Experimental methods
Human sample preparation
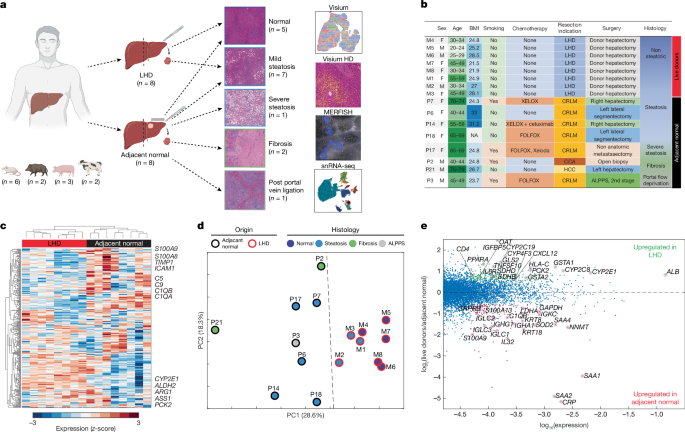
Human samples and clinical data were obtained from patients undergoing either hepatectomy surgery due to a hepatic pathology or a donor hepatectomy for a living donor liver donation (Fig. 1b). In the hepatectomies that were performed due to liver pathologies, a sample was taken at least 5 cm from the closest margin of the pathology. In the hepatectomies that were performed as donor hepatectomy for a living donor liver donation, a liver biopsy of 1 cm³ was obtained immediately upon abdominal opening, prior to mobilization of the liver and initiation of resection. All hepatectomies due to liver pathologies were performed at Sheba Medical Center, Israel with approval from the Sheba Medical Center Helsinki committee. All donor hepatectomies for a living donor liver donation were performed at Mayo Clinic, Rochester, MN, with approval from the Mayo Clinic IRB committee. Informed consent was obtained from all patients, and all experiments followed all the Helsinki committee guidelines and regulations. Samples were taken only in cases where no pathology was observed in the sampling area during pre-operative and intraoperative evaluations. In all cases ~1 cm of hepatic parenchyma distant from the liver capsule area were resected. Tissues were gently washed in PBS and were embedded in OCT for Visium and MERFISH or fresh-frozen for Visium HD.
Non-human sample preparation
Whole washed pig (Sus scrofa domesticus) livers were bought from a butcher at Kibbutz Lahav C.R.O., Israel, which sells meat products from surplus animals. Specimens were adult males (5.5 months, ~85 kg) and an adult female (5.5 months, ~75 kg) raised under farming conditions. Washed cattle (Bos taurus) liver sections were donated by a butcher in Haifa, Israel. Specimens were adult males (17 months, ~670 kg) raised under farming conditions. Wild Boar (Sus scrofa) liver samples were taken from the cadavers of animals euthanized as part of population control regulated hunting done by the Israel Nature and Parks Authority (INPA). The sampling permit was acquired beforehand under INPA regulation. The researchers did not influence the time, location or manner of death; they were solely permitted to take samples from the cadavers. Age and weight estimation of the specimens were evaluated by trained INPA rangers and a veterinarian who participated in the hunting expedition. The boars’ ages were approximately 1–2 years, and they weighed approximately 60 kg. Liver tissues were gently washed in PBS and were embedded in OCT for spatial transcriptomics (Fig. 3b–d and Extended Data Fig. 6).
10X Visium spatial transcriptomics
Fresh-frozen OCT embedded blocks were cryo-sectioned into 10-µm-thick slices and carefully placed on a Visium Spatial Gene Expression slide. Each slide contained 4 slices from 4 different patients, total of 4 Visium slides were used for the 16 human samples. Slides were fixed with pre-chilled methanol at −20 °C, stained for H&E as described in the Visium user guide and were captured using a Leica DMi8 widefield inverted microscope equipped with a Leica DFC7000T colour camera and an HC PL APO 20×/0.80 DRY objective (506530, Leica Microsystems). Permeabilization time was set according to the 10X Tissue optimization protocol, resulting in 24 min for all 16 human and 7 non-human mammalian samples. After tissue permeabilization and transcript capture, libraries were prepared according to the Visium Spatial Gene Expression User guide with 14–17 cycles of PCR for cDNA amplification. Four Libraries for each slide were pooled and loaded at a concentration of 400 pM to a SP100 flow cell and sequenced on Illumina NovaSeq 6000. The Fastq reads generated from the sequencer were preprocessed by 10X Genomics space-ranger software (version 2.0.1) which included spatial de-barcoding, read-alignment to hg38 and UMI-generation. Further preprocessing was done in MATLAB R2022a.
