Experimental model and subjects
Plant materials and growth conditions
Arabidopsis thaliana Col-0 was used as the wild-type ecotype. The genotypes RAP2.121–28–FLuc, RAP2.12–GFP and Δ13RAP2.12–GFP were as previously described2,3. Seeds were sown in a 3:1 soil/vermiculite mixture, stratified at 4 °C in the dark for 3 days, then germinated at 22 °C/20 °C with a 16 h:8 h light/dark photoperiod and 100 μmol photons m−2 s−1 intensity. For in vitro propagation, seeds were sterilized using 70% ethanol for 1 min and incubated in 10% sodium hypochlorite (NaClO) for 10 min, followed by 6 washes in 1 ml sterile distilled water. For growth in liquid medium, 100 μl of seed suspension, corresponding to 20–40 seeds, was inoculated in 1 ml of sterile half-strength MS medium (basal salt mixture 2.15 g l−1, pH 5.7) supplemented with 1% sucrose in each well of 6-well plates. For growth in solid media, seeds were incubated in the dark at 4 °C for 2 days and subsequently on half-strength MS medium59, supplemented with 1% (w/v) sucrose and 0.8% (w/v) agar, and grown at 22 °C with a 16:8 day/night photoperiod at 100 μmol photons m−2 s −1 intensity.
Yeast strains and culture
A haploid parental strain BY4742 (Matα; His3-Δ1; Leu2-Δ0; Lys2-Δ0; Ura3-Δ0; Euroscarf #Y10000) was cotransformed following the LiAc/SS carrier DNA/PEG method60 with PCO4–pAG415GPD and different versions of the DLOR–pAG413GPD (C-, D- or R-DLOR). All plasmids used for yeast expression were produced in previous work61. Before transformation, cells were grown at 30 °C on YPDA (20 g l−1 peptone, 10 g l−1 yeast extract, 20 g l−1 glucose (Duchefa) and 20 mg l1 adenine hemisulfate (Sigma-Aldrich), supplied with 20 g l−1 agar (Duchefa) when necessary). Transformants were selected on SD medium containing 6.7 g l−1 yeast nitrogen base (DIFCO), 1.37 g l−1 yeast dropout medium (Sigma-Aldrich) and 20 g l−1 glucose, plus supplements (0.16 M uracil, 0.8 M histidine–HCl, 0.8 M leucine and 0.32 M tryptophan (Sigma-Aldrich) when complete), with 20 g l−1 agar when solid.
Bacterial strains
Bacterial strain Escherichia coli BL21 (DE3) was used for expression of recombinant PCOs. Bacteria were cultured in 2YT medium at 37 °C until the optical density at 600 nm (OD600) reached 0.6. Protein expression was induced by addition of 0.8 mM isopropyl-β-d-thiogalactoside (Sigma-Aldrich) at 18 °C for 16 h with shaking at 170 rpm in an incubator (Eppendorf).
Method details
DNA construct generation
For generation of the 35S:RAP2.3–nLuc construct, an 806-nucleotide synthetic string containing an Arabidopsis codon-optimized nLuc sequence including the RAP2.2 intron (Supplementary Table 7) was synthesized in the pMK-RQ backbone by GeneArt (Thermo Fisher Scientific). The destination vector pK7GWnL2 was generated through ligation between pK7GW2 (ref. 62) and GWnLuc-intron after restriction using XbaI and MluI (Thermo Fisher Scientific). The Arabidopsis RAP2.3 CDS was amplified from Col-0 complementary DNA without stop codon (RAP2.3Δstop), with overlapping AttB sites introduced by PCR using GoTaq DNA polymerase (Promega). Entry clone vector was then generated as a BP reaction between the RAP2.3 CDS PCR product and pENTR/D-TOPO (Life Technologies). The resulting entry vector was recombined into the generated pK7GWnL2 destination vector using LR clonase mix II (Thermo Fisher Scientific). Primers used for RAP2.3Δstop cloning and screening are listed in Supplementary Table 8. For generation of the 35S:RAP2.3–GFP construct, the entry vector containing RAP2.3 CDS was recombined with a pK7GW2F62 destination vector using LR clonase mix II (Thermo Fisher Scientific).
