Patients
This study was approved by the University Medical Centre (UMC) Utrecht ethical committee, carried out in accordance with the ethical guidelines and regulations and all patients provided written informed consent. FFPE specimens for immunohistochemistry and spatial transcriptomics were requested from and provided by the UMC Utrecht pathology department. Patient inclusion for the organoid biobank was managed by the Utrecht Platform for Organoid Technology (https://uport.umcutrecht.nl/researcher/en/). The biobank participants were 16 patients suspected of having early-stage CRC who underwent surgery for removal of the primary tumour, instead of endoscopic removal, owing to inaccessibility of the tumour. Clinical data from patients featured in this study can be found in Supplementary Table 1.
GeoMx bulk spatial transcriptomics
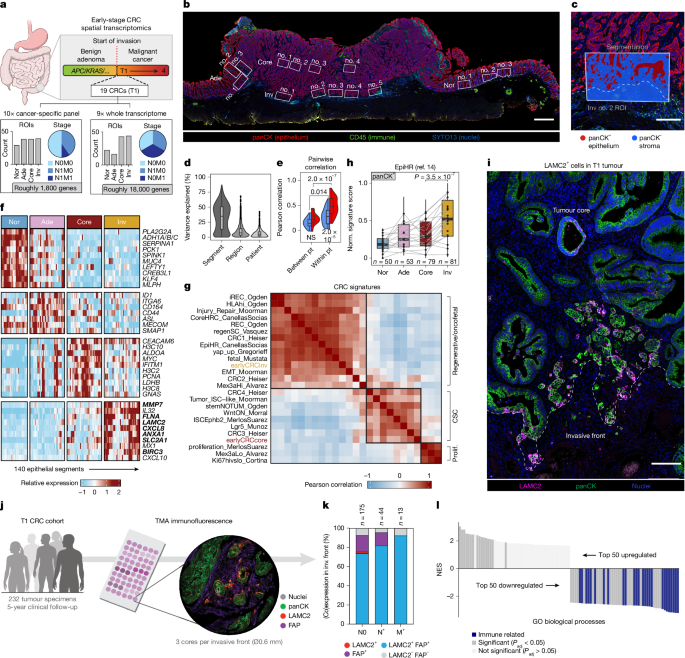
Nanostring GeoMx experiments were conducted with the Utrecht Sequencing Facility (USEQ) and performed as previously described in ref. 65. In brief, 10 T1 CRCs (5× T1N0M0 and 5× T1N1M0) were analysed using the GeoMx CTA (Cancer Transcriptome Atlas) panel and 9 T1 CRCs (3× T1N0M0, 3× T1N1M0 and 3× T1N0M1) were analysed using the GeoMx WTA panel. The specimens analysed by CTA were selected such that risk factors, including lymphovascular invasion, tumour budding, location and morphology were similar between metastatic and non-metastatic primary tumours. Specimens were stained for PanCK (Novus Biologicals, NBP2-33200AF532, 2 µg ml−1) to visualize epithelium, CD45 (Novus Biologicals, NBP2-34528AF594, 5 µg ml−1) to visualize immune cells and SYTO13 (Invitrogen, S7575, 500 nM) to visualize nuclei. ROIs containing 100 to 1,000 nuclei were placed in 4 histopathological regions per tumour: normal tissue adjacent to the tumour, adenomatous tumour component, tumour core and invasive front. Invasive front ROIs were consistently placed, with epithelial tumour strands penetrating the supportive tissue for roughly three-quarters of the ROI edge perpendicular to the tumour border. After ROI placement, PanCK immunofluorescence was used to segment epithelial (PanCK+) and stromal (PanCK−) compartments for separate transcriptomic profiling. For the CTA cohort, CD45 negative and positive areas within the stromal compartment were profiled separately, but were summed during analysis for comparability with the WTA experiment. Standard quality control (unified quality control threshold) was applied to both experiments and can be viewed in Supplementary Reports 1 and 2. In total 426 (CTA) and 285 (WTA) ROIs were sampled across all specimens of which 373 and 281 ROIs were retained after quality control for the CTA and WTA experiments, respectively. At the gene level, 1,781 out of 1,812 and 18,441 out of 18,677 genes were retained after quality control for the CTA and WTA experiments, respectively. Probe counts were aggregated per gene target, Q3 normalized, batch corrected (with ‘slide name’ as the batch to be corrected for) and log2 transformed. For all downstream analyses, sample pt17 (T1_NANO_013) was excluded, because it is classified as a T3 tumour. For variance partition analysis the VariancePartition66 (v.1.38.1) R package was used. To compare different tissue regions within a specimen and across different specimens, we used a linear mixed model approach to model the normalized expression separately for epithelial and stromal segments: log2(gene) ~ tissue region + (1 + tissue region | patient ID). For gene set enrichment analysis (GSEA), two methods were applied: preranked GSEA (fgsea67 v.1.24.0) and single-sample GSEA (ssGSEA68 implemented in GSVA v.1.46.0). Gene sets tested originated from MsigDB (https://www.gsea-msigdb.org/gsea/msigdb), from this study or from published literature (summarized in Supplementary Table 3).