10X Visium HD spatial transcriptomics
Fresh-frozen tissue samples were processed into FFPE blocks. Sections of 5 μm thickness were mounted onto Visium HD slides and deparaffinized according to the Visium HD Spatial Gene Expression User Guide. H&E staining was performed to visualize tissue morphology, and high-resolution brightfield images were captured using a Leica DMi8 widefield inverted microscope equipped with a Leica DFC7000T colour camera and an HC PL APO 20×/0.80 DRY objective (506530, Leica Microsystems). Following imaging, sections were destained and underwent de-crosslinking as described in the Visium HD FFPE Tissue Preparation Handbook. Three specific probes for each targeted gene were added to the deparaffinized, destained, and decrosslinked tissues.
MERFISH spatial transcriptomics
Liver OCT blocks were sectioned at width 10 μm. Sections were mounted on MERSCOPE glass slides (Vizgen) and were fixed with 4% paraformaldehyde pre-warmed to 47 °C for 30 min, rinsed, permeabilized, and preserved in sterile 70% ethanol. The procedures of probe hybridization, gel embedding, and tissue clearing were performed according to the MERSCOPE FFPE sample preparation user guide, using the predesigned pan cancer pathways 500-gene panel. In brief, tissues were dried for 1 h in room temperature to improve slide adherence. Then, tissues were photobleached for 3 h using the Vizgen photobleacher, followed by a de-crosslinking step, gel embedding and tissue clearing for 3 days. Next, sections were incubated with MERFISH probe mix for 48 h at 37 °C.
After probe hybridization, the samples were labelled with DAPI and poly T and placed in the MERSCOPE flow chamber for imaging. The raw image files obtained were processed via the MERlin image analysis pipeline61. Cellpose software version 1.x was used to segment cells from the nuclear DAPI signal. The decoded RNA molecules were then partitioned into individual cells to generate single-cell count matrices.
Immunofluorescence staining
Immunofluorescence was performed on 5-µm paraffin-embedded sections using using the BOND RXm research detection platform (Leica, DS9455). Tissue sections were deparaffinized using BOND dewaxing solution. Antigen retrieval was carried out in heat induced antigen retrieval HIER1 (pH 6) for 10 min.
Subsequently, sections were permeabilized and blocked for non-specific binding in 20% normal horse serum (Vector labs, S-2000) and 0.1% PBST. Primary antibodies against GS (Abcam, ab64613 and ab73593; 1:100), GLUT2 (Abcam, ab54460; 1:100) and DPP4 (a combination of AF1180 and TA500733, 1:100) were diluted in the 2% normal horse serum and 0.1% PBST and incubated for 30 min. After three washes, sections were incubated for 30 min at room temperature with secondary antibodies and Hoechst (invitrogen, H3570) in PBS. The secondary antibodies used were: Cy3-conjugated Donkey anti-Mouse IgG (Jackson ImmunoResearch, 715-165-151; 1:100), Cy5-conjugated Donkey anti-Rabbit IgG (Jackson ImmunoResearch, 711-175-152; 1:100) and Cy3 Donkey anti Goat (Jackson ImmunoResearch, 705-165-147, 1:100).
Slides were washed three times in PBS, mounted with aqua poly mount (polysciences, 1860620), Imaging was done using the Phenoimager HT system (Akoya Biosciences) at 20× magnification.
PhenoCycler antibodies and imaging
Our antibody panel consisted of DAPI nuclear staining and 48 antibodies targeting proteins expressed in diverse immune, stromal and epithelial liver cell types: CD45, CD3, CD69, CD90, CD11c, Ki67, CD8, CD4, HLA-DR (MHCII), FN1, CD31, VIM, PDPN, EGFR, CD279, SOX4, CD14, CD138, PDGFRA, CA9, GLUT1, CD44, CHI3L1, PanCK, CD163, NDRG1, APOD, SOX2, APOE, MT1MMP, S100B, SNAP25, aSMA, CD19, SOX9, TNC, CD24, CD83, SPARC, CD1c, ATRX, POSTN, VCAN, COL1, CD15, TNR, DCLAMP and HSPG2. Staining was performed on two LHD livers (M1, M3) and four adjacent normal livers (P2, P6, P17 and P4).