For HRPE–nLuc construct design, DNA containing the SacI-attR1-ccdB-attR2-NanoLuc-HindIII sequence (Supplementary Table 7) was de novo synthesized by the GeneArt service (Thermo Fisher Scientific) and cloned into the pBGWL7 Gateway destination vector63 The entry vector containing the HRPE:5′-UTR 35S sequence64 was recombined into the destination vector by Gateway cloning.
For Flag-tagged RAP2.12, the Golden Braid cloning system65 was used. The RAP2.12 CDS was resynthesized to substitute the 129–195 nucleotide (non-conserved) region with a plant-optimized sequence coding for a 3×Flag tag, and a 3×HA tag was added to the end of the CDS before the stop codon (Supplementary Table 7). This DNA string was cloned in the pUPD2 plasmid (elements B3–B4) using BsmBI and T4 ligase. The promoter of RAP2.12 (2,196 nucleotides upstream of the first ATG codon) was synthesized with the BsaI site (483 nucleotides) removed and cloned into pUPD2 to generate an A1–B2 element level 0 plasmid. An Alpha2-level vector was generated by assembling the RAP2.12 promoter (A1–B2), the double-tagged (Flag/HA) RAP2.12 CDS (B3–4), an extra eGFP tag (B5) and the NOS terminator (B6–C1), using BsaI and T4 DNA ligase. The resulting expression cassette was assembled with the promOLE1:gOLE1–TagRFP:Tnos cassette (Alpha1) into a binary Omega1 plasmid using BsmBI and T4 DNA ligase.
The DNA constructs to overexpress fragments of RAP2.12 were also generated the Golden Braid cloning system. Level 0 was created using BsmBI and T4 DNA ligase with PCR products corresponding to CDS portions coding for the desired RAP2.12 fragments. Alpha2 plasmids were generated to include the expression cassette prom2x35S:RAP2.12 fragment–nLuc:Tnos using BsaI and T4 DNA ligase. The resulting expression cassette was assembled with the promOLE1:gOLE1–TagRFP:Tnos cassette (Alpha1) into a binary Omega1 plasmid using BsmBI and T4 DNA ligase.
For recombinant protein production, the five pco genes from Arabidopsis had previously been cloned into the NdeI and XhoI sites of pET28a (Novagen) and transformed into E. coli NEB5α competent cells (New England Biolabs), and the sequences were validated by Sanger sequencing (Source Biosciences), as previously described66.
Arabidopsis transformation
Agrobacterium-mediated transformation was performed to obtain RAP2.3–nLUC, RAP2.3–GFP and HRPE:nLuc stable transgenic lines using floral dip medium as previously described67. T0 seeds were selected for resistance on agarized half-strength MS medium supplemented with the corresponding antibiotic and subsequently transferred in soil. The presence of the transgene was detected by PCR using GoTaq DNA polymerase (Promega). T3 generation plants were used for the experiments.