GeoMx CMS classification
Regions from the WTA cohort were used for CMS19 and iCMS classification20. For CMS classification, raw transcript counts of adjacent PanCK+ and PanCK− segments were summed per area of interest and thereafter summed by patient ID and tissue region. Patient F was excluded from this analysis, because the PanCK− and PanCK+ segments were not located within the same areas of interest. These pseudo-bulk samples were used as input for CMScaller19 (v.2.0.1), which was run with ‘RNAseq = TRUE’ alongside default parameters. Finally, the fraction of stromal nuclei for each area of interest was calculated. For iCMS classification CMScaller was run with raw PanCK+ gene counts only and ‘RNAseq = TRUE’. CMS2 and iCMS3 Up gene sets20 were used as templates to classify the segments.
GeoMx CNA prediction
Copy number alteration (CNA) profiles of epithelial cells from the different histopathological regions were estimated using inferCNV (v.1.14.2; ‘cutoff = 0.1’; using normal tissue as a reference group and excluding chromosome XY and mitochondrial genes). Chromosome arm gains and losses were defined as an average residual expression of more than 1.1 or less than 0.9 across all genes on that arm, respectively. Short arms of acrocentric (13p, 14p, 15p, 21p, 22p) and both arms of sex chromosomes were excluded. To calculate pairwise cosine similarities among ROIs from the same tumour, the average residual expression per chromosome arm was rounded to the nearest decimal.
Immunohistochemistry of CRCs
Spatial transcriptomics findings were validated with immunohistochemistry labelling on consecutive slides of the selected T1 tumours. Here 5-µm thick FFPE-embedded tumour sections were mounted on glass slides and baked in at 60 °C for 1 h. Deparaffinization and rehydration was performed as follows: xylene (3 min, 1 change), 96% ethanol (3 min, 1 change), 70% ethanol (3 min, 1 change), rinse in deionized water and rinse in tap water. Heat-mediated antigen retrieval was performed for 20 min in 50 mM Tris/1 mM EDTA pH 9.4 buffer at 95 °C. The following primary antibodies were used: SFRP2 (PA5-29390, Invitrogen, 1:200), LAMC2 (AMAb91098, Atlas Antibodies, 1:500), PanCK (AlexaFluor 532 conjugated; NBP2-33200 Novus 1:500 and NBP3-08398 Novus 1:300) and DNA Syto 13 (S7575, Invitrogen, 1:10,000). The following secondary antibodies were used: Alexa 594 anti-rabbit (Invitrogen A11037; 2 µg ml−1) and Alexa 594 anti-mouse (Invitrogen A11032; 2 µg ml−1). Slides were scanned on the GeoMx Digital Spatial Profiler (Nanostring) with a ×20 0.45 numerical aperture objective and analysed using the QuPath (v.0.6.0) Instanseg extension69. In brief, we quantified all cells within the invasive front (1 mm deep, measured from tumour margin), irrespective of tumour width. Within invasive fronts, nuclei and epithelial cell bodies were segmented on the basis of Syto13 and PanCK pixel intensities, after which percentages of LAMC2+ cells (in epithelium) and the percentages of SFRP2+ and FAP+ cells (in stroma) were calculated. Quantifications were visualized with GraphPad Prism (v.10.4.1).
