Lap-NET1 clinical trial
Study design and participants
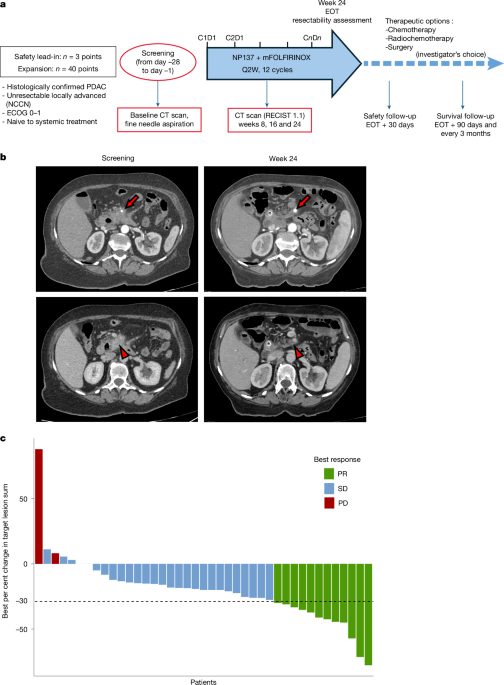
Lap-NET1 is a multicentric, prospective, single-arm phase 1b trial. This study included a safety lead-in phase to confirm the recommended dose of NP137 in combination with mFOLFIRINOX (ClinicalTrials.gov identifier: NCT05546853). Patients were enrolled and treated between March 2023 and June 2024 at nine sites in France.
Eligible patients were 18 years and older with histological or cytological confirmed diagnosis of locally advanced PDAC. The tumour was considered unresectable according to the NCCN Guidelines v2.2021. Prior systemic therapy, radiation therapy or resection for pancreatic cancer were not allowed. Patients had an ECOG performance status of 0 or 1 and a measurable disease per RECIST v1.1.
Ethics statement
The protocol was approved by the ethical committee and the French National Agency for Medicines and Health Products Safety (ANSM); the approvals were recorded via the EU Clinical Trials Information System (CTIS) (CTIS number 2022-500905-42-00). The trial was registered at http://www.clinicaltrials.gov under ID NCT05546853. All participants provided written informed consent prior to enrolment.
Procedures
During the safety lead-in phase, 3 to 12 patients would be enrolled to confirm the dose of 14 mg kg−1 (intravenously) every 14 days for NP137 in combination with mFOLFIRINOX (oxaliplatin (85 mg m−2), calcium levofolinate (200 mg m−2), irinotecan (150 mg m−2) on day 1 and 5-fluorouracil (2.4 g m−2 over 46 h)) administered every 14 days.
After the safety lead-in, 40 additional patients were treated with intravenous NP137 at the recommended dose, associated with modified FOLFIRINOX every 14 days until progression or the maximum treatment duration is reached (6 months, 12 cycles), or unacceptable tolerability. After completion, the investigator chose whether to stop the treatment and follow the patient, or use a follow-up treatment such as radiotherapy, radio-chemotherapy, maintenance chemotherapy or surgical resection (for eligible patients). Patients were followed up until progression and death.
End-points
The primary end-point was the proportion of patients experiencing adverse events of any grade and grade 3/4 adverse events as defined by the National Cancer Institute–CTCAE v5.0 related to the experimental treatment.
Secondary end-points included best overall ORR, disease control rate, duration of response, proportion of surgical resection, PFS and OS.
Exploratory analysis included the effect of specific mutational and expression status, including EMT markers, associated with the patient’s benefit. Descriptive study in spatial transcriptomics and IHC of the evolution of EMT markers, netrin1 expression and tumour microenvironment by comparing pre-therapeutic biopsies to tumour samples obtained during surgical tumour resection.
Statistical analysis
The data cut-off date was May 2025, and the safety analysis population includes all patients who received at least one dose of study drug (n = 43). Safety end-points were tabulated by organ class and grade, using NCI-CTCAE whenever possible. The efficacy evaluable set includes all ITT patients except for ORR including treated patients with at least one post-baseline tumour imaging scan. Descriptive statistics were used to summarize trial results. Continuous variables were summarized by their median and interquartile range, and qualitative variables by counts and percentages. Objective response was expressed as empirical proportions with their exact 95% confidence interval calculated using the Clopper–Pearson method. Duration of response, PFS and OS were summarized using the Kaplan–Meier method, and median survival times with their 95% confidence interval were provided. All analyses were done using R v4.3.