Low oxygen and reoxygenation treatments
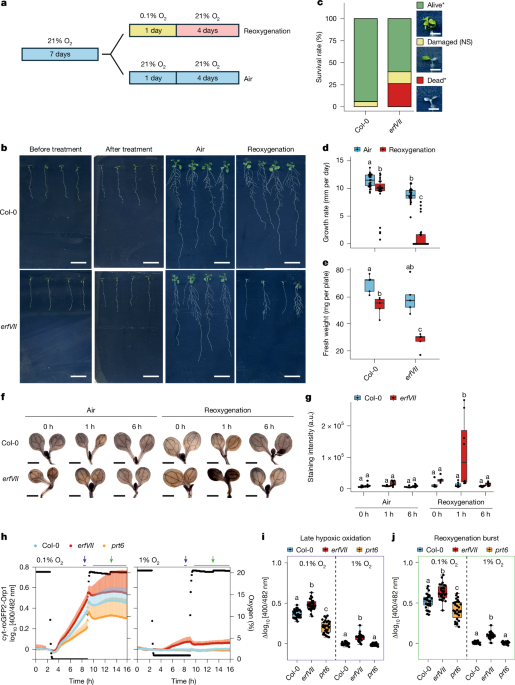
For hypoxia treatments, seedlings were grown in six-well plates in liquid media and subjected to anaerobic conditions inside Hypoxic Workstations (Whitley) continuously flushed with an artificial humidified atmosphere containing a mixture of oxygen (1%) and nitrogen gases (99%) at 22 °C for 6 h. For severe hypoxia treatment, seedlings were grown vertically in square plates and treated with 0.1% O2 v/v O2/N2 for 24 h. During the hypoxic treatments, the seedlings were maintained in the dark to avoid oxygen release by photosynthesis. Seedlings used for control samples were maintained under aerobic conditions (21% O2 v/v O2/N2) in the dark for equal times. After low oxygen treatment, plants were transferred to aerobic growth conditions for reoxygenation treatment.
Root length and survival rate measurements
To assess reoxygenation tolerance, 7-day-old seedlings were treated as previously described. Four or five plates, each containing five to seven seedlings, were used to test for each condition. Primary root length was measured both before and after 4 days of recovery, and fresh weight and survival rate were assessed following 4 days of recovery. Transparent squared plates containing Arabidopsis seedlings were scanned using an EPSON Perfection V750 PRO scanner with a resolution of 720 dots per inch. Growth rate was measured as increase in length of the primary root divided by days of recovery. Primary root length and lateral root density were assessed using ImageJ68 (v.1.54j).
TBHP treatment
Oxidative stress in plants was induced by treatment with 1 mM TBHP diluted in Milli-Q water for 6 h in normoxia and 6 h in hypoxia. After 3 h, the medium was replenished with TBHP.
Histochemical H2O2 staining and quantification
ROS were visualized with DAB (Fluorochem) staining to detect H2O2 using methods described previously69 with minor modifications. Seedlings were incubated with 1 mg ml−1 DAB, vacuum infiltrated for 5 min and incubated for 4–5 h in the dark with shaking. After staining, seedlings were washed with distilled water and bleached in several washes of 70% ethanol. Five to eight seedlings were analysed per condition using a Leica M165C stereo microscope with ×2.5 magnification, followed by quantification of pixel intensity using ImageJ68 (v.1.54j).
Evans blue staining for cell viability
Approximately 25–30 Arabidopsis seedlings per treatment were collected at designated time points during hypoxia and subsequent reoxygenation for both Col-0 and erfVII genotypes. Seedlings were incubated in 0.25% (w/v) aqueous Evans blue solution prepared in 0.1 mM CaCl2 (pH 5.6) for 15 min in the dark at room temperature. Following staining, seedlings were washed three times with Milli-Q water to remove excess dye. Root tissues were then visualized using a Leica M165C stereo microscope.
Fluorescent biosensing of oxidative stress in Arabidopsis
Arabidopsis wild-type plant lines stably expressing protein sensor roGFP2-Orp1 with cytosolic and nuclear localization were as described previously16. Sensor lines in erfVII and prt6 background70 were generated by Agrobacterium-mediated transformation using floral dip with a pH2GW7:cyt-roGFP2-Orp1 expression construct. Positive transformants were selected on selection medium with resistance marker hygromycin B on the basis of fluorescence. Measurements were performed using two independent sensor lines to control for any potential effects of sensor insertion loci. Replicates from the two lines were then combined for each genotype. Leaf discs of 5-week-old plants and 7-day-old seedlings were submerged in wells of a 96-well plate filled with 200 µl standard assay medium (10 mM MES (pH 5.8 with KOH), 10 mM MgCl2, 10 mM CaCl2, 5 mM KCl). A single leaf disc was placed in each well with the abaxial side up for the leaf disc experiment, whereas five or six seedlings were used per well in seedling experiments. Ratiometric readout of the biosensor was performed using a multiwell fluorimeter (ClarioStar Plus, BMG Labtech) in top optics mode, using 30 excitation flashes distributed in a 3-mm orbital average diameter (leaf discs) and a 3-mm spiral average diameter (seedlings). Samples of both genotypes were measured side by side using the same gain for the fluorophore channels (Ex1: F:400-10, Ex2: F482-16; dichroic mirror F:LP504; Em: F:520-10) for all samples for maximum comparability. Wild-type plants without sensor expression were included for background correction. Emissions of each sample were collected every 189 s (leaf discs) or 243 s (seedlings).