T1 CRC tissue microarray
A cohort of 261 patients with non-pedunculated T1 CRC were selected from a Dutch multicentre CRC cohort study37. This case cohort consists of 50% random patients of a larger T1 cohort, supplemented with 50% of patients with an endpoint of interest (lymph node metastases and/or recurrence) as previously described. For each tumour specimen, three cores (Ø 0.6 mm) were punched out of both the tumour centre and invasive front and set into paraffin blocks using an automated tissue microarray. Tissue microarray blocks were cut into 4-µm thick sections and stained with antibodies against nucleus, PanCK, LAMC2 and FAP as described in the ‘Immunohistochemistry of CRCs’ section. After quality control, 232 tumours were analysed and quantified (175 N0M0, 44N+, 13M+), using QuPath (v.0.6.0) software for visualization and GraphPad Prism (v.10.4.1) for visualization.
Organoid biobank
Organoid cultures were generated from punch biopsies (Ø roughly 3 mm) of fresh, surgically removed CRCs. Sampled histopathological regions included: normal tissue adjacent to the tumour, adenomatous tumour component, tumour core (carcinoma) and the invasive front. After sampling of a fresh tumour by punch biopsies, the remaining tumour specimens were fixed, embedded in paraffin, sliced and stained with H&E for validation of accurate sampling by histopathological examination of the tissue surrounding the holes resulting from the punch biopsies. For organoid derivation, punch biopsies were minced with scissors and subjected to enzymatic digestion at 37 °C for 15–25 min with 1 mg ml−1 collagenase (Sigma C9407) and 1 mg ml−1 Dispase II (Gibco 11510536) in basal medium (advanced DMEM (Gibco) supplemented with 1% HEPES buffer (Gibco), 1% GlutaMAX (Gibco) and 1% Penicillin/Streptomycin (Lonza)). The resulting tissue fragments were washed 3 times by means of centrifugation (500g, 4 min) and resuspension in 2 ml of basal medium, and then split into a fraction used for cryogenic preservation in Recovery Medium (Gibco, 11560446) and a fraction used for organoid derivation. The latter was resuspended in ice cold Matrigel (Corning) and plated in domes in prewarmed plastic culture plates. Following solidification (37 °C, 15 min) of the Matrigel, organoid culture medium (basal medium with 0.5 nM Wnt surrogate-FC fusion protein (U-Protein Express), 20% R-spondin conditioned medium (in-house production), 10% Noggin conditioned medium (in-house production), 1× B27 (Invitrogen), 1.25 mM N-acetylcysteine (Sigma-Aldrich), 50 ng ml−1 recombinant human EGF (Invitrogen), 50 ng ml−1 recombinant human insulin-like growth factor 1 (IGF1) (Biolegend), 50 ng ml−1 recombinant human FGF2 (FGF-basic, Peprotech) and 500 nM A83-01 (Tocris)), supplemented with 100 µg ml−1 Primocin (InvivoGen) and Rho-kinase inhibitor 10 µM Y-27632 (Gentaur), was added. Organoids were maintained in culture medium without Primocin at 37 °C with 5% CO2 and passaged weekly by trypsinization (37 °C, 1–4 min, Trypsin-EDTA, Sigma T3924). After trypsinization for passaging, medium was supplemented with Y-27632 for 3 days. Cultures were regularly tested for mycoplasma contamination.
The availability of the organoid lines that have been generated in this study is restricted by the UMC Utrecht ethical committee. To receive these organoid lines, a request with the appropriate forms has to be made through this committee, which will determine whether the request corresponds with the informed consent of the patient.
Organoid growth factor-dependency screens
To assess growth factor dependency of organoid lines, organoids were plated as single cells and cultured for 9 days in the presence or absence of indicated growth factors and inhibitors (Nutlin-3 (Sanbio 10004372), 5 ng ml−1 recombinant human TGFB1 (Immunotools 11343160), 20 ng ml−1 recombinant human BMP2 (Immunotools 11343273) and 20 ng ml−1 recombinant human BMP4 (Immunotools 11345043), 1 µM afatinib (SelleckChem)). In brief, organoids were trypsinized with Trypsin-EDTA, filtered with a 30-µm cell strainer (Sysmex), seeded at 3,000 cells per condition in 10 µl Matrigel (Corning) drops on glass bottom 96-well angiogenesis culture and imaging plates (IBIDI), and overlayed with 70 µl of medium. Medium was refreshed on days 3 and 6 after seeding. Organoid growth was monitored by brightfield imaging using an EVOS imaging system (Invitrogen). To assess outgrowth efficiency per condition, the total organoid area on the brightfield images of the ninth day after seeding was determined. For this, images were segmented with OrganoSeg64 software and analysed with a custom ImageJ/Fiji macro.
