Destination vector cloning
We built a specific custom destination vector for efficient Golden Gate cloning of enhancer fragments. This included the reporter gene cassette NanoLuc-Ires-mNGreen-pA, downstream of a mCMV promoter. A bacterial suicide ccdB cassette spanning the enhancer position enabled assembly of a single enhancer for selection and efficient cloning. This was based on the EMMA Golden Gate cloning system43. For the GSC long-term differentiation experiments, the same vector was built but included PiggyBac transposase recognition sites flanking the entire cassette.
Bioinformatics to design a SOX2 enhancer oligonucleotide pool
We re-analysed previously published1 SOX2 ChIP–seq and H3K27ac GSC cell line data to identify GSC-specific SOX2 peaks that were overlapping with H3K27ac and absent in differentiated cells (cells in serum culture). The resulting shared peaks were then combined into one set and manually curated to remove centromeres. This resulted in 1,721 peaks with an average length of 402 bp. These peaks were then used to design a set of 160 bp sequences. Next, 20 bp adapters were included to flank the sequences and these sequences were synthesized as an oligonucleotide pool (Twist Bio).
Construction of an arrayed plasmid library of enhancer fragments
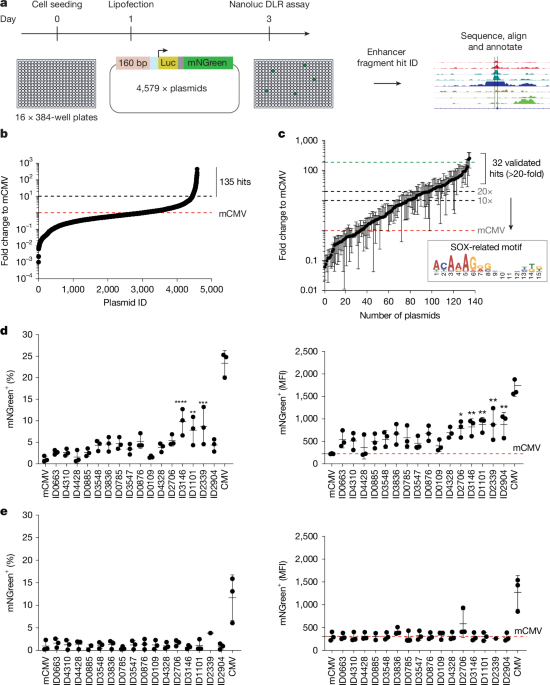
The oligonucleotide pool (9,523 unique sequences) was first amplified for 10 cycles with 0.25 µl (2.5 ng input) volume, and 0.5 µl of this reaction was used for the subsequent 15 cycles of amplification to reduce PCR ‘jackpot’ amplification. The final products were cloned into the expression vector using an efficient Golden Gate reaction. We used KAPA HiFi Hotstart polymerase with GC buffer (Roche, KK2501), 68 °C annealing temperature and 5 s of extension time at 72 °C. A total of 4,579 individual plasmids were then randomly picked, miniprepped and plated as an arrayed plasmid DNA library on 96-well plates. This was deemed a practical number for arrayed library screening. A magnetic-bead-based SPRI purification method was used to minimize loss and to clean up the DNA before Golden Gate cloning. Sanger sequencing for quality checks of a sample of the plasmids confirmed that they were diverse, and all sequences could be mapped back to the original library design.
Screening platform for 384-well plates
Cells were seeded in 384-well plates using a multidrop and transfected the following day using CyBio Felix (CyBio). Two days later, a NanoGlo DLR assay was used to identify hits with a fold change to mCMV of >10. These hits were Sanger sequenced to determine the specific sequence and mapped back to the genome to validate overlap with the original SOX2 ChIP–seq data. The GREAT tool predicted the target gene for hit sequences44.
General cell culture procedures
The GSC lines GCGR-E17 (E17), GCGR-E21 (E21), GCGR-E27 (E27), GCGR-E28 (E28), GCGR-E31 (E31), GCGR-E34 (E34), GCGR-E37 (E37) and GCGR-E55 (E55) and the human NSC lines NS9FB_B (NS9), NS12ST_A (NS12) and NS17ST_A (NS17) were generated in the Pollard Laboratory and are available upon request from the Glioma Cellular Genetics Resource. Informed consent was obtained for use of patient tissue. All procedures on patient brain tissue received ethics approval from the NHS Health Research Authority, East of Scotland Research Ethics Service (REC reference 15/ES/0094), and all procedures on embryonic and fetal brain tissue received ethics approval from the Lothian NHS Board, South East Scotland Research Ethics Committee (REC reference 08/S1101/1).
Cells were grown as adherent monolayers under serum-free conditions as previously described13. All cells lines tested negative for mycoplasma at passage 3 using a MycoAlert Mycoplasma Detection kit (Lonza, LT07-318). Cell line authentication by STR profiling was performed as a service by the European Collection of Authenticated Cell Cultures (ECACC). Reports are available upon request from the Glioma Cellular Genetics Resource. Analysis was conducted using the Promega Fusion system (DC2402), analysing differences at 24 distinct hypervariable genetic loci, 16 of which are used for the final STR profile and report. The ioGlutamatergic (io1001S) and ioGABAergic (ioEA1003S) neurons and ioMicroglia (ioA021) were obtained from bit.bio and grown per the manufacturer’s specifications. GSC7 was previously characterized22.
For lipofection, Plus reagent and Lipofectamine LTX (Life Technologies, 15338030) were each diluted in half the volume of Opti-MEM I reduced-serum medium (hereafter referred to as Optimem) (Life Technologies, 31985062). Following this step, the Plus reagent and Optimem premix was added to all DNA samples followed by the Lipofectamine LTX and Optimem premix. The transfection mix was incubated for 5 min at room temperature and then carefully dropped onto the cells. Generally, no change in medium was carried out. Cells were analysed 2 days after transfection.
HEK293 cells were seeded at the specified density in the respective plate format. The next day GMEM medium, DNA and polyethylenimine were mixed and incubated for 15 min at room temperature to facilitate formation of the complex. The transfection mix was subsequently dropped onto the cells, and analysis was carried out 2 days later.