Oxygen gradients were performed by targeted influx of N2 into the reader system using an atmospheric control unit. For leaf disc experiments, different O2 concentrations were tested on consecutive days using material from the same plants, with consistent measurement parameters. For seedling experiments, different batches of plants of the same age were used for each O2 concentration. In vivo responsiveness of the roGFP2-Orp1 protein sensor was routinely validated after experiments using the same tissue by subsequent treatment with 20 mM DTT to drive the sensor to a fully reduced state, followed by two washes with standard assay medium before addition of 20 mM 2,2′-dithiodipyridine to oxidize the sensor. For data analysis, wild-type autofluorescence was subtracted from biosensor intensities before calculation of 400 nm/482 nm ratios for each time point. Ratio data were log10-transformed to increase symmetry.
Confocal imaging
Seven-day-old seedlings were used for GFP detection after treatment. For nuclear localization, seedlings were stained in phosphate-buffered saline (PBS) containing 1 μg ml−1 4′,6-diamidino-2-phenylindole (DAPI, Thermo Fisher Scientific) and washed three times in PBS. Imaging was performed using a ZEISS LSM 880 Airyscan microscope (Department of Biology, University of Oxford), equipped with a ×25 objective lens, upon laser excitation at 405 nm and collection at 410–495 nm for DAPI imaging, and excitation at 488 nm and collection at 498–560 nm for GFP imaging. Confocal images were analysed using ZEISS ZEN Lite software (v.3.11).
Western blot
Equal amounts of total protein (100 μg) were resolved by 10% SDS–PAGE and transferred to a polyvinylidene difluoride membrane (Power Blotter Pre-cut Membranes) using Power Blotter 1-Step Transfer Buffer (Invitrogen). Membranes were probed with an anti-Flag M2-Peroxidase (HRP) antibody (Sigma-Aldrich, catalogue. no. A8592) at 1:5,000 dilution overnight at 4 °C. Following incubation, membranes were washed three times with PBST (1× PBS containing 0.1% Tween-20) for 5 min each at room temperature. Immunoblots were developed on film using SuperSignal West Atto Ultimate Sensitivity Substrate (Thermo Fisher Scientific) and imaged on the iBright CL1500 Imaging System (Thermo Fisher Scientific). To verify equal protein loading, the membranes were subsequently rinsed with distilled water and stained with 0.1% (w/v) Ponceau S solution in 5% acetic acid for 5 minutes at room temperature to visualize total protein bands. Excess stain was removed by washing the membrane with distilled water until clear background was obtained. The stained membrane was then imaged for documentation.
Luciferase assay
Total proteins were extracted in passive lysis buffer (Promega). Firefly Luciferase activities were measured using a ONE-Glo Luciferase Assay kit (Promega), and the Nano-Glo Luciferase Assay System (Promega) was used to measure the activity of the nLuc enzyme. The luciferase signal was normalized on the basis of the total protein concentration using the Bradford assay71.
ROS pretreatment
For ROS pretreatment, 7-day-old seedlings grown vertically were incubated with 1 mM TBHP for 2 h in the dark. Following pretreatment, seedlings were used for tolerance assays as described above.
