974), anti-DsRed (rabbit, 1:1,000; Takara Bio, 632496, RRID: AB_10013483), anti-tdTomato (goat, 1:1,000; Arigobio, ARG55724), anti-human GLP1R (rabbit, 1:200; Invitrogen, PA5-97789, RRID: AB_2812404), anti-HA (rabbit, 1:1,000, Cell Signaling, 3724), anti-Th (rabbit, 1:500; Chemicon, AB152) and anti-GFP (goat, 1:500; Rockland, 600-101-215). The secondary antibodies (Jackson ImmunoResearch) used were Cy2-conjugated donkey anti-rabbit (1:250; 711-225-152, RRID: AB_2340612), Cy3-conjugated donkey anti-rabbit (1:250; 711-165-152, RRID: AB_2307443), Cy5-conjugated donkey anti-rabbit (1:250; 711-175-152, RRID: AB_2340607), Cy3-conjugated donkey anti-goat (1:250; 705-165-147, RRID: AB_2307351) and Alexa-Fluor 488 donkey anti-goat (1:250; 705-545-003, RRID: AB_2340428). RNA fluorescent in situ hybridization was performed on fixed brain slices with a probe to detect Glp1r RNA (RNAscope Multiplex Fluorescent Reagent Kit v2 Assay, Advanced Cell Diagnostics, 1297311, 24065D). All procedures were performed according to the manufacturer’s instructions.
Antigen retrieval for human GLP1R staining
Antigen retrieval was performed before immunohistochemistry staining of human GLP1R, by incubating the sections in the following solutions sequentially in room temperature: 1% NaOH + 0.3% H2O2 in PBS for 20 min, 0.3% glycine in PBS for 10 min and 0.03% sodium dodecyl sulfate in PBS for 10 min. Then, antigen-retrieval-treated sections were stained following the immunohistochemistry staining procedures described.
FOS analysis pipeline
FOS images were uploaded to ImageJ (Fiji) and cropped on the basis of brain regions outlined in the Allen Brain Atlas. The areas of the cropped regions were measured and recorded. Image thresholds were set per image and particles were analysed within the size restriction of 50–500 pixels. FOS particles were analysed per image, and total particles of each image were divided by total area of the image. At least three FOS images per region for each mouse were quantified and averaged per mouse and per genotype (WT or Glp1rS33W). Ratios of NTS-to-AP FOS activation in Glp1rS33W mice were calculated by dividing FOS per unit area of NTS by FOS per unit area of AP for each mouse, and averaged per injection. The heat map was generated by first normalizing each condition to WT control by region and clustered by column.
Single-nucleus RNA sequencing analysis
Using a previously published single-nucleus RNA sequencing atlas39, we calculated the number of Glp1r+ cells and other markers of interest and quantified their overlap, defining positive cells as those with mRNA counts greater than 0.
Metabolic analysis in the comprehensive laboratory animal monitoring system
Indirect calorimetry in the comprehensive laboratory animal monitoring system (CLAMS, Columbus Instruments) was used to evaluate metabolic parameters of WT and Glp1rS33W mice. All WT and Glp1rS33W mice were single-housed and maintained on a 12:12-h LD cycle with ad libitum access to food (PicoLab Rodent Diet 5053) and water. Metabolic measures of respiratory exchange ratio and energy expenditure were averaged over 3 days per mouse and per genotype (n = 10 or 11 mice per genotype). Averaged LD cycle and total 24-h respiratory exchange ratio and energy expenditure were analysed per genotype and per sex.
GTTs
WT or Glp1rS33W mice were fasted for 16 h overnight before the start of the experiment (ZT10–ZT2). Mice received a tail snip and blood glucose measure using a glucometer (OneTouch Ultra Test Strips for Diabetes), along with injection or oral gavage of danuglipron (3 mg kg−1) or liraglutide (0.3 mg kg−1) 15 min before dextrose (d-glucose) injection, or orforglipron (1 mg kg−1) 240 min before dextrose. Orforglipron was intentionally administered 4 hours earlier because of its partial agonist properties, which mean that it takes longer for it to act. At time point 0, mice received a blood glucose measure and injection of dextrose (1 g kg−1). At 15, 30, 60, 90 and 120 min after injection, blood glucose levels were measured.