Screening assay using the Nano-Glo Dual-Luciferase reporter assay system
This assay consists of two steps. Transfection was performed using a normalization plasmid PGK-Firefly-Luciferase (Promega, E5011) transfected in a 1:10 ratio with the plasmid of interest. Therefore, in this assay, the firefly luciferase activity is first measured, which enables normalizing of the transfection efficiency. Cells were washed 3 times with PBS and 20 µl was left in a 96-well plate (25 µl in a 384-well plate). Oneglo buffer was added, and the plate was shaken for 5 min at 480 rpm to allow cell lysis. Next, 20 µl cell lysate in a 96-well format (25 µl cell lysate in a 384-well format) was transferred into an opaque white 96-well format (or 384-well format) and light was measured for 0.1 s per well (Ensight Multimode Plate Reader, Perkin Elmer). For the subsequent NanoLuc reaction in a 96-well format, 2 µl cell lysate was transferred into an opaque white plate containing 40 µl PBS, and 20 µl Stopglo buffer, supplemented with substrate, was added to each well. In a 384-well format, 20 µl Stopglo buffer containing substrate was added on top of the undiluted cell lysate. The plate was shaken again for 5 min at 480 rpm to quench the firefly luciferase reaction and to ensure good mixing. NanoLuc activity was measured for 0.1 s per well. Data obtained by the NanoLuc reaction were normalized to the Firefly reaction to account for well-to-well variability of transfection efficiency. Normalized data were used to calculate the fold change compared with the empty vector control, which contained mCMV only and no enhancer fragments.
Immunocytochemistry
Cells were washed twice with PBS and fixed in 4% paraformaldehyde (PFA) for 10 min. Cells were permeabilized with 0.1% Triton-100 in PBS (PBST). Cells were blocked with blocking solution (1% BSA in PBST with 3% goat serum) and incubated with the primary antibody at 4 °C overnight. The next day, cells were washed 3 times in PBST, and the respective secondary antibody was applied in blocking solution and incubated for 45–60 min at room temperature. Cells were washed with PBS and incubated for 5 min with a DAPI nuclear counterstain at 1 µg ml–1 final concentration. Images were acquired using a Nikon TiE microscope and NIS elements software (Nikon).
Flow cytometry
For flow cytometry, cells were detached, pelleted and resuspended in an appropriate volume of flow cytometry buffer (1% BSA in PBS, v/v) or PBS. Cells were stained with Draq7 (Abcam, ab109202, final concentration of 0.1 µM) or DAPI (ThermoFisher Scientific, D3571) as a live/dead stain and analysed using a BD LSRFortessa cell analyser (4 lasers, BD Bioscience). Analysis of flow cytometry data was carried out using FlowJo Analysis software (v.10.6.2) or FCS Express 7 flow cytometry software (v.7.22.x).
RNA isolation, cDNA synthesis and RT–qPCR
RNA was extracted using a MasterPure RNA Purification kit (Epicentre). RNA was stored at −80 °C, and concentration was determined using a NanoDrop spectrophotometer. cDNA synthesis was carried out using a SuperScript III Reverse Transcriptase kit (Life Technologies) according to the manufacturer’s instructions. Around 200–500 ng RNA was used for the cDNA reaction. The same amount of RNA was used for each experiment. After the reaction, cDNA was diluted to the required volume using nuclease-free water.
For RT–qPCR, TaqMan Universal PCR master mix (Applied Biosystems) and TaqMan gene expression assays (Life Technologies) were used on a Quant Studio7 Flex Real-Time qPCR machine. RNA samples that did not undergo reverse transcription (to assess DNA contamination) and water controls were used on every plate. RT–qPCR was carried out in technical duplicates. Data analysis was performed using the ddCt method, which assumes 100% PCR efficiency and is guaranteed with TaqMan assays. In brief, the mean was calculated for technical replicates and normalized to the housekeeping gene GAPDH, which produces the dCt value.
Western immunoblotting
Cell lysates were prepared by resuspending cells in RIPA buffer (50 mM HEPES pH 7.7, 150 mM NaCl, 1% NP-40, 0.5% DOC and 0.1% SDS), incubating on ice for 5 min, centrifuging at maximum speed for 10 min in a tabletop centrifuge (5415D, Eppendorf) and collecting the supernatant (lysate). Protein extracts were quantified using a Pierce BCA Protein Assay kit (Thermo Scientific, 23225). SDS–PAGE was performed using homemade 4–12% Bis-Tris gels. Gels were transferred onto PVDF membranes (Millipore, IPVH00010), previously activated in methanol, by wet electroblotting or semi-dry blotting using a Bio-Rad Trans-blot turbo system. Western blots were revealed using HRP-conjugated antibodies, homemade ECL solutions or Clarity Western ECL (Bio-Rad), and imaged using X-ray films or a Bio-Rad ChemiDoc Imaging system.
Precipitation of interacting proteins
PCR amplification with biotinylated primers and purification were used to generate templates that could pull down bound proteins. Enhancer fragments were amplified with 5′ biotinylated primers using PrimeStar Max (Takara) according to the manufacturer’s instructions.The following primers were used: forward primer 595: TGATCCGTCTCGCCCTACTAGGTTACTGGTGCATGC; reverse primer 596: ACTAACGTCTCGGAGCACCCAAACTATTGGAGCGAG (bold sequences, adaptors).
PCR purification using Agencourt AMPure XP magnetic beads (Beckman Coulter) was carried out according to the manufacturer’s instructions. PCR product quantification using Tapestation and reagents was carried out according to the manufacturer’s instructions.
All the buffers were prepared the day before, passed through a 22 µm filter (except the dialysis buffer) and left at 4 °C overnight. DTT and protease inhibitors were added immediately before use. Cell pellets were resuspended in 5 ml per 40 million cells of ice-cold buffer A (10 mM HEPES pH 7.9, 1.5 mM MgCl2, 10 mM KCl, 0.5 mM DTT and protease inhibitors (complete, Roche, 11697498001)) and incubated on ice for 10 min. The cell suspension was transferred to a glass Dounce homogenizer and dounced 40 times on ice, transferred to a Falcon tube and centrifuged at 1,350 rcf for 10 min at 4 °C. The supernatant was discarded (cytosolic extract) and the pellet was resuspended in 100 µl per 10 million cells in ice-cold buffer B (20 mM HEPES pH 7.9, 5% glycerol, 1 M NaCl, 1.5 mM MgCl2, 0.2 mM EDTA pH 8.0, 0.5 mM DTT and protease inhibitors (complete, Roche, 11697498001)). The suspension was rotated for 30 min at 4 °C and transferred into a dialysis membrane (SnakeSkin, Thermo Scientific, 68100). Dialysis was carried out in 500 ml dialysis buffer (20 mM HEPES pH 7.9, 5% glycerol, 100 mM KCl, 0.83 mM EDTA pH 8.0, 1.66 mM DTT and protease inhibitors (complete, Roche, 11697498001)) for 2 h at 4 °C with rotation. The dialysis buffer was changed (500 ml), and dialysis was continued overnight at 4 °C. The extract was collected in a 1.5 ml tube and spun at maximum speed for 15 min at 4 °C (centrifuge 5415D Eppendorf). The supernatant was collected into new 1.5 ml tubes and quantified (Pierce BCA Protein Assay kit, Thermo Scientific, 23225). Next, 100 µg aliquots were flash-frozen in liquid nitrogen and stored at −80 °C. The following primary antibodies were used for western blotting: anti-SOX2 (rabbit) 1:1,000 Abcam (ab92494); anti-SOX9 (rabbit) 1:500 Millipore (ab5535); and anti-GAPDH (mouse) 1:10,000 Ambion (AM4300). The secondary antibodies used for western blotting were anti-rabbit 1:5,000 Novex (A16110) and anti-mouse 1:5,000 Novex (A16027).