WGS of organoids
For WGS, DNA was extracted from organoid cultures as early as possible (always before the eighth passage) using the DNA micro kit (Qiagen) according to the manufacturer’s instructions. Truseq DNA nano WGS library preparation and sequencing (Illumina NovaSeq 6000 or X; 2× 150 bp; coverage 15–30×) were performed by the USEQ. Somatic variants were called using the nf-core implementation (oncoanalyser v.1.0.0: https://github.com/nf-core/oncoanalyser of the Hartwig Medical Foundation pipeline (https://github.com/hartwigmedical/pipeline5). The pipeline was run in TUMOR_GERMLINE mode (‘mode’, ‘wgts’). Relevant reference data, prebuilt indices and reference genome (Hartwig human reference GRCh38) were downloaded from the public repository before running the pipeline. SMAD4 heterozygous loss was manually annotated based on CNA data of chromosome 18q.
For construction of phylogenetic lineage trees, short variants shared by many samples from the same patient were called and filtered using joint variant calling by GATK HaplotypeCaller (v.4.1.3, part of the NF-IAP pipeline; https://github.com/UMCUGenetics/NF-IAP). SMuRF (v.3.02, https://github.com/ToolsVanBox/SMuRF) was used to filter somatic variants (absent in the normal samples) from the multi-sample VCF files. High-confident somatic small variants with a variant allele frequency of more than 0.25 in at least 1 sample were included to generate a binary mutation table. The R package ape (v.5.8) was used to construct and visualize the lineage trees.
Plate-based scRNA-seq
To characterize cell type composition in early-stage CRC, we performed scRNA-seq on tissue fragments of five CRCs that were cryopreserved in parallel to organoid establishment of the punch biopsies mentioned in the section ‘Organoid biobank’. For this, tissue fragments were thawed, washed with basal medium and trypsinized to single-cell suspensions using TrypLE (Gibco 12604013) supplemented with 10 µM Y-27632 for 5 min at 37 °C. To distinguish epithelial, immune and stromal cell populations and sort equal amounts of these three populations, single-cell suspensions were stained with DRAQ7 (Invitrogen, 1:200), phycoerythrin anti-human CD326 (EpCAM) (324205 9C4, Biolegend, 1:200) and fluorescein isothiocyanate (FITC) anti-CD45 (368507 2D1, Biolegend, 1:200) in advanced DMEM/F12 for 30 min on ice. Viable single cells (DRAQ7−) were sorted (BD FACSAria III) into 384-well cell-capture plates from Single Cell Discoveries, which contain a 50-nl droplet of well-specific barcoded primers and 10 µl of mineral oil (Sigma M8410). After sorting, plates were briefly centrifuged (500g) and then kept on dry ice until further storage at −80 °C. scRNA-seq was performed by Single Cell Discoveries according to an adapted version of the SORT-seq protocol70 with primers described in ref. 71. Cells were heat-lysed at 65 °C followed by complementary DNA (cDNA) synthesis. After second-strand cDNA synthesis, all the barcoded material from one plate was pooled into one library and amplified using in vitro transcription. Following amplification, library preparation was performed following the CEL-Seq2 protocol72 to prepare a cDNA library for sequencing using TruSeq small RNA primers (Illumina). The DNA library was sequenced by paired-end sequencing on an Illumina NextSeq 500, high output, with a 1× 75 bp Illumina kit (read 1, 26 cycles; index read, 6 cycles; read 2, 60 cycles).
scRNA-seq analysis
For alignment of reads, an adapted version of the nf-core scrnaseq pipeline (v.2.4.0)73 was used (https://github.com/gowanaka/nf-core-scrnaseq). In brief, STARsolo (v.2.7.10b) was used to align reads to a custom GRCh38 human reference transcriptome including External RNA Controls Consortium (ERCC) spike-ins. Following mapping, count matrices were generated with STARsolo (v.2.7.10b). Gene expression was analysed using Seurat (v.5.0.1)74. Cells with less than 25% mitochondrial content, less than 25% exogenous ERCC spike-in content, more than 1,000 transcript counts (nCount_RNA) and more than 500 unique detected genes (nFeature_RNA) were selected for downstream analysis. Mitochondrial transcript counts were removed before count normalization and scaling by the Seurat NormalizeData and ScaleData functions, respectively. Unsupervised clustering was used to cluster cells according to the standard Seurat workflow. Gene expression signature scores were calculated with the Seurat AddModuleScore function. Differential expression analysis was performed with the FindAllMarkers function.
