In utero electroporation
All animal procedures in this study were performed with approval from the St. Jude Institutional Animal Care and Use Committee. IUE was performed as described previously27, and plasmids were prepared with a NucleoBond Xtra Maxi Plus EF kit (Takara Biosciences). After anaesthesia with 4.5% isoflurane, pregnant CD1 mice at E16.5 were subjected to abdominal incision to expose the uterus. A DNA plasmid cocktail (1 μg μl−1 pBCAG-HA-ZRFUS1, 1 μg μl−1 pbCAG-eGFP-Luciferase, 1.5 μg μl−1 pX330-sgTp53, 2 μg μl−1 GLAST-PBase, 1.5 μg μl−1 mPlagl1 single-guide RNA, 0.5 μg μl−1 TrackerSeq library, FastGreen dye) was injected into the lateral ventricles with a glass pipette. Electric pulses were then delivered to the embryos by gently clasping their heads with forceps-shaped electrodes. Six 33-V pulses of 55 ms were applied at 100-ms intervals. The uterus was then repositioned into the abdominal cavity, the abdominal wall was sutured and the skin was stapled. Following birth, pups were monitored for clinical signs of tumour growth (such as seizures, circling, and head doming), as well being monitored by magnetic resonance imaging every 2 weeks. At end point, mice were collected for isolation of nuclei or immunofluorescence staining. In accordance with our St. Jude Institutional Animal Care and Use Committee protocol, end point was defined on the basis of a set of neurological symptoms (gait, hunching, kyphosis, squinting), and these limits were not exceeded in any of the experiments. Mice for isolation of nuclei were perfused with 10 ml cold PBS, and tumours were frozen in isopentane and stored at −80 °C. For survival curves, pups from a minimum of 2 mothers were included. Randomization and blinding were not applicable.
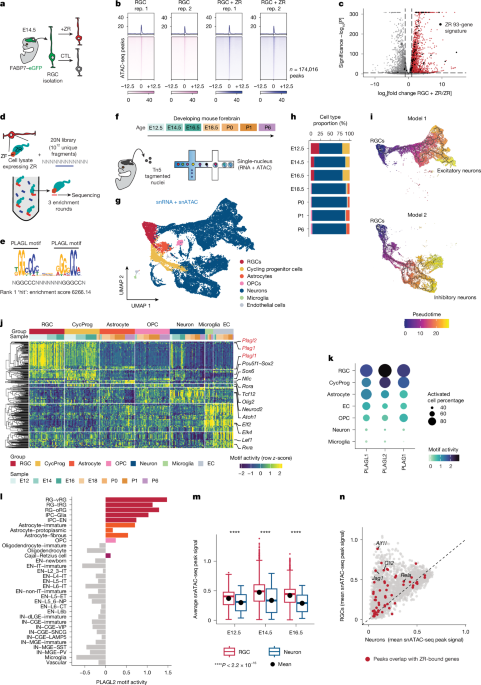
RGC isolation
RGCs were previously isolated from Ink4a-knockout mice with GFP expressed from the Blbp promoter using a Worthington Papain Dissociation system (LK003150). Cells were grown in neural basal medium (Invitrogen) supplemented with sodium pyruvate, glutamine, B27, N2, bFGF (10 ng ml−1) and rhEGF (20 ng ml−1) and tested for mycoplasma monthly. Cells were grown on treated cell culture dishes coated with Matrigel (Corning). RGCs were made ZR positive using a lentivirus generated by the Viral Vector Core at St. Jude. Mouse tumour cells were seeded into 10-cm plates 24 h before infection and infected with lentivirus with 8 μg ml−1 polybrene for 24 h. Infected cells were selected with 2 μg ml−1 puromycin for 3 days. ZR expression was confirmed by western blotting. Single-guide RNAs for Plagl1 were generated by the Center for Advanced Genome Engineering with an Addgene 52961 backbone, which included RFP. ZR-positive RGCs were infected by the same method and sorted for RFP using a BD FACSAria Fusion system. Knockout of more than 90% was confirmed by targeted deep sequencing.
OPC isolation
Primary OPC cultures were performed as previously described56,57. In brief, cortical tissues from E14.5 mouse embryos were collected, and neural stem cells were cultured as neurospheres for 4 days. Neural stem cells were dissociated and plated on poly-d-lysine-coated dishes at a density of 1.5 × 104 cells cm−2 in OPC media. They were subsequently infected with either ZR-Lenti-Cherry or control virus for 14 h. After viral infection, cells were maintained in OPC medium for 4 days before being collected for ATAC-seq and RNA-seq.