Each antibody was validated using conventional immunofluorescence on control tissues (liver, duodenum and glioma) for which matched Visium spatial transcriptomic data were available. PhenoCycler Fusion runs were performed according to the manufacturer’s protocol. In brief, fresh-frozen sections were fixed in 4% paraformaldehyde for 20 min, followed by blocking with 4% bovine serum albumin and 0.25% Triton X-100 for 30 min. Primary antibodies were incubated overnight at 4 °C, and secondary antibodies for 2 h at room temperature. All antibodies, including pre-conjugated reagents from Akoya, were further validated on control samples through multiple PhenoCycler runs to optimize antibody concentrations and exposure settings for optimal signal-to-noise ratio.
Single-nucleus isolation form FFPE tissue samples
Three 25-µm-thick tissue sections were washed with 3 ml of Xylene for 10 min to remove paraffin. Subsequently, they were rehydrated via a sequence of 1 ml ethanol washes, each lasting 30 s: 2 rounds in 100% ethanol, then 70%, and finally 50% ethanol. Next, the tissue sections were washed with nuclease-free water and finally with PBS. Tissue disruption was performed using the 37C_FFPE_1 gentleMACS in 2 ml of digestion mix composed of 1 mg ml−1 Liberase TM (5401119001, Roche), in RPMI1640. The separated nuclei were filtered through a 40-μm mesh, double-rinsed with 1× PBS and centrifuged at 850g at 40 °C for 5 min. They were then resuspended in 500 μl Quench Buffer B (10X genomics PN2001300). The concluding nuclear count was performed using the Denovix cell counter.
Single-nucleus RNA profiling from FFPE tissue samples
Gene expression libraries were prepared using the GEM-X Flex Gene Expression Human (1000793 10X Genomics), in accordance with the user guide (CG000787 revision B). Additional optimizations in the pre-amplification and Indexing cycles tailored for nuclei derived from FFPE samples. Over 300,000 nuclei per sample underwent a 20-h hybridization with the BC01-BC04 probe sets. This was followed by three washes using the Post-Hyb Buffer as recommended in the guide. Post-hybridization, nuclei were resuspended in the Post-Hyb Resuspension Buffer, counted, combined to achieve target of 20,000 cells per sample, and loaded onto Chromium X chip for capture, adhering to the guide’s procedures. An eight-cycle pre-amplification and nine-cycle indexing PCR was conducted. Library final concentration of 150 pM with the addition of 15 pM from PhiX library was loaded on NovaX (Illumina) sequencing machine aiming at 20,000 reads per cell with the following setting: Read1, 28 bp; Index1, 10 bp; Index2, 10 bp; Read2, 90 bp. Raw reads were de-multiplexed using Bcl-conver (v4.2.4) and aligned using Cellranger multi v9.0.0. GRCh38 (build 2024-A, 10X Genomics) and Chromium Human Transcriptome Probe Set v1.1.0 references were used for probe read mapping.
Mouse immunofluorescence
Mouse experiments were approved by the Institutional Animal Care and Use Committee of the Weizmann Institute of Science (approval number 05220824–2) and performed in accordance with institutional guidelines. Two-month-old C57BL/6 male mice, housed under regular 12 h:12 h light:dark cycle and fed ad libitum were used in the experiments. Mice were euthanized by cervical dislocation and tissues were collected and fixed in 4% paraformaldehyde in PBS in 4 °C. Livers were incubated for 24 h, transferred to 70% Ethanol, embedded in paraffin, and stored at room temperature before staining.
Computational methods
Genome alignment
Human samples were aligned to genome assembly hg38. Domesticated pig samples were aligned to genome assembly Sscrofa11.1. Cattle samples were aligned to genome assembly BTauARS-UCD1.2. Boar samples were aligned to genome assembly Sscrofa11.1.
Spatial transcriptomics preprocessing
To remove outlier spots, for each patient, spots with log10 of total UMI counts lower than 3 s.d. from the mean log10 of total UMI counts across all spots, or mitochondrial fraction over 5 s.d. from mitochondrial fraction mean across all spots, were filtered out. Portal spots with prominent fibrosis, identified histologically based on the H&E staining, were manually marked and excluded from the analyses (Fig. 2a, patients M8, P7, P17, P2 and P21). In addition, spots located in areas where the tissue anatomy integrity was unclear were manually annotated and filtered out. The remaining spots were used for analysis. To correct for high abundant transcript diffusion into neighbouring spots51, we calculated for each gene the mean UMI count in spots within the fiducial frame that were not under the tissue to compute a background vector of expression. Next, the background vector was subtracted from each spot under the tissue, and negative values were set to zero. Finally, for the remaining spots, the raw data was normalized to relative counts per spot by dividing by the sum of all genes.