CosMx single-cell spatial transcriptomics
To map spatial distribution of cell types identified with scRNA-seq, we performed Nanostring CosMx single-cell spatial transcriptomics75 on one T1 CRC included in the organoid biobank (pt5/ptD; Human CosMx Universal Cell Characterization Panel; 1,000 gene targets; Fig. 3) and 11 CRC specimens temporally surrounding the moment of malignant transformation (3× intramucosal carcinoma, 5× T1 sm1 and 3× T1 sm3; Human CosMx 6,000 Discovery Panel; 6,000 gene targets, Fig. 5). Slides were stained with segmentation markers (Human Universal Cell Segmentation Kit, RNA, Bruker Spatial Biology, 531-121500020) for nuclei (4,6-diamidino-2-phenylindole (DAPI)), cell membranes (CosMx Hs CD298/B2M Segmentation Marker Mix, Ch2 RNA), epithelial and immune cells (CosMx Hs PanCK/CD45 Marker Mix Ch3/Ch4, RNA, Bruker Spatial Biology) and macrophages (CosMx Hs CD68 A La Carte Marker, Ch5 RNA, Bruker Spatial Biology, 531-121500022, second experiment only). After filtering on the basis of standard quality control, cells were labelled according to predicted cell type using label transfer from the Seurat package (v.5.0.1)74, with our early-stage CRC scRNA-seq dataset as a reference. Query and reference datasets were downsampled to only include overlapping gene targets before label transfer and both were normalized and scaled using the SCTransform method. Principal component analysis (PCA) was performed for the scRNA-seq data. FindTransferAnchor() and TransferData() were used to anchor the scRNA-seq PCA reference data to the CosMx query data and transfer cell type labels. After label transfer, raw CosMx data were normalized and scaled again using SCTransform. PCA was performed on normalized data. Uniform manifold approximation and projection (UMAP) (30 principal components, min.dist = 0.01) was used for dimensionality reduction. Nearest neighbour graphs were constructed using the first 30 principal components. Unsupervised clustering was performed using the Seurat default implementation of the Louvain algorithm (resolution 0.7). In plots where cell type labels are shown, only cells that were annotated with prediction.score.max ≥ 0.6 are shown.
Subclustering of FAP+ CAFs and epithelial clusters (pt5; Fig. 3) was performed using the Louvain algorithm with resolutions 0.2 and 0.05, respectively. FAP+ CAF subclusters were assigned to a CAF subtype on the basis of marker gene expression. The epithelial HRC subcluster was annotated on the basis of marker gene expression. Epithelial subclustering of the other 11 CRC specimens was restricted to the cancer epithelial clusters identified by means of clustering per specimen (resolution 0.7). We selected clusters with high HRC program expression within each specimen separately by reclustering cancer epithelium (resolution 0.7 and 0.2). We did not detect a HRC cluster in specimens T1_NANO_022 (incomplete invasive front), T1_NANO_030 and T1_NANO_031 (both intramuscosal carcinomas). For single-cell spatial plots of epithelial cells, cells were filtered by PanCK staining intensity (lowest tenth percentile excluded).
Neighbourhood analysis
Profiling spatial context of cancer cells, we performed cellular neighbourhood analysis for the oncofetal and cancer stem cells of the 11 CRC specimens analysed with the Nanostring CosMx 6,000 gene panel. In brief, we ran RANN’s nn2() function per sample to find the neighbours of a cancer cell within a 50-µm radius. The output cells × clusters matrix was used to count neighbouring cell types for composition analysis (sccomp76), sum expression profiles across all neighbours for neighbourhood differential expression analysis and to cluster cells on the basis of neighbour cell composition using k means clustering (k = 10).
NicheNet analysis
NicheNet analysis was performed on the GeoMx WTA invasive front segments and CosMx ‘niche3’ cells (oncofetal niche) with nichenetr77 (v.2.0.0; receivers = epithelial segments; senders = stromal segments). Genes with expression below the 25th quantile across all sender or receiver segments were excluded. Ligands of interest were prioritized on the basis of cumulative interactive potential across all the coreHRC genes.