Homeostatic (SD) food intake
Home cages were changed and food was removed from the home cage 1 hour before the start of the experiment. Mice were injected with vehicle or drug (danuglipron 30 mg kg−1, liraglutide 0.3 mg kg−1 or orforglipron 1 mg kg−1) at ZT11.5 (ZT8 for orforglipron), and two pellets of SD (PicoLab Rodent Diet 5053) were placed on the home-cage floor at ZT12. All experiments were done in light-tight boxes equipped with programmable lighting, allowing dark-cycle conditions to be maintained during the daytime without exposing the mice to ambient light. Food intake measurements were taken at 1, 2 and 4 hours after ZT12 using infrared night vision goggles (Nightfox Swift Night Vision Goggles). For the danuglipron dose–response experiment, 3 mg kg−1, 10 mg kg−1 or 30 mg kg−1 danuglipron was injected at ZT12 and SD intake was measured two hours later. For post-fast refeeding experiments (Extended Data Fig. 1l–q), mice got new bedding and 16 h of food deprivation (ZT10–ZT2) followed by an injection of drug or vehicle injection at ZT2 and refeeding with SD (30 min after injection for danuglipron and liraglutide; 4 hours after injection for orforglipron). For all food intake measurements, except Fig. 4v,w and the panels in Fig. 5, values were rounded to the nearest 0.1 g; otherwise, data were not rounded.
Hedonic (HFD) food intake
Mice were habituated to HFD (Open Source, D12451; 4.73 kcal g−1; 45% fat, 20% protein, 35% carbohydrate; 17% sucrose) for 1 hour over 2 days before testing days. SD was removed from the home cage 1 hour before the start of the experiment. Mice were injected with vehicle or drug (danuglipron 30 mg kg−1, liraglutide 0.3 mg kg−1 or orforglipron 1 mg kg−1) at ZT1.5 and one pellet of HFD was placed on the home-cage floor at ZT2 (ZT5.5 for orforglipron). Food intake measurements were taken at 1, 2 and 4 hours after HFD delivery. The same parameters were used in oral gavage experiments with danuglipron and orforglipron. All feeding experiments in Fig. 4a–k used a within-individual paired design. Within each viral cohort, the same mice received vehicle and danuglipron on separate, counterbalanced test days. The reported n values therefore correspond to paired vehicle and drug datasets for each mouse, and the plotted intake values reflect one test session per condition per mouse, averaged across mice.
Optogenetic food intake
For CeAGlp1r soma stimulation, mice were single-housed for at least 5 days and habituated daily for 1 hour to a patch-cord and for 1 hour to HFD pellets over 2 consecutive days before testing for the hedonic feeding paradigm. On subsequent testing days, mice underwent optogenetic stimulation at frequencies of 0 Hz (control), 5 Hz, 10 Hz and 20 Hz for 1 hour each, which took place on separate days. For homeostatic feeding assessment using SD, mice were fasted overnight, whereas for hedonic feeding assessment (HFD), mice had ad libitum access to standard chow before testing. Food intake was quantified manually by weighing pre-measured SD or HFD pellets at the beginning and at the end of each 1-hour stimulation session. The laser stimulation protocol consisted of 473-nm blue light delivered at 20 Hz in a 1 s on, 3 s off pulse pattern. Light power exiting the fibre-optic cable was measured using an optical power meter (Thorlabs) and maintained at 8–10 mW across all experiments. All experiments were performed during the light phase, between ZT3 and ZT4. Mice with missed virus injections or off-target fibre placements were excluded from the analysis.