RNA-seq and ATAC-seq analyses
RNA-seq and ATAC-seq analyses were performed using Genialis Expression software (https://www.genialis.com) deployed locally on St Jude HPC infrastructure. Briefly, the RNA-seq pipeline run on the Genialis platform comprised the following steps. Raw reads were filtered to remove adaptors and poor-quality reads using BBDuk (v.37.9; https://sourceforge.net/projects/bbmap/). The resulting reads were mapped to the reference genomes (Ensembl 92) using STAR (v.2.7.0; RRID SCR_015899). FeatureCounts (v.1.6.3; RRID SCR_012919) was used for quantification of gene expression levels, followed by DEseq2 (RRID SCR_000154) for differential gene expression analysis. Genes with low expression (expression count summed over all samples of less than 10) were filtered out from the input matrix to DESeq2. The paired-end reads from ATAC-seq were trimmed using BBDuk (v.37.9) and mapped to reference genome mm10 using Bowtie2 (v.2.3.4.1). MACS2 (v.2.1.1.20160309) was then used to call peaks on the aligned reads using a P value cutoff of 0.01 (parameters –shift −75 –extsize 150 –nomodel –call-summits –nolambda –keep-dup all −P = 0.01).
TrackerSeq library generation and validation
TrackerSeq library cloning was carried out generally as described in ref. 19. In brief, the pCAG-SacB plasmid was digested with BstXI, and the 8-bit barcode was cloned into it using NEBuilder HiFi master mix (six reactions in total), followed by isopropanol purification. Purified reactions were electroporated into Endura DUOs (Lucigen) using a MicroPulser with program Ec1 (Bio-Rad). Four electroporations were carried out then recovered for 1 h at 37 °C in 2 ml of recovery media. Next, cells were plated overnight at 32 °C on 245-mm plates (Corning). The following morning, plates were scraped and cells were collected in Luria Broth (Miller), and library plasmids were purified using Endofree midiprep kits (Qiagen). For validation, 10 ng of the library plasmid prep was amplified using 2xPhusion (NEB) and sequenced by the Hartwell Center for Genome Sequencing Facility at St. Jude.
CSI DNA binding assay
HEK293T cells were transiently transfected with HA-tagged ZR fusion plasmids using Lipofectamine 2000 reagent according to the protocol (Thermo Fisher 11668019). Following expression, cells were lysed with RIPA buffer (Thermo Fisher 89900) and spun down for collection of supernatant. A DNA library (Integrated DNA Technologies) containing randomized central regions of 20 bp flanked by constant sequences complimentary to primers was converted to double-stranded DNA and brought to 74 ng μl−1 before being combined with 1% w/v bovine serum albumin, 500 ng μl−1 poly dI-DC (Thermo Fisher 20148E), 1% NP-40 (Thermo Fisher 85124), and 10× PBS. ZR-positive and ZR-negative cell lysates were incubated with this mixture for 1 h at room temperature. The mixture was then added to anti-HA beads (Thermo Fisher 88836) washed in binding buffer (10× PBS, 1% bovine serum albumin, 1% NP-40) and incubated for 30 min. Solutions were washed in binding buffer and aspirated on a magnetic plate three times before being resuspended in a PCR master mix (Lucigen Econo Taq 2×30035-1 and custom primers). Library fragments attached to beads were amplified on a Bio-Rad thermal cycler and then purified using a New England Biolabs Monarch PCR & DNA clean-up kit (T1030L). Eluted DNA library fragments from each sample were diluted to a concentration of 74 ng μl−1 and checked on a gel before being incubated with cell lysates again. Following three rounds of incubation and amplification, all purified library fragments from each round for each sample were given a unique barcode and a sequencing adaptor and then sequenced with a NovaSeq short-read sequencing amplicon kit, yielding approximately 500 million reads. Sequencing results were sorted by barcode, and the 20-bp library regions selected in each sample were ranked by enrichment and normalized to fusion negative lysate samples. Primer sequences were as follows: forward 5′-CTGATCCTACCATCCGTGCT-3′, reverse 5′-CCGCTCGGTACGAAGCTG-3′.