Fibroblast immortalization and culture
Fibroblast lines were derived from early passage cultures of the punch biopsies that were used to establish organoids (‘Organoid biobank’ section). In brief, fibroblasts adhering to the plastic bottom of the organoid culture plates were maintained with DMEM supplemented with 10% fetal bovine serum (Bodinco) and 1% penicillin/streptomycin (Lonza) after organoid removal for passaging and subjected to simultaneous lentiviral transduction with hTERT (third-generation adaptation of Addgene no. 85140) and BMI1 (no. 12240) overnight78. Fibroblast lines were passaged weekly by trypsinization.
Organoid–fibroblast cocultures
Organoids were cocultured with fibroblasts in a transwell setup (Polycarbonate Cell Culture Inserts with 0.4 µm pore size in a six-well plate format, ThermoFisher) for 48 h in growth factor depleted medium (basal medium, B27 (Invitrogen) and 1.25 mM N-acetylcysteine (Sigma-Aldrich)). Fibroblasts were trypsinized, counted and seeded as a single-cell suspension (300,000 cells per well) in fibroblast culture medium (above) on plastic or in 200 µl of Matrigel (Corning) 1 day before coculture to allow for adherence to the plastic substrate. To start coculture, 5-day old organoids were plated in 150 µl of Matrigel (Corning) on top of the transwell membranes. To harvest RNA, transwell culture inserts with organoids were removed and organoids and fibroblasts were lysed separately, followed by RNA extraction using the Nucleospin RNA isolation kit (Macherey-Nagel 740955), according to the manufacturer’s instructions. To investigate matrix-induced and juxtacrine effects, organoids and fibroblasts were seeded simultaneously in collagen-Matrigel (25%/25%) (Collagen Type I Corning 354236) mixtures and cocultured in growth factor depleted medium for 2 days or 5 days before flow cytometric quantification of EMP1-mNeon+ cells.
RNA-seq of organoid–fibroblast cocultures
RNA-seq library preparation was performed by the USEQ according to the Illumina TruSeq stranded PolyA protocol. Libraries were sequenced in two runs on an Illumina NextSeq 2000 (run 1: 20 samples, 2 × 50 bp paired-end sequencing, index 1: 17 cycles, read 1: 50 cycles, index 2: 8 cycles, read 2: 50 cycles and run 2: 11 samples, 1 × 50 bp single-end sequencing, index 1: 17 cycles, read 1: 50 cycles, index 2: 8 cycles). For alignment of reads, the nf-core RNA-seq pipeline (v.3.14.0) was used (https://doi.org/10.5281/zenodo.1400710, ref. 79) with the option ‘star_salmon’. Briefly, FASTQ files underwent quality control (FastQC v.0.12.1), adaptors were trimmed (Trim Galore! v.0.6.7), reads were aligned to the GRCh38 human reference transcriptome (STAR v.2.7.9a) and a gene expression matrix was generated (Salmon v.1.10.1). Differential expression analysis at the gene and gene set level (ssGSEA/GSEA) was performed using DESeq2 (v.1.38.3). Genes that had at least a count of 10 in at least 4 samples were retained, VST normalized and a PCA was conducted. Organoid and fibroblast samples were batch corrected by sequencing run and Line_ID, respectively.
Generation of EMP1
mNeon organoid knock-in
EMP1mNeon knock-in organoids (pt5 inv) were generated by in-trans paired Cas9 targeting as described in ref. 80. SpCas9 (Addgene no. 48139) locus-specific expression vectors were generated according to published protocols81 (guide 5′-TCCTGAGAAAGAAATAAGGC-3′). The targeting vector was generated by introducing 449-nucleotide homology arms and flanking EMP1 guide sequences into a custom-made vector (IRES-mNeon-NLS-P2A-iCasp9-WPRE-pA-PGK-PuroR-pA; Addgene no. 251175) using golden gate assembly. For transfection, organoids were trypsinized to cell clumps containing roughly 5 cells (around 1 × 106 cells in total) and coelectroporated with 4 μg of SpCas9 DNA and 11 μg of targeting vector using the NEPA21 Super Electroporator (Nepagene) following the conditions described in ref. 82. Electroporated cell clumps were plated in Matrigel overlayed with organoid culture medium supplemented with 10 µM Y-27632 Rho-kinase inhibitor for the first 3 days. Targeted cells were selected using 1 μg ml−1 puromycin and maintained as polyclonal populations. To confirm EMP1-mNeon-NLS fluorescence and nuclear localization, live organoids were incubated with Hoechst 33342 (ThermoFisher Scientific 62249, 1:5,000, 30 min, 37 °C with 5% CO2) to visualize nuclei and imaged with a Leica SP8 scanning confocal microscope using LAS X software (v.3.5.7.23225).