Streptavidin magnetic beads (NEB, S1420S) were used according to the manufacturer’s instructions. In brief, 10 µl beads were aliquoted into low-bind tubes and washed 3 times with binding buffer (20 mM Tris-HCl pH 7.5, 0.5 M NaCl and 1 mM EDTA) on a magnetic stand. Next, 20 µl biotinylated DNA (around 20 pmol) was mixed with 200 µl binding buffer and was added to the beads. The suspension was rotated at room temperature for 2 h. The beads were washed 3 times with binding buffer, and 50 µg nuclear extract was added (total volume of 50 µl). Beads, DNA and nuclear extract, to which 150 µl dialysis buffer (20 mM HEPES pH 7.9, 5% glycerol, 100 mM KCl, 0.83 mM EDTA pH 8.0, 1.66 mM DTT and protease inhibitors (complete, Roche, 11697498001)) was added, were rotated overnight at 4 °C. The following morning, beads were washed 3 times with wash buffer (20 mM HEPES pH 7.9, 5% glycerol, 250 mM NaCl, 0.83 mM EDTA pH 8.0 and 1.66 mM DTT). For western blotting, proteins were eluted in 20 µl loading buffer (lithium dodecyl sulfate buffer containing 50 mM DTT) and boiled for 5 min to denature the proteins. For mass spectrometry, the beads were sent dry to the Institute for Genetics and Cancer core facility (University of Edinburgh) for analyses.
SOX2 and SOX9 ChIP–seq library preparation and analysis
In brief, GSCs cultured in 150 mm dishes to 70% confluence were crosslinked in 1% formaldehyde for 10 min at room temperature. Excess formaldehyde was quenched by incubating cells with 0.25 M glycine for 5 min at room temperature. The cell pellet was washed twice with ice-cold PBS before storage at −80 °C. For each GSC line, cell pellets from eight 150 mm dishes were combined before proceeding for cell lysis and sonication. The cell pellet was lysed in buffer LB3 (10 mM Tris-HCl pH 8.0, 100 mM NaCl, 1 mM EDTA pH 8.0, 0.5 mM EGTA pH 8.0, 0.1% sodium deoxycholate, 0.5% N-lauroylsarcosine and 0.1% SDS) and chromatin was fragmented to 200–600 bp size on a Covaris M220 sonicator (total of 4 cycles, with the following treatment settings: 600 s per cycle, peak power 75, duty factor 10, cycles/burst 200; temperature: minimum 5 °C, set point 7 °C, maximum 9 °C). For each ChIP assay, 15–20 µg fragmented chromatin was incubated overnight in a cold room with 10 μg antibodies (R&D SOX2 AF2018, Millipore, SOX9 AB5535) and 30 μl Protein G Dynabeads (10003D). The next day, magnetic beads were washed 5 times with RIPA (50 mM HEPES-KOH pH 7.5, 0.5 M LiCl, 1 mM EDTA pH 8.0, 1% NP-40 and 0.7% sodium deoxycholate), once with TE NaCl (10 mM Tris-HCl pH 8.0, 1 mM EDTA pH 8.0 and 50 mM NaCl) and finally eluted in 200 μl ChIP elution buffer (50 mM Tris-HCl pH 8.0, 10 mM EDTA pH 8.0 and 1% SDS). Immunoprecipitated chromatin was reverse crosslinked by incubating samples at 65 °C for 10 h and extracted using the phenol–chloroform–IAA (11896714) method. To account for intertumoral heterogeneity, ChIP–seq was performed on seven independent primary GSC lines. Libraries were prepared for two technical ChIP replicates and one input control using a NEBNext Ultra II DNA Library Prep kit for Illumina (E7645S). The libraries were barcoded using NEBNext Multiplex Oligos for Illumina (Dual Index Primers Set 1 kit, E7600S) and sequenced on an Illumina Novaseq (50 bp read length).
Paired-end reads were aligned to the hg38 genome using BWA, filtering out poor-quality (MAPQ < 10), duplicates, mitochondrial genome and blacklisted regions (https://www.encodeproject.org/annotations/ENCSR636HFF/). Technical replicates were merged, and peaks were called using MACS2 with default settings. Consensus SOX2 and SOX9 peak sets were derived by taking the overlap of peaks occurring in 5 out of 7 GSCs for each TF set. The overlap significance between the consensus SOX2 and SOX9 peak sets was determined using a circular permutation test in regioneR (ntimes = 10,000). Genomic regions near SOX2 and SOX9 peaks were annotated using HOMER45.
To perform overlapping analysis with GSC super-enhancers, we downloaded raw fastq files from publicly available H3K27ac datasets on GSCs (GSE119834, GSE74529, GSE121601 and GSE92458) and called super-enhancers using ROSE with default settings. A consensus set of GSC super-enhancers was derived by considering super-enhancers that occurred in at least two GSCs. The overlap significance between the consensus co-bound SOX2–SOX9 sites and consensus GSC super-enhancers was performed using a circular permutation test as described above. Gene ontology term association analysis was performed using GREAT. To find centrally enriched de novo motifs at SOX2 and SOX9 peaks in co-bound SOX2–SOX9 enhancers, and to identify the spacing between the most significant de novo motifs from each peak set, we used CentriMo and SpaMo from the MEME-ChIP suite of tools46.
scRNA-seq sample preparation
On the day of cell seeding, cells were detached as described above. Cells were counted using a haemocytometer and seeded into a 6-well Corning plate at 400,000 cells per well in 2 ml. Plates were incubated at 37 °C with 5% CO2. The next day, AAV vector stocks were thawed at room temperature. The appropriate amount of virus stock was added to the culture medium to achieve a final multiplicity of infection (MOI) of 1,000,000, which ensured high transduction efficiency. Viral-containing medium was added to cells without replacing the existing medium. The cells were then returned to the incubator at 37 °C with 5% CO2.