For axon-stimulation experiments, mice expressing ChrimsonR or mCherry in either NTSGcg terminals in the CeA or CeAGlp1r terminals in the VTA were single-housed for at least 5 days and habituated daily for 1 hour to a patch-cord and for 1 hour to HFD pellets over 2 consecutive days before testing for the hedonic feeding paradigm. On different days, mice received 1-hour light stimulation at 0 Hz (control), 20 Hz (CeAGlp1r axons) or 40 Hz (NTSGcg axons) with a 1 s on, 3 s off pulse pattern. Simulations were bilateral for NTSGcg axons in the CeA and unilateral for CeAGlp1r axons in the VTA. Light power exiting the fibre-optic cable was measured using an optical power meter (Thorlabs) and maintained at 8–10 mW across all experiments. Homeostatic and hedonic feeding paradigms were assessed as explained above. All sessions were done during the light phase (ZT3–ZT4), and any mouse with mistargeted viral expression or fibre placement was excluded from analysis.
Weight-loss experiment
Male Glp1rS33W mice (at least 8 weeks old) were placed on a HFD (Open Source, D12451; 4.73 kcal g−1; 45% fat, 20% protein, 35% carbohydrates; 17% sucrose) for at least 8 weeks before the start of the experiment. Mice that did not gain at least 20% of their baseline body weight were excluded from testing, disqualifying female mice from this study. Male mice were randomly assigned to 21 days of vehicle or orforglipron (1 mg kg−1) injection and were retested with the opposite treatment after 10 days of rest. Glp1rS33W mice were injected daily at ZT6 and food and body weight were measured.
Conditioned taste avoidance
Mice were water deprived from ZT9 to ZT2 the next day and habituated to two water bottles for 2 days. Measurements of water bottles were taken from ZT2 to ZT3 to ensure mice were drinking. On the third day, mice got two 0.15% saccharin (98+%, Thermo Fisher Scientific Chemicals, 149001000) bottles for 1 hour and were immediately injected or orally gavaged with vehicle or drug (LiCl (125 mg kg−1), danuglipron (30 mg kg−1), liraglutide (0.3 mg kg−1) or orforglipron (1 mg kg−1)). Normal water bottles were restored for the next 2 days. Water was again deprived at ZT9, and at ZT2 the next day, one water bottle and one 0.15% saccharin bottle was counterbalanced and placed in each cage. Measurements were taken after 24 h. Preference ratios were calculated as (0.15% saccharin consumed)/(water consumed + 0.15% saccharin consumed).
Anxiety testing
Single-housed mice were put into the behaviour room to habituate 1 hour before the experiment and were injected with vehicle or danuglipron (30 mg kg−1) at ZT11.5 or orforglipron (1 mg kg−1) at ZT8.5. Experiments were performed from ZT12.5 to ZT13.5. Mice were placed individually in the centre of the arena (50.8 cm diameter with 24 cm walls) with dim light (around 25 lux) and allowed to explore freely for 5 min. In the open field test, the arena is a circular box with a diameter of 50 cm. In the elevated plus maze test, the arena is an elevated cross (height of 40 cm), with two closed arms (5 × 30 cm), and two open arms of the same dimension. Mouse movement was captured by a camera, and nose points and centre points were tracked by Ethovision software. The arena was cleaned with 70% ethanol between trials. In the open field test, the total centre area (33.9 cm diameter) was defined as the centre zone. Total distance travelled and percentage of time in the centre zone were measured. In the elevated plus maze, head dip zone was defined as within 5 cm outside of the open arm. Total distance travelled, time spent in each arm and head dip activity were measured in Ethovision.
Home-cage monitoring of Glp1r
S33W mice
Mice were single-housed and acclimatized to PhenoTyper home cages (Noldus) for 5 days before testing. Each cage was placed inside a light-tight behavioural box equipped with programmable lighting, allowing precise control over light exposure. Cages were maintained on a 12:12-h LD cycle. During acclimatization, mice had ad libitum access to standard chow (SD) in a food hopper, and to water bottles, running wheels, shelters and bedding. Mice that failed to meet a baseline criterion of at least 50 food-hopper head entries between ZT12 and ZT14 were excluded to ensure sufficient engagement with the feeding set-up. Of 59 mice tested, 12 did not meet this threshold and were excluded.