EMP1
mNeon organoid reporter-based screen
To screen for ligands that induce oncofetal tumour cell states, EMP1mNeon organoids were trypsinized (TrypLE), plated as single cells (300 cells per μl, filtered with a 40-μm strainer) and treated with candidate ligands 5 days after plating. Single candidate stimuli or combinations were added in growth factor-deprived medium (basal medium with B27 (Invitrogen) and 1.25 mM N-acetylcysteine (Sigma-Aldrich) after 2 washes with basal medium and included: TGFβ1 (5 ng ml−1; Immunotools 11343160), TGFβ3 (5 ng ml−1; Immunotools 11344483), PGE2 (10 μM; Tocris 2296), PGD2 (10 μM; Merck 538909), CXCL12 (40 ng ml−1; Immunotools 11343363), FGF2 (50 ng ml−1; Peprotech 100-18B), IGF1 (50 ng ml−1; Biolegend 590904), FGF7 (50 ng ml−1; Peprotech 100-19), GREM1 (100 ng ml−1; Peprotech 120-42-50UG), SFRP1 (100 ng ml−1; Peprotech 120-29), SFRP2 (100 ng ml−1; Biotechne 1169-FR-025), GDNF (50 ng ml−1; ThermoFisher 450-10-10UG), IL-36A (50 ng ml−1; ThermoFisher 200-36A-2UG), IL-36B (50 ng ml−1; ThermoFisher 200-36B-2UG), CXCL14 (40 ng ml−1; Immunotools 11345190), BMP2 (20 ng ml−1; Immunotools 11343273), BMP4 (20 ng ml−1; Immunotools 11345043), hepatocyte growth factor (50 ng ml−1; ThermoFisher 100-39-10UG), vascular endothelial growth factor (50 ng ml−1; ThermoFisher 100-20-2UG), WNT5A (20 ng ml−1; Biotechne 645-WN-010), IL-6 (100 ng ml−1; Stem Cell Technologies 78050.1), OSM (50 ng ml−1; R&D Systems 295-OM-010), IL-1B (20 ng ml−1; ThermoFisher 200-01B-10UG), tumour necrosis factor (10 ng ml−1; Knoll AG), IL-27 (100 ng ml−1; ThermoFisher 200-38-2UG) and interferon-gamma (100 ng ml−1, ThermoFisher 300-02-20UG). The percentage of EMP1mNeon positive cells among live cells was measured 24 h after addition of candidate stimuli as described below.
Flow cytometry
Single-cell organoid suspensions were prepared by trypsinization with Trypsin-EDTA for 5 min at 37 °C. Flow cytometry measurements were performed on a BD FACSCelesta CellAnalyzer. Single live cells (DAPI−) were gated in the BV421 channel, mNeon and phycoerythrin fluorescence were measured in the FITC-A and PE-A channels, respectively. Gates were set on the basis of negative control samples, that is, parental organoid line or unstained cell suspensions. To separate organoid and fibroblast cells in juxtacrine cocultures (Extended Data Fig. 8g), cells were stained with phycoerythrin anti-human CD326 (EpCAM) (324205 9C4, Biolegend, 1:400). To measure MHCI levels (Extended Data Fig. 8i,j), cells were stained with phycoerythrin anti-human HLA A/B/C (311405 W6/32, Biolegend, 1:400). Flow cytometry data were analysed and visualized using BD FACSdiva software and the free online tool https://floreada.io.