Pseudo-bulk analyses of human samples
Mitochondrial (^MT-) and ribosomal (^RPL, ^RPS) genes were removed from the variable gene list to minimize batch effect contributions to the analysis. Background subtracted counts were normalized, and a mean expression level was computed for each gene in every patient, creating a pseudo-bulk value for each gene. A z-score was calculated for each gene across patients, and hierarchical clustering (Fig. 1c) and PCA (Fig. 1d) were performed on the resulting z-scores for all genes with maximal normalized expression above 5 × 10−4. Spearman correlation distance and average linkage were used for hierarchical clustering. Comparison with NDD liver (Extended Data Fig. 1f) was performed on Visium data from Andrews et al.14. Pseudo-bulk was computed as the mean of spots annotated as hepatocytes and that further showed ALB expression above 0.003 of summed UMIs. GSEA29 was performed using version 3.0 with default parameters using the gene sets Hallmark and KEGG.
Human portal–central axis spot annotation
For Visium, we first extracted for each gene the spearman correlation coefficient between the gene’s expression level and CYP2E1 expression level over all spots. The 20 genes that were most correlated or anticorrelated with CYP2E1 over all spots were defined as our pericentral and periportal landmark gene sets (pericentral landmark gene set: CYP3A4, ADH1B, CYP1A2, CYP2E1, APOA2, APOC1, ADH4, ADH1A, APOH, AMBP, GSTA2, ADH1C, SLCO1B3, AOX1, APOA5, DCXR, RBP4, OAT, CYP2C19, GC; periportal landmark gene set: SERPINA1, APOA1, ALB, C7, NNMT, HAMP, ALDOB, ASS1, CYP2A7, MGP, A2M, FXYD2, CCL21, HAL, IGFBP2, SDS, AQP1, CYP2A6, FBLN1, PTGDS). Those landmark gene sets’ normalized summed expression levels enabled assigning a percentile-categorized zonation score to each spot, computed as the sum of periportal expression divided by the sum of periportal expression plus the sum of pericentral expression (\(\frac{\mathrm{sum}\,\mathrm{of}\,\mathrm{periportal}\,\mathrm{expression}}{\mathrm{sum}\,\mathrm{of}\,\mathrm{periportal}\,\mathrm{expression}\,+\,\mathrm{sum}\,\mathrm{of}\,\mathrm{pericentral}\,\mathrm{expression}}\)). This zonation score was then binned into eight zones with equal percentiles, and a zone index was assigned to each spot. To correct for spurious annotations, we applied a median filter by re-assigning the zone of each spot as the median zone of all spots surrounding it (relative distance of under 150 µm; Fig. 2a). Subsequently, the zonation bias P value of each gene was computed using the Kruskal–Wallis test, followed by Benjamini–Hochberg correction for multiple hypotheses. Centre of mass was calculated for every orthologous gene per sample as the weighted average of the zones26,28. Gene zonation scores were calculated as the linear combination of the zonal layer with the highest gene expression and the gene’s centre of mass (Fig. 2). Gene zonation scores were non-defined for genes that exhibited low expression in our Visium datasets. For the MERFISH analysis, we used NOTCH3 as a periportal landmark gene and reconstructed cell zonation using the distance from a NOTCH3 positive cell. For the Visium HD analysis, we worked on the 8-μm spot size data. Non-parenchymal spots, identified histologically based on the H&E staining, were manually marked and excluded from the analyses, as were spots at the section periphery and spot with fewer than 50 UMIs. Zonation score was initially assigned to each spot using the same landmark gene method used for the Visium slides and binned into eight zones based on equal percentiles over the analysed spots. Subsequently, zone values were assigned the median zone of 100 μm ×100 μm areas. Spots were binned and averaged, yielding the Visium HD-reconstructed zonation profiles.
Human hepatocyte marker gene identification
To identify genes that serve as markers for hepatocytes, we utilized the single-cell atlas generated by Massalha et al.17. We defined marker genes as those with a normalized expression level higher than 10−5 and with a ratio of expression in hepatocytes to their maximum expression in all other cell types greater than 0.5. Since hepatocytes contain more than tenfold total mRNA than non-parenchymal liver cells, this value ensures that the majority of transcripts for such genes in each spot are likely to originate from hepatocytes46. We identified 1,724 hepatocyte marker genes (Figs. 2b,c and 3h).