Yeast treatments
For TBHP treatments, colonies were inoculated in 5 ml of -His -Leu SD medium, grown overnight, diluted to half in fresh medium and further diluted to OD600 = 0.1. Cultures were grown for 5 h before the treatment. TBHP was then supplied for up to 30 min at different concentrations (0, 0.25, 0.5 and 0.75 mM). Samples of culture (50 µl) were collected for luciferase assays, centrifuged at 15,000 rpm for 5 min, frozen and extracted in 50 ml of PLB. Luciferase was measured using the Dual-Luciferase Reporter Assay System (Promega) as described previously25. One millilitre of culture was used for OD600 spectrometric measurements.
ROS scavenger and inducer treatments
Seven-day-old Arabidopsis seedlings were exposed to high light (1,600–1,800 µmol m−² s−¹) for 15 min to 1 h, or treated with ascorbate (10 mM), cadmium (10 mM), arsenic (10 mM), diuron (1 mM), antimycin A (200 µM) or methyl viologen (1 mM) for 4 h in the dark under air conditions.
Recombinant protein production
Expression and His6-tag affinity purification of AtPCOs were performed as previously described66. Following affinity purification, the His6-tag was cleaved using TEV protease, and the cleaved tag was removed using a HisTrap HP column (GE Healthcare). Proteins were then purified with a HiLoad 26/600 Superdex 75 prep-grade size-exclusion column (GE Healthcare) equilibrated with 50 mM Tris (pH 7.5) and 0.4 M NaCl. Protein purity was assessed with SDS–PAGE.
In vitro H2O2 oxidation assay of PCOs
Recombinant PCO enzymes (AtPCO1 to AtPCO5; 10 μM) were incubated with H2O2 or an equal volume of H2O in 50 mM HEPES buffer at pH 7.4 (herein termed reaction buffer) at 4 °C for 30 min. Excess H2O2 was removed using a Micro Bio-Spin P-6 chromatography column (Bio-Rad) equilibrated with reaction buffer. Then, 200 μM RAP22–15 peptide (CGGAIISDFIPPPR, purchased from GL Biochem, China) was reacted with 1 μM PCO (H2O2 treated or non-treated) at 25 °C for the required time. For determination of half-maximal inhibitory concentration, 2 µM AtPCO1, AtPCO2 or AtPCO4 was incubated with a series of H2O2 concentrations (0–2 mM) at 4 °C for 30 min. Subsequently, 100 µM RAP22–15 peptide was incubated with 1 µM PCO or with buffer alone (H2O2-treated or untreated) as a control at 25 °C for the specified reaction time. After each reaction, 5-μl samples were quenched in 45 μl of 5% formic acid to stop the enzymatic reaction. Peptide masses were subsequently analysed using an Agilent RapidFire RF360 sampling robot connected to an Agilent 6530 Accurate-Mass Q-ToF mass spectrometer operated in positive electrospray mode. Product distributions were assigned on the basis of the relative integrated areas of peaks corresponding to products of interest. Spectra were visualized using Qualitative Analysis (v.B.07.00), and Agilent RapidFire Integrator (v.4.3.0.17235) was used to calculate integrated peak areas.
H2O2 oxidation assay of RAP22
–15 peptide
Stock solutions of RAP22–15 (200 µM) were prepared in reaction buffer and treated with 400 µM freshly prepared DTT for 45 min at room temperature (25 °C) to ensure peptides were monomeric (in all experiments unless otherwise specified). Peptides were then treated with H2O2 under the conditions required by the experiment. Time-course experiments were conducted in 2-ml deep-welled plates and analysed in real time using RapidFire mass spectrometry as described above.
Peptide fragmentation by LC–MS/MS
DTT-reduced RAP22–15 (20 µM) was treated with 1 mM H2O2 at room temperature for 1 h. A 5-µl sample was diluted with 45 µl 5% formic acid for measurement. LC–MS/MS was carried out using an Acquity UPLC system coupled to a Xevo G2-XS Q-ToF mass spectrometer on a Chromolith Performance RP-18e 100-2 mm HPLC column (Merck) at 40 °C as above. Ions with an m/z ratio of 1,474.7 (+32 Da of AtRAP22–15) were selected for sequential fragmentation under a collision energy of 80 V. Spectra were visualized using Qualitative Analysis (v.B.07.00). Fragments were assigned by comparison of the obtained spectrum with computationally predicted fragment patterns, calculated using the web tool Protein Prospector (v.6.3.1; University of California, San Francisco).