To capture the peak behavioural effects of each drug, injections were timed on the basis of their pharmacokinetics and previous results (Fig. 1), and all behavioural assays were done at the same ZT, with paired controls accounting for differences in injection timing. For testing, lithium chloride (125 mg kg−1) or vehicle was administered 5 min before recording. Danuglipron (30 mg kg−1) or vehicle was administered at ZT11.5, liraglutide (0.3 mg kg−1) or vehicle at ZT10 and orforglipron (1 mg kg−1) or vehicle at ZT8. Injections were counterbalanced across mice. Behaviour was recorded for 2 hours starting at ZT12. In the fed condition, mice had access to a HFD placed on the cage floor from ZT11 to ZT12. At ZT12, the HFD was removed and only SD remained in the food hopper. Fed mice were habituated to HFD exposure (1 hour per day) for 3 days before cage habituation. All mice were also habituated to handling and saline injections for 3 days before recording.
PhenoTyper sensors continuously recorded food-hopper entries, water licks and wheel rotations. Behavioural sessions were video-recorded from above using infrared cameras (960 × 540 pixels, 25 fps, greyscale). Videos were cropped to 2-hour segments using Adobe Premiere Pro and re-encoded using H.264 compression with FFmpeg for consistent playback and frame indexing.
Machine-learning-assisted behaviour classification
Mouse pose tracking was performed using SLEAP (v.1.3.3)24,25. Nine keypoints were tracked: nose, ears (left and right), tail base and five body centre points (Supplementary Fig. 1b). A total of 11,847 frames were manually labelled across 19 representative videos. These were split into training (9,693 frames; 80%), validation (1,077 frames; 10%) and test (1,077 frames; 10%). The test set was drawn from a separate video to ensure independence and to capture a variety of poses and behaviours. The model was trained using a U-Net architecture with max stride 32, 16 filters and ±180° rotation, using default parameters for the single-mouse pipeline. On the held-out test set, the model achieved a mean Object-Keypoint Similarity (OKS) of 0.92, an OKS-based Visual Object Classes (VOC) mean average precision of 0.84, and an OKS-based VOC mean average recall of 0.89.
Representative heat maps of the nose keypoint were generated to visualize the spatial occupancy patterns across the home-cage PhenoTyper. These heat maps were created by cropping to exclude cage walls and binning into 62 × 27 spatial grids.
Although only the nose keypoint was used for heat-map visualization, Keypoint-MoSeq24,25 was applied to the full set of nine tracked keypoints to infer behavioural syllables from 80 h of keypoint data. After alignment and centring, four latent dimensions explained 90% of variance. To determine the appropriate timescale for behavioural segmentation, we ran an automatic κ scan in Keypoint-MoSeq and identified values that yielded syllable durations that were consistent with previous mouse studies25. We then fitted models using the three closest κ values and selected the one that produced syllables most consistent with recognizable behavioural motifs. This final model used a κ value of 105 and identified 91 syllables. Syllables that comprised at least 0.5% of frames were included. Those between 0.01% and 0.5% were retained; syllables less than 0.01% were excluded. Syllables were grouped into behavioural categories (Supplementary Fig. 3) by two trained raters. Some syllables captured blended actions (for example, groom/sniff), probably owing to overhead view limitations. Low-quality and background syllables (around 1%) were excluded.
Syllable analysis
Before manual curation, syllable data generated by MoSeq were analysed to assess behavioural differences across treatment groups. For each drug condition (LiCl, danuglipron, liraglutide and orforglipron), syllable use was compared with those of their respective paired control groups. In addition, syllable transition patterns were examined by comparing bigram-normalized transition probabilities between drug-treated and control mice.
All analyses were performed using code based on previous MoSeq analyses25,53. Owing to the large number of syllables detected, only statistically significant (P < 0.05) or frequently occurring syllables were visualized in most figures. However, all syllables were included in the full analysis.