After 3 days of incubation with the viral particles, cells were detached as described above. Next, 10% of the cells were prepared for flow cytometry using DAPI as a live/dead stain (see Supplementary Information for the gating strategy). The remaining 90% of the cells were processed for single-cell transcriptome sequencing.
Cells were fixed using a Parse Evercode v.2 Cell Fixation kit following the manufacturer’s instructions, with an average of around 400,000 cells per sample. Fresh reagents were prepared for each fixation, and 7.5% Gibco BSA fraction V was added per the manufacturer’s recommendations. In brief, cells were washed with PBS and resuspended in cold Cell Prefixation buffer. Cells were filtered through a 40 µm strainer after 5 min of centrifugation at 300g at 4 °C. Following this step, Cell Fixation solution and Cell Permeabilization solution were subsequently added, with incubation on ice for 10 min and 3 min, respectively. Cell Neutralization buffer was then added, and cells were centrifuged again at 300g for 5 min at 4 °C. Finally, cells were resuspended in a volume of 50–90 µl Cell Buffer containing 1% DMSO based on the live cell number before fixation. Cells were filtered through a 40 µm strainer and stored at −80 °C for up to 3 months.
scRNA-seq library preparation
Libraries were generated using a Parse Evercode WT Mega v.2 kit according to the manufacturer’s instructions. In brief, 72 samples were loaded into a 96-well RT barcoding plate for reverse transcription, with each well containing a unique barcoded RT primer. Following cDNA synthesis, cells were pooled and subjected to two additional rounds of barcoding, achieving three rounds of barcoding. Barcoded cDNA was then amplified via template switching and pooled into 15 sublibraries, each containing around 62,500 cells. A fourth barcode was incorporated during PCR amplification of the sequencing libraries. Sublibraries were then fragmented, ligated with Illumina adapters and purified using Ampure XP beads. Library quality was assessed on an Agilent BioAnalyzer 2100 before sequencing. Sublibraries, with 5% PhiX spike-in, were sequenced on an Illumina NovaSeq X using 300 cycles kits as paired-end, dual-index reads.
scRNA-seq analysis
Sequencing reads were aligned to the human (hg38) genome, and UMI counting data were generated following the standard Parse Bioscience pipeline, ‘splitpipe (v.1.1.2)’, with default parameters. Three custom genes were added to the genome (mCherry, HSV-TK and bGHployA) to capture ‘activated’ SSE-7 cells. In total, 345,495 cells were included in the initial dataset. Low-quality cells were filtered out based on the following thresholds: (1) the percentage of mitochondrial gene was >20%; (2) the number of genes was <300; (3) the number of uniquely aligned reads was <500; or (4) the number of uniquely aligned reads was >20,000. Genes detected in fewer than five cells were filtered out as low-quality genes. Potential doublets were identified using the R package ‘DoubletFinder’ (v.2.0.4), with an expected doublets rate of 3%, as guided by Parse. Doublets and clusters containing more than 20% doublets were removed. Ambient RNA-contaminated cells (cell score > 0.2) were removed using ‘decontX’ from the R package ‘celda’ (v.1.18.2). After filtration, 270,842 cells remained for further analysis.
The data were normalized using ‘LogNormalize’, and principal component analysis was performed based on the top 2,000 variant genes using the R package ‘Seurat’ (v.5.0.3). The first 15 principal components were used as input for Louvain-based graphing. SSE-7-activated cells (2,711 cells) were labelled according to the following criteria: mCherry > 1 or HSV-TK > 1 or bGHpolyA > 0. Cells with a single count of mCherry or HSV-TK from the virus-treated group were labelled as false negative (9,583 cells), whereas the remaining cells were labelled as SSE-7 non-activated cells (125,525 cells). Meanwhile, cells expressing mCherry or HSV-TK in the control group were labelled as false positive (1,998 cells). Transcriptional subtypes were predicted on the basis of a previously described gene signature27 using the function ‘sigScores’ and ‘as_four_state_gbm’ from the R package ‘scalop’ (v.1.1.0). Cell cycle was predicted using ‘CellCycleScoring’ from the R package ‘Seurat’.
SCENIC analysis of scRNA-seq data
False-negative and false-positive cells were removed, and the dataset was divided into 15 libraries as per the sequencing library preparation. The dataset was then analysed following the standard ‘pyscenic (v.0.12.1)’ pipeline. Auxiliary human datasets (hg38 mc_v10_clust) were downloaded from the cisTarget resources website (https://resources.aertslab.org/cistarget/). Motif–TF annotation was based on 10 kbp upstream and downstream around the transcription start sit (20 kbp in total). SSE-7-enriched TF modules were selected on the basis of the following criteria: (1) Z score > 0.7 and (2) Z score in the SSE-7-activated group was higher than in other groups, as determined by repeated-measures ANOVA test. Enrichment for candidate TF modules across 15 libraries was as follows: SOX8 (found in 9 libraries), STAT1 (found in 15 libraries), IRF9 (found in 12 libraries), ETS1 (found in 13 libraries), MAF (found in 3 libraries), SMAD1 (found in 12 libraries), FOS (found in 11 libraries), HIF1A (found in 7 libraries), SOX9 (found in 10 libraries), STAT3 (found in 3 libraries), NEF2L2 (found in 13 libraries), JUN (found in 2 libraries) and SOX2 (found in 2 libraries).