Protein analysis
For protein mass measurement, 100 μM of recombinant AtPCO4 enzyme was incubated with 1 mM H2O2 or an equal volume of H2O in reaction buffer for 1 h at 25 °C. Excess H2O2 was removed using a Micro Bio-Spin P-6 chromatography column (Bio-Rad) equilibrated with reaction buffer. The total mass of the H2O2-treated or non-treated AtPCO4 was measured using RapidFire mass spectrometry as described above.
Cysteine oxidation detection
H2O2-treated or non-treated AtPCO4 was incubated with 1 mM BioDiaAlk in the dark for 1 h at 25 °C, followed by reduction with 10 mM DTT for 1 h at 25 °C. Equal amounts of each protein were separated by SDS–PAGE and transferred on to polyvinylidene fluoride membranes, followed by streptavidin–HRP blotting at 1:1,000 dilution or anti-his-HRP blotting at 1:10,000 dilution. The protein signal was visualized by chemiluminescence (ECL Plus, Pierce).
LC–MS/MS data acquisition
Recombinant AtPCO4 enzyme (15 μg) was incubated with H2O2 or an equal volume of H2O in reaction buffer for 1 h at 25 °C. After removal of excess H2O2, the enzyme was reduced with 85 mM DTT in 50 mM ammonium bicarbonate (Ambic) for 40 min at 56 °C, followed by incubation with 55 mM iodoacetamide in 50 mM Ambic for 30 min in the dark at room temperature. For elimination of excess iodoacetamide, samples were reduced again with 85 mM DTT in 50 mM Ambic for 10 min in the dark at room temperature. In-solution trypsin digestion was performed by addition of trypsin in a 1:50 (w/w) ratio overnight at 37 °C, followed by desalting using C18 ZipTip. The resulting tryptic peptides were resuspended in 40 μl of Milli-Q water with 2% acetonitrile and 0.1% formic acid, and 2 µl was analysed on a nanoAcquity UPLC system (Waters) connected to an Orbitrap Elite mass spectrometer (Thermo Fischer Scientific) possessing an EASY-Spray nano-electrospray ion source (Thermo Fischer Scientific). The peptides were trapped on an in-house packed guard column (75 μm internal diameter × 20 mm, Acclaim PepMap C18, 3 μm, 100 Å) using solvent A (0.1% formic acid in water) at a pressure of 140 bar. The peptides were separated on an EASY-spray Acclaim PepMap analytical column (75 μm internal diameter × 50 mm, RSLC C18, 3 μm, 100 Å) using a linear gradient (length: 100 min, 3% to 60% solvent B (0.1% formic acid in acetonitrile), flow rate: 300 nl min−1). The separated peptides were electrosprayed directly into the mass spectrometer, which was operated in data-dependent mode using a collision-induced dissociation (CID)-based method that performed beam-type CID fragmentation of the peptides. The instrument was controlled using Orbitrap Eclipse Tune 3.5/3.1 and Xcalibur 4.5/4.4. Full scan mass spectra (scan range: 350–1,500 m/z; resolution: 120,000; AGC target: 1e6; maximum injection time: 250 ms) and subsequent CID MS/MS spectra (AGC target: 5e4; maximum injection time: 100 ms) of the 10 most intense peaks were acquired in the ion trap. CID fragmentation was performed at 35% of normalized collision energy, and the signal intensity threshold was kept at 500 counts.