Behaviour localization and categorization
The locations of the food hopper, water spout and shelter were identified using OpenCV (2024), and corresponding regions of interest were defined. The nose keypoint was used to detect entries into the food and water regions of interest; centre point 3 was used for shelter entry. Behaviours were labelled contextually (for example, ‘sniff by food’ versus ‘sniff’ elsewhere). This produced 22 distinct behaviours, grouped into 5 broader categories on the basis of behavioural similarity and transition frequency: rest or groom in shelter; groom; move or explore; food-motivated; and drink. In some analyses (Extended Data Figs. 5c and 6k,q,w), movement within the shelter was analysed separately. ‘Pause’ outside the shelter was excluded owing to lack of behavioural relevance. These behaviours captured 95.7% of all behavioural time across videos. For each 2-hour recording, time spent per behavioural category was normalized to total behavioural time.
Grouped behaviour analysis
Behavioural proportions were analysed using beta-distributed generalized linear mixed-effects models, with mouse as a random effect. Different link functions (for example, logit or cloglog) were used depending on the distribution of each behaviour. Proportions in the control condition were normalized to mean = 1; treatment data for each mouse were scaled using this normalization to highlight the magnitude of change. For network analysis, transition probabilities between behaviours were computed for each mouse and normalized by the total outgoing transitions from the starting behaviour. Probabilities were then combined across both directions (A↔B) to generate undirected transition measures and averaged per group. Behavioural transition networks were generated using NetworkX54, with line colour scaled by the global maximum-normalized transition probability across all plotted groups. Node size reflects average bout length and was normalized to the global maximum across all groups, then manually scaled in Illustrator (for example, a value of 0.86 corresponds to 86% of the maximum node size). PCA was conducted on selected standardized behavioural feature data to reduce dimensionality. The main analysis code is available on GitHub (see ‘Code availability’).
Electrophysiology recordings
Brain slice preparation
At least a month before recordings, AAV-DIO-mGlp1r or AAV-DIO-hGLP1R was co-injected 1:1 with AAV-DIO-eYFP into Glp1r-Cre mice so that YFP marked Cre-positive neurons competent to express the receptor. Preparation of acute brain slices for patch-clamp electrophysiology experiments was modified from standard protocols previously described55,56,57. Mice were anaesthetized with isoflurane and decapitated. The brains were rapidly removed and kept in modified chilled artificial cerebrospinal fluid (ACSF) (1 °C) containing 93 mM N-methyl-d-glucamine, 2.5 mM KCl, 1.25 mM NaH2PO4, 20 mM HEPES, 5 mM l-ascorbic acid (sodium salt), 2 mM thiourea, 3 mM sodium pyruvate, 0.5 mM CaCl2, 10 mM MgSO4, 25 mM d-glucose, 12 mM N-acetyl-l-cysteine and 30 mM NaHCO3, with pH adjusted to 7.2–7.4 using HCl (osmolarity 310 mOsm). Slices were continuously oxygenated with 95% O2 and 5% CO2 throughout the preparation. Coronal brain sections (300 μm) were prepared using a Leica Microsystems VT1200 vibratome. Slices were collected and placed in ACSF containing 125 mM NaCl, 2.5 mM KCl, 1.25 mM NaH2PO4, 2 mM CaCl2, 1 mM MgCl2, 0.5 mM l-ascorbic acid, 10 mM glucose, 25 mM NaHCO3 and 2 mM Na-pyruvate (osmolarity 310 mOsm), warmed to 37 °C for 30 min and then kept at room temperature for up to 5 hours.