Cross-species zonation reconstruction
To remove outlier spots, for each non-human sample spot with log10 of total UMI counts that are either higher or lower than 1.5 standard deviations from the mean log10 of total UMI counts across all spots were filtered out. For the non-human livers, each spot’s zone was determined based on landmark gene expression in cow and mouse26 and based on distances from the portal fibrotic septa in pigs and boars; Spots were then binned into 6 distinct zones, by equally proportioned percentiles and the assigned zone was median filtered using spots closer than 150 µm. Zonation profiles per species were computed as the weighted means of the zonation profiles of individual samples, weighted by the median number of UMIs per spot. When comparing zonation trends of specific gene subsets (Fig. 3h), the ratios between the expression levels in the most periportal and pericentral zones were computed. Only hepatocyte-specific genes that were highly expressed in at least one non-human species and in humans (maximum expression above 10−5 in each) were analysed.
Cross-species orthologous genes analysis
To allow for a common ground analysis, we sub-setted our data to genes that show 1:1 orthologue across all species (1 orthologue per gene per species). In total, 12,410 human, mouse, pig, boar and cattle orthologous genes were found using the Ensembl biomart tool55 (Supplementary Table 7) 1,249 hepatocyte-specific genes with orthologues across all species genes were found (Fig. 3h). Only human patients with steatosis level below 10% were used (n = 7, M2, M3, M4, M5, M6, M7 and M8). For the analysis of WNT-activated and WNT-repressed genes we utilized the data from a published WNT-hyperactivating liver-specific Apc-knockout model mice43 and used the 20 genes that were most WNT-upregulated and the 20 genes that were most WNT-downregulated.
When comparing the zonal trends of NPCs (Fig. 4) we considered the subset of genes that were expressed in the respective cell type at a level above 5 × 10−5 and threefold higher levels than the maximal expression in any other cell type. For human liver we used the pseudo-bulk data of Massalha et al.17, for mouse we used the data of Ben-Moshe et al.62.
Liver cell atlas using Visium HD
Visium-HD was performed on liver tissue sections from two donors, M2 and M6. Single-nucleus segmentation was performed on the H&E image using the Stardist algorithm63 implemented within QuPath, utilizing the pre-trained H&E model he_heavy_augment. Segmented nuclei were expanded by up to 6 µm or until contacting neighbouring nuclei. Each segmented object was classified as either hepatocyte or NPC using a random forest object classifier trained on morphological features. The 2 µm × 2 µm Visium HD bins were next assigned to the segmented cells if their centroids resided within the cell area. Each cell received classification based on object classifier and the gene expression values that equal the sum of UMIs of the enclosed bins.
For downstream single-cell analysis, cells with fewer than 200 UMIs were excluded. To generate a single-cell uniform manifold approximation and projection (UMAP), the data from both patients were combined. Gene expression was log-normalized, and highly variable genes (HVGs) were identified using thresholds of min_mean = 0.095 and min_disp = 0.5. z-scored HVG expression values were concatenated with a weighted classification binary output vector from the object classifier, which encoded the hepatocyte versus NPC label. PCA was applied, 25 components were selected and batch correction was performed using the Harmony algorithm (Harmonypy 0.0.10). The Harmony-corrected PCs were used to build the neighbourhood graph and UMAP visualization, using Scanpy (version 1.10.0) standard functions (n_neighbors = 20, resolution = 1.7).
The Visium HD single-cell atlas was used to extract landmark genes for each major cell type: hepatocytes, LSECs, Kupffer cells and fibroblasts. To this end, the periportal and pericentral landmark genes from the Visium experiments were first used to assign a zonation score, which was then divided into three equal percentile bins. Next, the zonation profiles of each cell type were computed as the mean expression of all cells in the specific bin. The centres of mass were computed for each gene and sorted, and the top and bottom 20 genes with the most extreme centres of masses were selected as the cell-type-specific portal or central landmark gene set respectively. This calculation was performed over all genes for hepatocytes, and only for genes with maximal zonal expression above 10−4 and twofold higher expression in the respective cell type compared to the maximal expression in all other liver cell types, based on the human liver cell atlas of Massalha et al.17.