Lentivirus production and titration
To produce SOX2, SOX9 and rtTA2M2 individual lentiviral supernatant samples, nearly 2.4 million HEK293T cells per 15 cm plate were cultured for 24 h in GMEM medium supplemented with 10% fetal calf serum (FCS), 1 mM sodium pyruvate, 1 mM glutamine and non-essential amino acids (HEK medium) at 37 °C and 5% CO2. At 24 h after seeding, HEK293T cells were transfected with a plasmid cocktail containing 7.5 μg expression plasmid, 5.1 μg psPAX2 packaging vector, 2.4 μg envelope vector mixed in 45 μl Fugene6 (Roche) and 855 μl Opti-MEM medium (Invitrogen, Thermo Fisher Scientific). The cells were further cultured for 16 h at 37 °C and 5% CO2 before the medium was changed for HEK medium. At 65 h after transfection, virus-containing supernatant was collected and cleared by centrifugation and filtered through a 0.45 μm syringe filter (Merck Millipore). Virus was concentrated (roughly 100-fold) by transferring to thin-walled ultracentrifuge tubes and pelleted by ultracentrifugation at 25,000 rpm for 2.5 h using a SW32-TI rotor in a Beckman Optima XPN ultracentrifuge (Beckman Coulter). The pelleted virus was resuspended in 300 μl plain GMEM medium and incubated at 4 °C for 16 h before aliquoting and flash-freezing in liquid nitrogen for storage at −80 °C. The viral titre, calculated by flow cytometry, was 6 × 107 infectious units per ml.
Overexpression of SOX2 and SOX9
Early passage (passage 4) human fibroblasts (hFibs) were seeded, 1.5 million per 15 cm plate, and cultured in hFib medium (GMEM medium supplemented with 10% FCS, 1 mM sodium pyruvate, 1 mM glutamine, MEM 1× non-essential amino acids solution, 50 μM 2-mercaptoethanol (Gibco, Thermo Fisher Scientific) and 2 ml penicillin–streptomycin (Invitrogen, 15140122)) 24 h before transduction at 37 °C and 5% CO2. The cells were transduced by exchanging the growth culture medium with lentiviral medium, without antibiotics, containing 8 μg ml–1 polybrene, and 65 μl concentrated rtTA2M2 and either SOX2 or SOX9 virus, for single factor overexpression, or 75 μl rtTA2M2, SOX2 and SOX9 viruses for double factor overexpression. At 48 h after transduction, the cells were split 1:2 and cultured for 24 h, after which the culture medium was replaced with induction medium containing 1 μg ml–1 doxycycline (induction day 0). After 48 h of induction, the cells were collected for lysis and nuclear protein extraction.
Nuclear extraction of GSC7 and hFib cells
GSC7 cells were cultured as adherent monolayers13. For differentiation, cells were cultured without the supplements EGF and FGF and in 5% serum for 2 weeks. GSC7 and hFib nuclear lysates were prepared in the same way; cells were resuspended to 1 million cells per 100 μl 10 mM HEPES pH 7–8, 1.5 mM MgCl2, 10 mM KCl, 0.5 mM DTT, supplemented with complete Ultra EDTA-free protease inhibitors (Roche, 05892970001) and homogenized using a dounce and tight-fit pestle with 1 stroke per 1 million cells. Samples were centrifuged at 1,350 rcf for 5 min at 4 °C. The nucleus pellet was resuspended to 10 μl per 1 million starting cells in 20 mM HEPES pH 7–8, 30% glycerol, 420 mM NaCl, 1.5 mM MgCl2, 0.2 mM EDTA, 0.5 mM DTT and complete Ultra EDTA-free protease inhibitors (Roche, 05892970001). Samples were dialysed for 4 h into a minimum of 1–1,000 volume excess of 20 mM HEPES pH 7–8, 30% glycerol, 100 mM NaCl, 0.83 mM EDTA, 1.66 mM DTT, 0.2 mM PMSF and complete Ultra EDTA-free protease inhibitors (Roche 05892953001) using Slide-A-lyzer mini dialysis units with 3,500 MWCO cassettes (Thermo). Samples were divided into single-use aliquots (11 μl) and flash-frozen in liquid nitrogen for storage at −80 °C.
Western blot analysis of overexpression
The protein concentrations of the lysates were quantified using a Pierce BCA Protein Assay kit according to the manufacturer’s 96-well format instructions (Thermo Fisher Scientific). The following overexpression (OE) values were used: SOX2 OE = 10.54 mg ml–1, SOX9 OE = 5.56 mg ml–1, SOX2 and SOX9 OE = 7.56 mg ml–1, GSC7 GSCs = 1.02 mg ml–1 and GSC7 differentiated cells = 0.43 mg ml–1. Verification of SOX protein overexpression, relative to untransfected hFib lysate, was resolved by SDS–PAGE and electroblotting onto a PVDF membrane for western blotting. The primary antibody incubations with goat anti-human SOX2 antibody (1:363, AF2018, Abcam) and rabbit anti-mouse SOX9 (1:2,000 AB5535, Chemicon) were performed at 4 °C for 16 h. The secondary antibody incubations with donkey anti-goat IgG-HRP (1:2,000 dilution of ab97110, Abcam) and goat anti-rabbit (1:2,000 of 32460, Invitrogen) were performed for 1 h at room temperature. Blots were visualized using SuperSignal West Pico Chemiluminescent substrate (Thermo Fisher Scientific) and HRP was visualized using a Bio-Rad ChemiDoc MP imaging system.
Recombinant SOX protein production
His-tagged human SOX2 and SOX9 were expressed from a pET28a backbone (cloned by G. Roberts in the Soufi Laboratory) using Rosetta 2(DE3) pLys competent cells (Novagen).