Laser capture microdissection
Tumour samples were formalin-fixed paraffin-embedded (FFPE) for further histologic analysis. Paraffin blocks were sectioned at 3 μm and dried for 1 h at 65 °C. One slide was submitted to HPS histological staining in order to determine tumour cell clusters. Five additional slides were stored at –80 °C prior to LCM. For each experiment, slides were deparaffinized and stained as follows: 2 min xylene, 1 min ethanol (100%), 30 s ethanol (95%, 75%, 50%), 30 s cresyl violet (1%), followed by 30 s ethanol (50%, 75%, 95%) and 2× 30 s ethanol (100%).
LCM was performed on a PALM MicroBeam system (v4.6, P.A.L.M. Microlaser Technologies) with RoboSoftware. Regions of interest were identified on serial scans at 5× magnification and precisely dissected at 20×. Because archived glass slides without membrane coating were used, dissection was carried out in ROBO-LPC mode, allowing automated catapulting of tissue fragments by multiple laser pulses. Laser settings (power and duration) were optimized for capture efficiency.
Microdissected material was collected in 0.5 ml Zeiss adhesive cap tubes (Carl Zeiss Microscopy; one tube per slide). Dissections were completed within 60 min, with dissected areas ranging from ~7 × 104 to 1.4 × 106 µm2. RNA was extracted in 150 µl PKD buffer using the RNeasy FFPE Micro Kit (Qiagen) according to the manufacturer’s protocol. RNA integrity (RIN) and DV200 were assessed with the Agilent 2100 Bioanalyzer (Agilent Technologies).
Whole transcriptome was captured from 1 ng of tumoural RNA. Libraries were prepared using Takara Bio SMART-Seq mRNA LP reagents following the manufacturer’s recommendations. Libraries were sequenced using the NovaSeq6000 Illumina sequencer in 100 bp paired-end.
Bioinformatic analysis
After Illumina sequencing, standard bioinformatics analysis were used to generate fastq files, followed by quality assessment (fastqc v0.11.9 and MultiQC v1.23). Sequencing reads were trimmed to remove adapter sequences and low-quality bases using Trim Galore! v0.6.10. Trimmed reads were then aligned to the human reference genome (GRCh38/hg38) using the align function of Rsubread v2.18.0. Following alignment, gene-level read counts were quantified from the aligned BAM files using the featureCounts function from the Rsubread package.
The counts were then loaded into a DGEList object using the edgeR package v4.6.2 for downstream differential expression analysis. Genes with low expression were filtered out using the edgeR filterByExpr function. The calcNormFactors function was used to normalize for library size and RNA composition bias. DEGs were extracted based on an FDR-adjusted P value < 0.05. Counts per million (CPM) values were calculated with a prior count of 2 and log-transformed for visualization and estimation of signature scores using singscore v1.28.0.
Molecular classification was performed using tte PDACMOC R package v2.5.2 (ref. 26) to classify samples based on several established PDAC molecular subtyping schemes, including those from Collisson27, Moffitt22, Bailey28, Puleo29, Chan-Seng-Yue30 and a PDAC consensus classifier. In addition, the PurIST (Purity Independent Subtyping of Tumors) algorithm21, specifically developed for pure tumour samples, was applied to the count data to distinguish between basal-like and classical subtypes. A PurIST basal-like score (PBP) greater than 0.5 was used to classify samples as basal-like, while scores less than or equal to 0.5 were classified as classical.
GSEA was performed to identify biological pathways and functions enriched in the DEGs between experimental groups (for example, pre- versus post-treatment, or PR versus SD, according to the RECIST 1.1 classification). The analysis was conducted using the GSEA function of clusterProfiles v4.16.0 and the MSigDB H (Hallmarks) gene sets.
Raw counts of the dataset GSE253260 corresponding to the study by Nicolle et al.23 were downloaded directly from the GEO website and processed with edgeR, as described above.
Survival curves were estimated using the Kaplan–Meier method, and survival outcomes were compared between groups with a log-rank test, using the survival v3.8-3 package.
All R/Bioconductor analyses were based on R v4.5.1 (2025-06-13) (https://cran.r-project.org/) using Rstudio software (https://posit.co/download/rstudio-desktop/). Most visualizations are based on ggplot2 (https://ggplot2.tidyverse.org/).
To infer tumour purity, we used the ESTIMATE algorithm31, which uses single-sample GSEA (ssGSEA) with stromal and immune cell signatures. To this end, ESTIMATE scores were calculated in normalized counts of the Lap-NET1 dataset and next transformed to relative purity scores for visualization.