Recordings
Brain slices were placed in a chamber superfused (around 3 ml per min) with continuously oxygenated ACSF solution warmed to 32 ± 1 °C. Human-GLP1R- or mouse-GLP1R-expressing amygdala neurons were identified by video microscopy on the basis of the expression of eYFP and regional markers. Whole-cell electrophysiology recordings were performed using a Multiclamp 700B amplifier with signals digitized by a Digidata 1550B digitizer. Currents were amplified, low-pass-filtered at 2 kHz and sampled at 35 kHz. Borosilicate electrodes were fabricated using a Brown–Flaming puller (model P1000, Sutter Instruments) to have pipette resistances between 2.5 mΩ and 4.5 mΩ. Current-clamp recordings of membrane potentials were collected in an ACSF solution identical to that used for preparation of brain slices. The internal solution contained the following: 120 mM K-gluconate, 10 mM NaCl, 2 mM MgCl2, 0.5 mM K2EGTA, 10 mM HEPES, 4 mM Na2ATP and mM 0.3 NaGTP, pH 7.2 (osmolarity 290 mOsm). Resting membrane potential was recorded as previously described55,56. After 5 min of baseline membrane potential recordings, 30 μM of danuglipron was perfused for 5 min. Action potentials were generated using a current injection ramp (100 pA s−1) before and after 5 min of danuglipron perfusion to determine effects on time to action potential threshold. Danuglipron powder (Selleck, S9851) was dissolved to 30 mg ml−1 in 100% ethanol with gentle sonication, then diluted to 3 mg ml−1 (5.4 mM) in vehicle (1 M NaOH, 2% Tween 80, 5% PEG 400, 5% dextrose). For electrophysiology recordings, 280 µl of this stock was added to 50 ml ACSF (1:180 dilution) to yield a final danuglipron concentration of 30 µM, corresponding to final concentrations of approximately 0.056% ethanol, 0.011% Tween 80 and 0.028% PEG 400.
Statistics
Electrophysiology recordings were analysed using ClampFit (v.11.2). All statistical comparisons were made using the appropriate test in GraphPad Prism (v.10.4.0). Membrane potential and action potential properties underwent descriptive statistics followed by normality and lognormality tests using a Gaussian distribution. Data were assessed for normality using the D’Agostino–Pearson omnibus normality test, Anderson–Darling test, Shapiro–Wilk test and Kolmogorov–Smirnov test with Dallal–Wilkinson–Lilliefors for P values. Cells with unstable resting membrane potentials were excluded from the analysis and no cells were considered statistical outliers. Initial testing was followed by Tukey’s test for drug effect comparison. Data are presented as individual data points and/or mean ± s.e.m.
Fibre photometry recordings
Calcium recordings (GCaMP)
Mice underwent 20-min daily habituation sessions over 2 consecutive days to acclimatize to the fibre-optic cable (Doric Lenses, Ø 400-μm core, 0.57 numerical aperture). On the test day, mice were injected with vehicle, danuglipron (30 mg kg−1) or liraglutide (0.3 mg kg−1) 2 hours before recording. The order of injections was randomized to avoid order effects. After the 2-hour post-injection period, mice were connected to patch cables that were interfaced with rotary joints to enable free movement. Recordings were done for 1 hour in the mouse home cage without food and water. Fibre photometry data were recorded using fluorescent signals from both calcium-dependent (465 nm) and calcium-independent isosbestic (405 nm) excitation wavelengths (Doric). The isosbestic (405 nm) signal served to control for artefacts. The light power of the fibre-optic cable was measured before each experiment and maintained at approximately 20–30 μW for both the calcium-independent isosbestic (405 nm) and the calcium-dependent (465 nm) signal.
Calcium analysis (GCaMP)
The isosbestic signal (405 nm) was fitted to the calcium-dependent (465 nm) signal using a linear least squares method implemented in a custom MATLAB script, and ΔF/F was calculated as (465 nm – fitted 405 nm)/fitted 405 nm. For z-score calculation, we then implemented a paired z-score normalization: for each mouse, we used the full ΔF/F time series from its vehicle session to calculate μ(vehicle) (mean) and σ(vehicle) (s.d.), and all ΔF/F values—both vehicle and drug—were converted to z-scores via [ΔF/F(t) – μ(vehicle)]/σ(vehicle), anchoring comparisons to a common baseline distribution. Significant calcium transients were detected on these z-scored traces using a threshold of median + 2 s.d. (of the entire recording) with a minimum duration of 1.5 s; events were counted per trial and displayed as heat maps of z-score to ensure full transparency of raw recording structure. Finally, the AUC of each mouse’s complete z-scored trace was calculated using a custom MATLAB script to validate overall activity differences between conditions. Heat maps were generated in MATLAB using min–max normalization, scaled to a range of 0–1. For each mouse, the normalization range was determined on the basis of the vehicle condition: the average of the lowest 360 data points was set as the minimum, and the average of the highest 360 data points was set as the maximum. This normalization range, derived from the vehicle condition, was then applied to the corresponding paired danuglipron or liraglutide data for the same mouse. A moving average with a window and bin size of 10 smoothed the data, which were then plotted as a heat map. Mice with missed virus injection or off-target fibre placement were excluded from analysis.