Processing data
Data were analysed with Peaks v.8.5. The raw MS file was searched against the TAIR database. Trypsin with a maximum of three missed cleavages and one unspecific end was selected as the protease. Carbamidomethylation (cysteine) was set as a fixed modification, and oxidation (methionine) and deamination (asparagine, glutamine) were set as variable modifications. The precursor mass tolerance was set to 15 ppm. Fragment mass tolerances for CID were set to 0.8 Da. All peptides present at −log10[P] > 20 and spectra were manually checked and validated or disqualified.
EPR of AtPCO4
EPR spectroscopy was performed on a Bruker EMXmicro spectrometer with a Premium bridge connected to an ER4122SHQE-W1 cavity fitted to an Oxford Instruments ESR900 cryostat. The microwave source was operated at 9.3891(17) GHz, and spectra were recorded at 10 K with liquid helium cryogen. Protein (200 µM) and control solutions were frozen in liquid N2. Spectra were obtained as two 5-min scans from 10 mT to 650 mT using a time constant of 20.48 ms, a microwave power of 200 µW, modulation amplitude of 1 mT and modulation frequency of 100 kHz.
RNA extraction and real-time qPCR analyses
Total RNA was extracted from 60–80 mg of plant material using the phenol–chloroform extraction method as described previously3. RNA concentration was quantified using a NanoDrop ND-1000 (Thermo Scientific), and RNA integrity was tested on a 1% agarose gel. One microgram of total RNA was subjected to DNase Treatment (Thermo Scientific) and retrotranscribed using a qPCRBIO cDNA Synthesis Kit (PCR Biosystems). Real-time qPCR was performed with a QuantStudio 5 Real-Time PCR System (Applied Biosystems) using Power SYBR Master Mix (Thermo Fisher Scientific). Ubiquitin-10 (AT4G0532) was used as a housekeeping gene for Arabidopsis analysis. Four biological replicates were extracted for each condition, each represented by two technical replicates, and the average expression was calculated. The primer pairs used for real-time RT–qPCR are listed in Supplementary Table 9. The relative expression of each individual gene was calculated using the \(2^-\rmC_\rmt\) method72.
ChIP assay
ChIP was performed using a modified version of the protocol described in ref. 73. Chromatin was extracted from 2 g of 7-day-old Δ13RAP2.12–GFP seedlings grown in sterile liquid half-strength MS medium, supplemented with 1% w/v sucrose, under controlled conditions (16 h:8 h light/dark photoperiod, at 22 °C). Seedlings were treated with 1 mM TBHP, or dimethyl sulfoxide (DMSO) as a control, in 1 ml of fresh liquid 1% w/v sucrose half-strength MS medium for 6 h in the dark. Seedlings were cross-linked by dipping in 1% formaldehyde for 10 min and quenched with 0.125 M glycine under vacuum infiltration for 5 min. Seedlings were blotted on paper tissue to dry them and immediately frozen in liquid nitrogen. Each sample was ground to powder and resuspended in 2.5 ml nuclei extraction buffer (100 mM MOPS pH 7.6, 10 mM MgCl2, 0.25 M sucrose, 5% dextran T-40, 2.5% Ficoll 400, 40 mM β-mercaptoethanol, 1× protease inhibitor cocktail (P8340); Sigma-Aldrich). The resulting suspensions were filtered twice through Miracloth (Millipore, 25 μm pore size), and the flowthrough was spun (10,000g, 5 min, 4 °C) for collection of the nuclei at the bottom of the tube. The supernatant was removed, and the pellet was resuspended in 75 μl nuclei lysis buffer (50 mM Tris-HCl pH 8.0, 10 mM EDTA pH 8.0, 1% SDS) and then incubated on ice for 30 min. Samples were diluted by addition of 625 μl ChIP dilution buffer (16.7 mM Tris-HCl pH 8.0, 167 mM NaCl, 1.2 mM EDTA, 0.01% SDS) and sonicated four times with 95% sonication amplitude (SONICS Vibracell VCX130 sonicator) for 30 s. The volume was adjusted to 900 μl with ChIP dilution buffer containing 1.1% Triton X-100, and the samples were centrifuged (10,000g, 5 min, 4 °C). Clean supernatants were transferred to fresh tubes for the subsequent immunopurification steps, and 18 μl (2%) of each sample was put aside to be used as an ‘input’ control. Then, 5 µg of GFP antibody (Roche, catalogue no. 11814460001) was added to the supernatant with a final concentration of 5.5 ng μl−1, and the antibody was pulled down from the nuclear lysate after sonication using Dynabeads Protein G magnetic beads (Thermo Scientific). At the end of the reverse cross-link step, DNA was purified using a QIAquick PCR Purification Kit (Qiagen) and eluted in a final volume of 30 µl. Enrichment of genes in the chromatin immunoprecipitate was detected through real-time qPCR using a CFX384 Touch Real-Time PCR Detection System (Bio-Rad), with a triple technical replicate for each of the four biological replicates, applying the percent input method. To calculate the ratio between immunoprecipitated DNA and input DNA, log250 was subtracted from the raw Ct values of the input, before the Ct immunoprecipitated value was obtained. The final enrichment was calculated as \(2^-\mathrmddC_\rmt\). The primer sequences used are listed in Supplementary Table 10.