In some analyses, the Lap-NET1 dataset was integrated at the count level with the Linehan-seq dataset (GSE131050) originally described by Nywening et al.32. This cohort includes RNA-seq data of LAPC samples obtained with fine needle aspiration, including pre-treatment samples for nine patients subsequently treated with FOLFIRINOX, five of which also having matched post-treatment RNA-seq data. To minimize batch effects, we used raw fastq data (Sequence Read Archive (SRA) study SRP197641) processed with the same alignment/mapping pipeline (Rsubread) for both datasets before integration. In the combined dataset, we isolated the unique effect of NP137 that goes beyond the effect of FOLFIRINOX, by using a paired interaction strategy (between pre and post timepoint and treatment).
Neogenin IHC
IHC was performed on the Leica Bond RX automated immunostainer (Leica biosystems) using Bond Polymer Refine Detection Kit according to the manufacturer’s instructions. Slides were dewaxed at 72 °C with Bond Dewax Solution (Leica Biosystems, AR9222) and antigen retrieval was performed using Bond Epitope Retrieval Solution 2 (ph9, Leica, AR9640) for 20 min at 97 °C. Sections were incubated with a rabbit recombinant monoclonal neogenin antibody (diluted at 1:1,000, Abcam, Ab324107) for 30 min. Staining was visualized with 3,3′-diaminobenzidine as a chromogenic substrate. Finally, the sections were counterstained with haematoxylin. Finally, sections were scanned with panoramic scan II (3D Histech, Budapest, Hungary) at 20×. All the scans were interpreted by two independent pathologists specialized in digestive oncology.
Cell line and cell culture
The mouse cell line Panc02 was grown in low-glucose Dulbecco’s Modified Eagle’s Medium (DMEM, Gibco) complemented with 10% fetal bovine serum (FBS, Gibco) and 1% penicillin/streptomycin (Gibco). Cells were maintained in culture at 37 °C under humidified atmosphere consisting of 20% O2 and 5% CO2.
RNA extraction and RT–qPCR
Total RNA was isolated from cells using NucleoSpin RNA Plus kit (Macherey-Nagel) according to the manufacturer’s instructions, after washing the cells with PBS pH 7.4 (Gibco). Expression of mRNA was measured using a NanoDrop1000 (Thermo Fisher Scientific). RNA was retrotranscribed using the T100 ThermoCycler (Bio-Rad) and the iScript cDNA Synthesis Kit (Bio-Rad) according to the manufacturer’s instructions. Quantitatyive PCR with reverse transcription (RT–qPCR) was performed using QuantStudio 3 (Thermo Fisher Scientific) and OneGreen Fast qPCR Premix (Ozyme) according to the manufacturers’ instructions.
Cell lysate collection, western blot and antibodies
Confluent cells were washed with cold PBS and scrapped in lysis buffer (Tris 10 mM pH 7.6; SDS 5; Glycerol 10%; Triton X-100 1%, DTT 100 mM). After sonication, proteins were quantified using Pierce 660 nm Protein Assay Reagent (Thermo Fisher Scientific) and loaded on 7.5% SDS–polyacrylamide gels (Bio-Rad) transferred to nitrocellulose membranes using Trans-Blot Turbo Transfer (Bio-Rad). Membranes were blocked for 1 h at room temperature with 5% of nonfat dried milk for netrin1 GAPDH and with 5% of BSA for neogenin. Staining was performed overnight with following antibodies: anti-netrin1 (diluted at 1:1,000, Ab126729, Abcam), anti-neogenin (diluted at 1:1,000, D7M8E, Cell Signaling Technology) and anti-GAPDH (diluted at 1:1,000, 5174, Cell Signaling Technology). After washing, membranes were incubated with secondary antibody, goat anti-rabbit coupled with HRP (diluted at 1:10,000, ARG65351, Arigo), for 1 h at room temperature. West Dura Chemiluminescence System (Thermo Fisher Scientific) was used to intensify the signal. Imaging was performed using Chemidoch Touch (Bio-Rad).
Wound-healing migration assays
Panc02 cells were transfected overnight with negative control (Life Technologies) or mouse NEO1-targeting siRNAs (L-047750-00-0005, Dharmacon) at a final concentration of 10 nM, using Lipofectamine RNAiMAX reagent (Thermo Fisher Scientific) and OptiMEM medium (Gibco). Cells were then plated in Incucyte ImageLock 96-well plates (Sartorius) at a final concentration of 25,000 cells per well and pretreated for 24 h with netrin1 antibody NP137 or isotypic control NP001 (20 µg ml−1). After 24 h, confluent cells were wounded and washed twice with PBS to remove detached cells. Medium was changed and supplemented with either control, NP137 and/or MEK1/2 inhibitor (PD0325901, 10 nM, Sigma-Aldrich). Wound recovery was monitored for 48 h using Sartorius Incucyte S3 Live Cell-Analysis System.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.