SOX2 was expressed as previously described47. Cells in LB medium (1% Bacto-tryptone, 0.5% yeast extract and 1% NaCl), supplemented with 30 µg ml–1 kanamycin and chloramphenicol, were grown at 37 °C overnight. SOX9 was expressed in 2× TY medium (1.6% Bacto-tryptone, 1% yeast extract and 0.5% NaCl), supplemented with 30 µg ml–1 kanamycin and chloramphenicol, at 16 °C overnight. After expression, bacteria cell pellets were lysed, 100 ml buffer per 1 l of culture, in GuHCl Denaturing buffer (6 M guanidine-HCl, 50 mM Tris-HCl (pH 8) and 500 mM NaCl, 5% (v/v) glycerol) overnight. Lysate was sonicated for 200–300 s at 25–30 microns using a Soniprep 150 with intermittent chilling on ice. The supernatant was clarified by centrifugation at 18,000 rpm in a FLA21-8x50y rotor for 30 min. The lysate was further sonicated for 120 s, as above, then filtered through a 0.45 µm filter before loading into a HisTrapHP column (Cytivia) for affinity purification of His-tagged proteins using an AKTA pure system. The column was equilibrated with 5 CV of denaturing 10 mM imidazole buffer (6 M urea, 500 mM NaCl, 50 mM Tris-HCl (pH 8), 10 mM imidazole and 5% (v/v) glycerol), after sample loading was washed with 20 CV of denaturing 50 mM imidazole buffer (6 M urea, 500 mM NaCl, 50 mM Tris-HCl (pH 8), 30 mM imidazole and 10% (v/v) glycerol), and bound proteins were eluted with denaturing 300 mM imidazole (6 M urea, 500 mM NaCl, 50 mM Tris-HCl (pH 8), 300 mM imidazole and 10% glycerol). Peak fractions of the elution were analysed by SDS–PAGE, and samples with appropriately sized bands (40 kDa for His–SOX2, and 60 kDa for His–SOX9) were combined and desalted into 2 M urea buffer using a HiPrep 26/10 desalting column (GE Healthcare). The SOX2 desalting buffer comprised 50 mM Tris-HCl (pH 8), 240 mM NaCl, 10 mM KCl, 2 mM MgCl2, 2 mM CaCl2, 0.8 M urea, 30% (v/v) glycerol, 0.1% NP40 substitute, 0.05% Triton-X-100, 2 mM EDTA and 5 mM DTT resuspended in 1× PBS. The SOX9 desalting buffer comprised 50 mM HEPES NaOH (pH 7.5), 240 mM NaCl, 10 mM KCl, 2 mM MgCl2, 2 mM CaCl2, 2 M urea, 30% (v/v) glycerol, 0.1% NP40 substitute, 0.05% Triton-X-100, 2 mM EDTA and 5 mM DTT. The fractions corresponding to a dip in the absorbance at A280 were collected and visualized by SDS–PAGE before combining. Protein concentration was determined by SDS–PAGE, comparing to rAlbumin standards. The gels were imaged by detecting Coomassie blue staining using a Bio-Rad ChemiDoc MP imaging system. The resulting images were visualized, and the bands quantified using ImageJ 2.8/1.54i.
Small-molecule inhibitor screening
E55 cells were screened using inhibitors included in the Kinase Screening Library (Cayman, 10505) as well as two YAP–TEAD pathway inhibitors, MGH-CP1 (HY-139330) and TED-347 (HY-125269). Around 18,000 cells were seeded per well in a 96-well plate and transduced with the corresponding AAVs at a MOI of 2,000,000 on the next day. Cells were incubated at 37 °C, 5% CO2. In the primary screen, the cells were treated with a single concentration of 3 μM per inhibitor at 24 h after transduction. After 48 h of inhibitor treatment at 37 °C and 5% CO2, the cells were detached and analysed using an Agilent NovoCyte Penteon five laser flow cytometer and FCS Express 7 Research (v.7.24.0030). The results were expressed as the fold change in mCherry median fluorescence intensity in the inhibitor-treated samples relative to the DMSO control. After performing the first round of screening, inhibitors that gave more than 20% changes were selected for the second round of screening.
In the second-round screen, the cells were seeded and transduced with AAVs using the same procedures as the primary screen. At 24 h after transduction, the cells were treated with 3 different doses (0.1, 1 and 3 μM), and incubated at 37 °C and 5% CO2 for 48 h. Cells were then detached and analysed using a Agilent NovoCyte Penteon five laser flow cytometer. Draq7 was used as the live/dead stain in both rounds of screening. The results of the second round screen were expressed as the fold change in mCherry MFI and as the percentages of mCherry-positive cells, each normalized to the DMSO control.
EMSAs
Cy5-labelled SSE-7 bait for EMSAs was generated by PCR amplification of SSE-7 from the plasmid using Cy5-labelled primers (Sigma): 5′-ACTAGGTTACTGGTGCATGCTTGTCCTGCCTTTGAGAACA-3′ and 5′-ACCCAAACTATTGGAGCGAGAGAAAGGAAAGAAAGAGGTC-3′. The following primers were used for Cy5-labelled fragments for EMSA: ID4310: 5′-ACTAGGTTACTGGTGCATGCTTGTCCTGCCTTTGAGAACA-3′ and 5′-ACCCAAACTATTGGAGCGAGCTCATTTGAAGCAGAAGAAT-3′; ID4428: 5′-ACTAGGTTACTGGTGCATGCTCTAAATGGAGATCCTCCCA-3′ and 5′-ACCCAAACTATTGGAGCGAGACAGGAGGAAGTAGTAAATC-3′; ID0785: 5′-ACTAGGTTACTGGTGCATGCAAGAATGAACTGGGCCCAGC-3′ and 5′-ACCCAAACTATTGGAGCGAGTGAAAACCAGAGGCATCTCA-3′; and ID3836: 5′-ACTAGGTTACTGGTGCATGCTACCCTTCCAGGGGAGCAGT-3′ and 5′-ACCCAAACTATTGGAGCGAGAGAAAGGAAAGAAAGAGGTC-3′.
The binding to Cy5-end-labelled SSE-7 was analysed in native 1% agarose gels (12 × 13 × 1 cm), which were prepared in 0.5× TBE (45 mM Tris-borate and 1 mM EDTA). Gels were stored overnight at 4 °C before pre-running at 120 V (approximately 10 V cm–1) for 1 h. For affinity analysis, a 10 μl mixture typically containing 50 nM Cy5-labelled full-length or fragmented SSE-7, 100 ng µl–1 poly(dA:dT) (InvivoGen, tlrl-patn) and 0, 0.5, 1, 2 or 4 μl of hFib lysates, with and without SOX overexpression (5.56 mg ml for OE lysates and 3.47 mg ml–1 for hFib control lysates), GSC7 GBM stem, or differentiated, cell lysates (at 0.43 mg ml–1), or purified recombinant SOX protein (at 3.17 µM). For SOX2 and SOX9 in combination, equimolar amounts of SOX2 and SOX9 were mixed for 0.5, 1, 2, 4 and 8 µl of protein in total. Next, 8 µl SOX protein was added to 25 nM SSE-7 DNA in a 20 µl volume, and 20 µl of sample was loaded for gels, which were prepared in 1× binding buffer (10 mM Tris HCl pH 7.5, 1 mM MgCl2, 10 μM ZnCl2, 10 mM KCl, 1 mM DTT, 5% (v/v) glycerol and 0.5 mg ml–1 BSA). The mixtures were incubated at 20 ± 1 °C in the dark for 1 h using DNA LoBind tubes (Eppendorf). The entirety of each sample was then loaded onto agarose gels and electrophoresis was performed at 120 V for 3 h at 4 °C. The gels were imaged by detecting Cy5 fluorescence using a Bio-Rad ChemiDoc MP imaging system. The resulting images were visualized, and the bands were quantified using ImageJ 2.8/1.54i.