Immunofluorescence of organoids
Organoids form coculture experiments were immunostained for LAMC2 protein levels as described previously83. In brief, organoids were dislodged from Matrigel matrix domes by incubation in basal medium supplemented with 1 mg ml−1 dispase for 30 min at 37 °C/5% CO2 and pelleted after several washing cycles with basal medium. Organoids were fixed in 4% paraformaldehyde in PBS on ice for 45 min. Fixed organoids were transferred to repellent plates (Greiner Bio-One). Permeabilization, blocking and antibody incubation steps were performed with organoid washing buffer (0.1% Triton X-100 in PBS and −0.2% wt/vol BSA) at 4 °C on a shaker. Primary antibodies used: LAMC2 (AMAb91098, Atlas Antibodies, 1:500) and beta-catenin (C2206, Sigma-Aldrich, 1:500). Secondary antibodies used: Alexa 647 anti-mouse (Invitrogen A21236; 1:500) and Alexa 568 anti-rabbit (Invitrogen A11011; 1:1,000) and Hoechst. Organoids were mounted in clearing solution (ddH2O, 60% (vol/vol) glycerol and 2.5 M fructose) and imaged on a Zeiss LSM880 confocal laser scanning microscope at ×40 magnification. Images were processed in Fiji software. Hoechst was used as a nuclear marker and beta-catenin to mark cell boundaries, to allow for LAMC2 quantification at single-cell resolution. Statistical analysis was performed in GraphPad Prism (v.10.4.1).
RNA-seq and qPCR of organoids treated with TGFβ and prostaglandins
Organoids were treated with a combination of TGFβ1 (5 ng ml−1; Immunotools 11343160), TGFβ3 (5 ng ml−1; Immunotools 11344483), PGE2 (10 μM; Tocris 2296) and PGD2 (10 μM; Merck 538909) in growth factor-deprived medium (basal medium with B27 (Invitrogen) and 1.25 mM N-acetylcysteine (Sigma-Aldrich) after 2 washes with basal medium, 5 days after trypsinization to single cells. After 24 h of induction, RNA was extracted using the Nucleospin RNA isolation kit (Macherey-Nagel 740955), according to the manufacturer’s instructions. Library preparation (directional messenger RNA; poly-A enrichment) and sequencing (NovaSeq X Plus Series PE150) were performed at Novogene and data were analysed with DESeq2 (v.1.38.3) and clusterProfiler (v.4.8.3) with method fgsea (v.1.24.0). Differential expression analysis was corrected for Patient ID. For quantitative PCR (qPCR), cDNA was generated from RNA using the iScript cDNA Synthesis Kit (Bio-Rad). For qPCR, 20 ng of cDNA was mixed with 0.5 µM forward and reverse primer each and 5 µl of PowerTrack SYBR Green (Applied Biosystems) per well. qPCR was performed on the Bio-Rad CFX96 and results were analysed with Microsoft Excel (v.16.95) using the ΔΔCt method with ACTB and PBGD as reference genes. Sequences of primers used for qPCR can be found in Supplementary Table 8.
Analysis of published scRNA-seq data
Published scRNA-seq data of human CRCs from refs. 45,55,54 (see ‘Data availability’ for accession codes) were integrated using the Seurat package (v.5.0.1)74 in R (v.4.2.0) with harmony integration according to the standard workflow. Clusters were annotated using cell type annotations included with the published datasets and marker genes of the clusters. For trajectory inference analyses, Monocle3 (ref. 84) (v.1.4.26), CytoTRACE2 (ref. 85) (v.1.1.0) and Slingshot86 (v.2.16.0) R packages were used to calculate single-cell potency and pseudotime scores. The CytoTRACEkernel from CellRank87 (v.2.0.7) was used to compute a transition matrix and construct pseudotime-based streamline plots featured in Fig. 5j. The bottom 2% low density areas in the UMAP space were excluded from these analyses.
Statistics and reproducibility
Statistical analysis was performed as noted in the figure legends using R (R base (v.4.2.0 or later), ggplot2 (v.3.5.1), ggpubr (v.0.6.0) Seurat (v.5.0.1)) and GraphPad Prism (v.10.4.1). Data distribution was assumed to be normal, but this was not formally tested. All statistical tests were two-tailed. Where not stated, P < 0.05 or false discovery rate (FDR) < 0.05 was deemed to be statistically significant. The Benjamini–Hochberg method was used to correct the P value for multiple testing. For comparisons between more than two sample groups, one-way analysis of variance (ANOVA) was performed, using Tukey’s HSD for post hoc analysis. Data are presented as mean ± standard deviation, unless otherwise stated in the figure legend. For GSEA results, an FDR < 0.25 was deemed to be statistically significant in line with ref. 88. Representative images (Fig. 4f and Extended Data Figs. 7a and 9b) depict consistent results that were observed in at least two independent experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.