Liver cell atlas using snRNA-seq
snRNA-seq data were generated from liver tissues of four individuals (M5, M6, M7 and M8) and combined into a single AnnData object. Cells with mitochondrial transcript fractions exceeding 40% were excluded. To reduce ambient RNA background, we performed background subtraction by estimating gene-wise mean expression in low-complexity cells (150–400 total UMIs) and subtracting this background from all cells, followed by transforming negative values to zero. All downstream analysis was performed on log-transformed expression data. To retain high-quality cells, we selected those with total UMI counts between 500 and 50,000 and with up to 7,000 detected genes. HVGs were identified using thresholds of min_mean = 0.095, max_mean = 6, and min_disp = 0.5. To refine the annotation, we removed likely contaminant non-hepatocyte cells with ALB expression greater than 0.01. A second round of HVG selection, PCA, Harmony batch correction, UMAP and Leiden clustering was then performed using 50 principal components, 15 neighbours, a resolution of 1.0 and a UMAP minimum distance of 0.5. Only clusters with more than five cells in each of the four patients were retained. Final cell-type annotations were curated based on canonical marker gene expression.
The lobule layer of the single nuclei was inferred based on the cell-type-specific landmark genes that were extracted from the Visium HD single-cell atlas. For each of the four main cell types, the periportal and pericentral landmark gene sets’ normalized summed expression levels enabled assigning a percentile-categorized zonation score to each cell, computed as the sum of periportal expression divided by the sum of periportal expression plus the sum of pericentral expression (\(\frac{\mathrm{sum}\,\mathrm{of}\,\mathrm{periportal}\,\mathrm{expression}}{\mathrm{sum}\,\mathrm{of}\,\mathrm{periportal}\,\mathrm{expression}\,+\,\mathrm{sum}\,\mathrm{of}\,\mathrm{pericentral}\,\mathrm{expression}}\)). This zonation score was binned into eight zones with equal percentiles, and a zone index was assigned to each cell. The zonation profiles were computed as the means across all cells belonging to the same zone. To examine the zonation of liver cell types (Fig. 4c–g and Extended Data Fig. 9), cell-type-specific markers were extracted as genes with mean expression higher than 10−4 and 20-fold higher expression in the respective snRNA-seq atlas cluster compared to the maximal expression in the other clusters, with the exception of hepatocytes, for which twofold higher expression threshold was used.
Ligand–receptor and transcription factor analysis
The database of Ramilowsky et al.30 was used as reference for ligands and matching receptors. Spearman correlations were computed between the zonation profiles of all ligands and matching receptors in each of the 16 pairs of cell types in the snRNA-seq dataset (Extended Data Fig. 9, Supplementary Table 9). Transcription factors are based on ref. 35.
Lipid droplet quantification
Analysis of gene expression changes in early steatosis was performed on four out of the seven donors who exhibited steatosis, and for which image quality was sufficient to reliably extract lipid content values (M1, M2, M3 and P6). To extract the lipid percentage of each spot in the H&E attained Visium slide, an artificial neural network pixel classifier was trained on the H&E images using the QuPath software v.4.064. The pixel classifier was trained to identify three histologic features: fat droplet, non-fatty liver tissue and the slide background. For each slide, the pixel classifier was trained on multiple fatty droplets and non-fatty tissue from the same H&E image (Fig. 5a), with the following parameters: resolution: ‘very high’, channels: ‘Hematoxylin and Eosin’, scales: ‘1,2,4’, features: all 12 available features (the default is Gaussian). The spot locations were uploaded from the Visium output, and the QuPath measurement tools was used to quantify the number of fat droplet pixels and non-fatty liver tissue pixels in each spot (Fig. 5c).
Comparison to mouse model of steato-hepatitis
Visium spots in the mouse data from Zhou et al.38 were assigned to one of 8 lobule layers following the procedure described in ‘Human portal–central axis spot annotation’, using the mouse landmark genes from Droin et al.27. For each mouse, pericentral expression was defined as the mean expression in the two most pericentral layers. The log2 of the ratio between the mean pericentral expression in the steatotic and non-steatotic mice was computed for each gene, and correlated with the log2 of the respective ratio between the pericentral spot expression in each of the steatotic human liver samples and the mean of the pericentral spot expression in all non-steatotic human samples.
Ethics statement
All work was conducted in compliance with the principles outlined in the Declaration of Helsinki. Human experiments were approved by the Sheba Medical Center Helsinki committee (Helsinki approval number 866521SMC) and by the Mayo Center Institutional Review Board number 22-008260. Informed consent was obtained from all participants. Mouse experiments were approved by the Institutional Animal Care and Use Committee of the Weizmann Institute of Science (approval number 05220824–2).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.