Adult human cortex and brain tumour slice cultures
Tumour and non-tumour brain tissue were transferred to a 10-cm2 tissue culture dish with sterile PBS and placed on ice. Visibly damaged tissue was removed and the remaining tissue was transferred into a 35-mm2 dish with pre-warmed 3% SeaPlaque agarose (50100, Lonza). After cooling in ice, the block was removed and cut using a scalpel into an approximately 2 cm cube around the brain. Before starting to cut, a 6-well plate was prepared. In each well, we introduced one cell culture insert (PICMORG50, Millicell) and added below it 1 ml culture medium in basal NSC medium, Dulbecco’s modified Eagle medium: F12 supplemented with EGF and FGF (Life Technologies). The embedded brain was placed in a circular vibratome plate with glue. The vibratome (VT1000 S, Leica) plate was fixed in the platform and filled with PBS and penicillin–streptomycin (15140-122, Gibco, 1:100). Then, 300-µm-thick slices were cut, with vibrating frequency set at 10 and speed at 1. Each slice was transferred using a small brush onto the top of a Millipore culture insert. The platform was maintained cool at all times. The 6-well plate was placed in an incubator at 37 °C with 5% CO2.
Addition of virus
A 5 µl volume of virus was pipetted onto the centre of each tissue slice, using a 10 µl tip without touching the tissue. Three repeat doses of 5 µl virus was added to each slice at 5 min intervals (20 µl in total per slice). Slices were incubated for 7 days. The medium was replaced on day 3 or 4.
Immunostaining
The medium was removed and exchanged for 1 ml freshly prepared 4% PFA; 1–2 ml PFA was also placed gently on top to cover the slice. After 2 h, PFA was removed, brain slices were washed 3 times with PBS and transferred using a brush to a 24-well plate. Slices were incubated at room temperature for 1.5 h in blocking solution (0.5% Triton X-100 and 3% goat serum; Sigma-Aldrich) followed by incubation with primary antibody for 2 days at 4 °C. Sections were washed five times with PBS, the respective secondary antibody was applied in blocking solution overnight. The next day, slices were washed 5 times with PBS mounted and cleared in RapiClear 1.49 (RC149001, Sunjin Lab).
Images were acquired using an Opera Phenix Plus high-content imaging system (Revvity) equipped with a ×40/1.1 NA water-immersion objective. Image analysis was performed in Harmony software (v.5.2; Revvity). For the full pipeline see Supplementary Information. In brief, nuclei were segmented using DAPI with the common thresholding method, and the nuclear mean intensity of channels SOX2 (488) and mCherry (555) were then calculated. Thresholds to define positive cells were set for each tissue slice individually based on 3–5 regions of interest. Nestin was segmented using ‘find image region’ with channel 647, and positive cells were defined as those with nuclei in these regions. Analysis was performed for a single z plane. Data were exported and analysed using GraphPad Prism. Representative images of the segmentation can be found in the Supplementary Information.
Zebrafish experiments
All embryos were obtained by natural spawning and collected in conditioned aquarium water in 0.00001% methylene blue. Embryos were raised at 28.5 °C in embryo medium (E3) on a 14 h light–10 h dark photoperiod and were treated with 200 μM N-phenylthiourea (Sigma) from 6 h post fertilization to inhibit pigmentation. Zygotes were injected at the one-cell stage of development. DNA constructs were created using the Tol2Kit system48,49. Approximately 2 nl of plasmid DNA (30 ng μl–1) containing Tol2-capped mRNA (20 ng μl–1), supplemented with 0.2% w/v phenol red (Sigma) to facilitate visualization of injected volume, was injected. To induce Akt1 overexpression in neural progenitor cells, a combination of Tol2-pDEST-NBT:DlexPR-lexOP-pA (20 ng μl–1) and Tol2-pDEST-lexOP:AKT1-lexOP:tagRFP (30 ng μl–1) plasmids was injected as previously described50. The newly generated tgSSE-7:eGFP was outcrossed to tg(Xla.Tubb:DsRed) and tg(olig1:mScarlet). Live imaging was performed using either a Leica MS205 stereomicroscope or a Leica SP8 confocal microscope with a ×40/NA 1.1 objective. Animal experimentation was approved by the ethics review committee of the University of Edinburgh and the Home Office in accordance with the Scientific Procedure Act 1986.
AAV medium-exchange transduction assay
HEK cells were seeded into a 6-well plate a day before transfection, aiming for 60–70% confluence the next day. On the day of transfection, the culture medium was exchanged and HEK cells were transfected with RepCap, pHelper and AAV-ITR transgene plasmids using polyethylenimine. Two days later, conditioned HEK culture medium (now containing rAAV particles) was collected and centrifuged at 1,300 rpm for 4 min to remove any cells. rAAV-conditioned medium was transferred to HEK or GSC7 cells seeded at low density (10–20%) in 6-well or 12-well plates. Two to three days later, cells were analysed for transduction. For AAV1 in vivo experiments, we purchased research-grade viral vectors from Vector Biolabs (iodixanol gradient ultracentrifugation purified).
AAV transduction assay in iPS cell-derived neurons
GSC7 and E55 cells were seeded in a 24-well plate format the day before transduction, whereas ioGlutamatergic neurons, ioGABAergic neurons and microglia (bit.bio) were seeded in a 48-well format before differentiation. After differentiation, cells were transduced with AAVs at a MOI of 5 × 105. Cells were incubated at 37 °C and 5% CO2 for 3 or 10 days.
Cell seeding and transduction for GCV killing assay
On the day of cell seeding, cells were detached using the method described above. Cells were counted using a haemocytometer and seeded into 96-well (1,000 cells per well in 50 µl), 24-well (30,000 cells per well in 500 µl) or 6-well (60,000 cells per well in 2 ml) Corning plate and placed in an incubator at 37 °C and 5% CO2. The next day, AAV stocks were thawed at room temperature, and an appropriate amount of virus stock was added to the required amount of culture medium to achieve the final MOI of 5 × 105. Culture medium containing viral particles was added to cells without the replacement of existing medium. Cells were returned to a 37 °C and 5% CO2 incubator.
Prodrug treatment
Lyophilized GCV was diluted in DMSO to achieve 100 mM stock concentration. GCV stocks were aliquoted and were stored at −20 °C for no longer than 1 month. To make a working stock concentration, GCV was diluted 1:100 in appropriate culture medium, and 20 µl was added to wells already containing 80 µl culture medium for a final concentration of 200 µM. As a negative control, DMSO was diluted 1:100 in appropriate culture medium to achieve a working stock. Next, 20 µl of working stock was added to wells already containing 80 µl culture medium. As a positive control, 20 µl DMSO was added to wells with 80 µl culture medium to achieve a final concentration of 20% DMSO.