RNA sequencing
For reoxygenation treatment, Col-0 and erfVII seedlings were grown for 7 days in a 6-well plate in vertical media and treated with severe hypoxia (0.1% O2) or air (21% O2) for 24 h in the dark, followed by 3 h or reoxygenation aerobic conditions, in the dark, for 3 h. For oxidative stress treatment, Col-0 and erfVII seedlings were grown for 7 days in a 6-well plate in liquid media, followed by 6 h treatment with 1 mM TBHP, in the dark, or mock treatment. At the end of the treatment, samples were collected and frozen in liquid nitrogen. RNA was isolated using a GeneJET RNA Purification Kit (Thermo Scientific) per the manufacturer’s instructions. RNA sequencing was performed in paired-end mode using Illumina Sequencing PE150 on the NovaSeq 6000 platform (Novogene). Transcriptomic analyses were conducted in R (v.4.3.1). After a quality check using FastQC, we aligned the reads on the A. thaliana full genome (TAIR 10) using Rsubread74 (v.2.16.1) and counted them using featureCounts75 (in the Rsubread package). A multidimensional scaling plot was used to assess similarities and differences between samples on the basis of their gene expression profiles. Differentially expressed genes were identified using edgeR76 (v.3.42.4). Differentially expressed genes with expression fold change of at least |1.5| and false discovery rate less than 0.05 (Supplementary Tables 1 and 2) were selected for subsequent analysis. Gene ontology enrichment analysis of the differentially expressed genes was conducted using clusterProfiler77 (v.4.10.1).
Motif discovery and enrichment
Overlapping downregulated genes in erfVII seedlings compared with the wild type under TBHP and reoxygenation treatments were used for DNA motif discovery with STREME (Sensitive, Thorough, Rapid, Enriched Motif Elicitation) in the MEME Suite (v.5.5.9)78. For each gene, a 2.5-kb genomic region upstream of the start codon was extracted from the A. thaliana TAIR10 reference genome and used as the input sequence set. A shuffled version of the input sequences served as the background control. Identified motifs were subsequently compared with known motif databases using TomTom (MEME Suite v.5.5.9)79. Full results are reported in Supplementary Tables 3–6.
Statistical analyses
Statistical analyses were performed and graphs were made and annotated using GraphPad Prism 10.2.3(403) and R Statistical Software (v.4.3.1). Normal distribution and homogeneity of variance of data were evaluated using by Shapiro–Wilk test and Levene’s test, respectively. Student’s t-test, Mann–Whitney test, analysis of variance or Kruskal–Wallis test followed by Tukey’s HSD post hoc test (P < 0.05) was performed to establish the statistical significance of differences. Additional information is provided in figure legends. Sample sizes were not statistically pre-determined. All statistical analyses are provided in Supplementary Table 11.
Materials availability
All unique and/or stable reagents generated in this study are available from the lead contacts without restriction.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.