Dopamine recordings (dLight)
Mice (WT, Glp1r-Cre or Glp1rS33W) were single-housed and habituated to the fibre-optic cable and HFD for 1 hour over 2 consecutive days. On the test day, mice received an injection of either a drug (liraglutide (0.3 mg kg−1), danuglipron (30 mg kg−1) or orforglipron (1 mg kg−1)) or vehicle on different days, with the order of drug versus vehicle injections randomized. Liraglutide and danuglipron were administered 2 hours before recording, whereas orforglipron was given 4 hours earlier. Fibre photometry data were recorded as described above. Fluorescent signals were collected from both dopamine-dependent (465 nm) and dopamine-independent isosbestic (405 nm) excitation wavelengths. During the testing sessions, small pellets of HFD (around 10 mg) were dropped into a cup at 2-minute intervals after the mice retrieved the pellet. Five to six trials were done per mouse. The recording session was video-recorded to time-stamp food retrieval time, and recordings were done during the light phase, between ZT3 and ZT6.
Dopamine analysis (dLight)
The isosbestic signal (405 nm) was fitted to the dopamine-dependent (465 nm) signal using a linear least squares method implemented in a custom MATLAB script. Then ΔF/F was calculated as (465 nm – fitted 405 nm)/fitted 405 nm. To account for inter-mouse differences in signal intensities, z-scores were calculated for the ΔF/F signals. The baseline period for each food trial was defined as the 30-s interval before food retrieval. The mean and standard deviation of the baseline period were used to compute the z-scores, with the formula: z-score = (F − Fμ(baseline))/s.d.(baseline), where F is the 405 nm corrected 465 nm signal (ΔF/F), μ(baseline) is the mean and s.d.(baseline) is the standard deviation of the baseline period. Video frames were analysed to determine the exact time stamp when the mouse retrieved the pellet, which was defined as time 0 for each retrieval. The 30-s window centred around the food retrieval time was extracted. The AUC and maximum fluorescence z-scored within the food retrieval window were further extracted and analysed for quantification of dopaminergic activity. Five food trials were averaged per mouse. Mice with missed virus injections or off-target fibre placements were excluded from the analysis.
Data and statistical analyses
Sample sizes were determined on the basis of our previous experiments and experience with these assays, with the goal of providing adequate power to detect biologically meaningful effects. No formal a priori statistical power calculations were performed. Investigators were not blinded to group allocation during data collection and analysis, except where explicitly indicated. All data are presented as mean ± s.e.m. unless otherwise noted. Statistical tests including paired or unpaired two-tailed t-tests, Kruskal–Wallis tests, Wilcoxon signed-rank tests, one-way ANOVA, two-way or repeated-measures ANOVA (with Bonferroni correction or Tukey’s HSD post-hoc tests) and linear mixed-effects models with beta regression (with Holm post-hoc test) were performed using RStudio (v.4.1.2, v.4.3.0), Python (v.3.11.5), JupyterLab (v.3.6.3), MATLAB (R2024b), GraphPad Prism (v.10.4.0) or Microsoft Excel (2024). Brief descriptions of all experiments in each figure panel, sample sizes, mean ± s.e.m, statistical tests, test statistics and P values are presented in Supplementary Table 1. *P < 0.05, **P < 0.01 and ***P < 0.001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.