Incucyte live-cell imaging
To track cell proliferation and morphological changes during treatments, cells were monitored using an Incucyte live-cell imaging system. Whole-well imaging of Corning 96-well plates was performed every 4 h. Basic confluence scoring analysis software (Incucyte) was used to estimate confluence. Images at specific time points were extracted to verify cell confluence and morphology.
MTT assay
On the day of assay, the culture medium was replaced with 0.3 mg ml–1 MTT solution (diluted in cell line-appropriate culture medium). Cells were placed in incubator at 37 °C and 5% CO2 for 3 h. After incubation, the medium was removed and 70 µl DMSO was added to each well. Each plate was kept in the dark at 37 °C for 20 min, shaking it occasionally. Before reading the plate, each well was visually inspected to make sure that all (formazan) crystals were dissolved. Plates were read with plate reader at 560 nm absorbance.
Statistical analysis
Data analysis was performed using Microsoft Excel (v.16.23 for Mac), GraphPad Prism (v7) and RStudio (v1.1.456). Error bars are shown as the s.d. of the mean. For illustrations, BioRender and Adobe Illustrator 22.0.1 were used.
Furthermore, we used open-source programmes such as bedtools, GREAT, fastasplitter, HOMER and MEME for various analyses45,46.
Tumour initiation by transplantation and intratumoral AAV delivery
All animal procedures were approved by the University of Edinburgh Animal Welfare and Ethical Review Body (AWERB) and conducted under UK Home Office licence (PPL number: PP8631583) in accordance with the Animals (Scientific Procedures) Act 1986 and ARRIVE guidelines. Male and female C57BL/6 mice (6–8 weeks old) were obtained from Charles River. Mice were housed in Individually ventilated cages with sterile bedding plus enrichment inside a pathogen-free facility on a 12-h light–dark cycle, at a temperature range of 20–24 °C and with relative humidity level of 45–65%. Transplantation experiments were performed as previously described25. We used 7-week-old male mice. Mice were transplanted with 200,000 NPE-IE cells in 2 µl PBS per mouse. NPE-IE cells have been previously described25 and have Nf1 and Pten inactivating mutations, alongside EGFvIII overexpression and a GFP-Luc reporter construct. At 2 weeks, when tumours were visible by IVIS imaging (bioluminescence) and of similar size, AAV1-SSE-7–HSV-TK–IL-12 virus (1.2 × 1013 viral genomes per ml, Vector Biolabs) was delivered directly into the mouse brain tumour (2.5 µl per mouse) using a microsyringe linked to an injection pump, at a rate of 0.17 µl min–1 and 2.3 mm depth, higher than the previous depth of 2.4 mm for tumour cells. Each mouse received 3.00 × 1010 viral genomes. The following day after virus injections, GCV was intraperitoneally injected (500 µl per mouse at 2 mg ml–1) once daily for 20 days. Mice were imaged by IVIS imaging (PerkinElmer) once a week for luciferase signals. Animals were observed regularly for any neurological symptoms. Mice with tumours were culled based on the following criteria; (1) body weight loss of ≥20% relative to baseline (weights of mice were recorded once weekly); (2) abnormal neurological signs, including ataxia, head tilt, circling behaviour, seizures, paresis or inability to right itself within 3 s; (3) hunched posture with reduced locomotor activity and lack of reach response; and (4) any extracranial tumours ≥ 15 mm in diameter or ulcerated as per the Workman (2010) guidelines51. All mice reaching the humane end point were culled using the schedule 1 method. The end-point limits were not exceeded in any of the experiments.
Immune characterization of mouse GBMs
Brains were collected and processed 5 days after treatment. First, they were microdissected under a Leica stereo microscope to avoid contamination with healthy tissue. Tissues were mechanically disaggregated using scissors, followed by 30 min of enzymatic digestion using a mix of 0.35 µg ml–1 Liberase TL (05401119001, Roche) and 0.23 µg ml–1 DNAse (101041590001, Sigma) in plain RPMI. After digestion, tissues were dispersed through a 70 nm cell strainer to obtain a single-cell suspension. After centrifugation, cells were resuspended in PBS for further flow cytometry staining. Cell suspensions were incubated for 10 min at 4 °C with a blocking mixture of mouse, rat and calf serum containing anti-CD16/32 blocking antibodies (clone 2.4G2, BioXcell, BE0307). Cells were washed with PBS and incubated for 20 min in antibody-containing staining buffer plus Fixable Viability Dye eFluor-780 (eBioscience, 65-0865-14) to distinguish live and dead cells. Cells were then washed and resuspended in fixation/permeabilization buffer (BD CytoFix/Cytoperm; 554714 or FOXP3 Transcription Factor Staining Buffer set, eBioscience) followed by intracellular target staining.
SABER–FISH detection of transgene copy number
SABER–FISH was performed as previously described52,53. Non-overlapping 36 bp probes were designed to target the HSV-TK transgene, and probes were extended using primer exchange reaction to approximately 500 bp. For imaging, cells were first stained with mCherry,then fixed with PFA and subsequently processed for immuno-SABER–FISH54.
Assessment of the fraction of the tumour cell population that activates AAV1-SSE-7–mCherry
AAV1-SSE-7-mCMV–HSV-TK-V5–mCherry was injected at around 3.58 × 1014 viral genomes per ml and 7.2 × 1013 viral genomes per ml (1:5 dilution) in PBS. AAV was injected into tumours as described above. After 7 days, tumours were imaged using a stereomicroscope and surgically excised.
Each tumour sample was added to 1 ml digest mix (RPMI + 1 mg ml–1 collagenase IV, 12.6 µg ml–1 DNase I and 1% penicillin–streptomycin) and incubated for 1 h in a shaking incubator at 40 rpm at 37 °C. Samples were filtered through a 100 μm cell strainer, washed with 9 ml FACS buffer and centrifuged at 250g for 15 min at room temperature. The pellet was then resuspended in 1 ml 37.5% Percoll in PBS and gradient centrifuged at 900g, room temperature, for 12 min with no brake. The supernatant was discarded, and the bottom layer of cells resuspend in 500 µl PBS and filtered into a FACS tube. Cells were stained with DAPI as a live/dead stain and analysed using a BD LSR Fortessa cell analyser (four lasers, BD Bioscience).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.