Nuclei isolation
Tissue (10–30 mg) was cut from human tumours and input into a 10x Nuclei Isolation Kit (PN-1000494). Kit instructions were followed, but the lysis buffer incubation time was increased to 15 min to isolate quality nuclei. Mouse embryonic forebrain was isolated with the 10x Demonstrated Protocol CG000366 RevD, but lysis buffer incubation was decreased to 2 min. IUE tumours were isolated with a Dounce homogenizer and iodixanol gradient using a modified version of the protocol described in ref. 58. Nuclei were resuspended in nuclei buffer and counted with a haemocytometer, and 10,000 were loaded on to a 10x Chromium Chip. Libraries were assessed with an Agilent TapeStation and sent for sequencing at St. Jude’s Hartwell Center on a NovaSeq 6000. Gene expression libraries had 400 million reads with a 28-10-10-90 cycle configuration, and ATAC libraries had 500 million reads with a 50-8-24-49 cycle configuration.
RNAscope
End-point ZR tumour-bearing mice were cervically dislocated, and brains were removed from the skull. Brains were flash-frozen in isopentane and frozen in OCT blocks, which were kept at −80 °C. Sections were cut on a Leica 3050 cryostat at 16 μm, and slides were kept at −80 °C until ready for use. The ACDBio Multiplex Fluorescent v.2 fresh frozen RNAscope protocol (revision B UM 323100) was followed with slight modifications: protease III was used for 30 min, opal fluorophores were used at 1:750, and DAPI was used at 1:5,000 for 20 min. The probes used were a custom C1 Zfta probe targeting [TGGCGTTTGGAGTATCTCATGGATTTCAACCCAGCGAGGCACGGCATGGTGTGCATGGTCTGCGGTAGCTCTTTGGCTACCCTGAAGCTGAGCACTATCAAACGCCACATCCGTCAGAAGCACCCGTACAGCCTGCAT], Cdk1 476081-C2, Pax6 412821-C3, Eomes 429641-C3, Grin1 431611-C3, Pdgfra 480661-C3, Mog 492981-C3 and Kif6 1760891-C3. Images were taken on a Zeiss LSM 780 at ×20 using a 9-square tile scan and maximum intensity projection of a z-stack.
Immunofluorescence
IUE surgery was performed with PBCAG-GFP at 4 μg μl−1. Brains were taken from P2 pups and placed in 4% paraformaldehyde at 4 °C with shaking for 24 h. They were then transferred to 30% sucrose until they sank (approximately 48 h). They were frozen in OCT and kept at −80 °C until ready for cutting. Brains were cut at 16 μm on a Leica 3050 cryostat, and slides were kept at −20 °C until ready for use. Slides were then permeabilized for 20 min (0.3% Triton in PBS), blocked for 1 h (5% normal donkey serum, 0.3% Triton in PBS), incubated with primary antibody overnight (GFP Aves, 1:2,000), incubated in secondary antibody for 1 h at room temperature (Thermo A78948), incubated in 1:5,000 DAPI (Thermo 62248) for 20 min, and mounted with Fluoromount G (Thermo 00-4958-02). For immunofluorescence of primary OPCs, cells were washed three times in PBS, permeabilized in PBST (0.3 % Triton/PBS) for 5 min, washed again, blocked with 10% normal goat serum in PBST for 1 h at room temperature, and incubated overnight at 4 °C with primary antibodies (Mouse OLIG2, Millipore MABN50, 1:500; Rabbit HA, sc-805, 1:1,000). After washing, cells were incubated with secondary antibodies for 1 h at room temperature, washed, stained with DAPI and mounted in VectaShield Antifade. All images were acquired on a Zeiss LSM 780 confocal microscope at ×20 using maximum intensity projections of z-stacks.
snMultiome data processing
For both human and mouse datasets, the ‘cellranger-arc count’ pipeline (10x Genomics, v.2.0.1, https://www.10xgenomics.com) was used for cell barcode detection, read alignment and quality assessment, following the standard 10x Genomics protocols. Human reads were aligned to the GRCh38 reference genome (refdata-cellranger-arc-GRCh38-2020-A-2.0.0, based on GENCODE v.32) and mouse reads to the mm10 reference genome (refdata-cellranger-arc-mm10-2020-A-2.0.0, based on GENCODE v.M23). The pipeline performed initial quality control by distinguishing intact nuclei from background and removing non-nucleus-associated reads. Subsequently, ambient RNA contamination per sample was assessed using SoupX (https://github.com/constantAmateur/SoupX) on the prefiltered data. Samples that passed both the initial quality control metrics reported in the Cell Ranger ARC QC summary and the SoupX contamination thresholds were retained for downstream analysis. We assessed data quality at the individual nucleus level and retained high-quality nuclei using Seurat (v.5.1.0, https://satijalab.org/seurat) and Signac (v.1.14.0, https://github.com/timoast/signac), applying the following criteria: total ATAC fragment count (nCount_ATAC) of at least 3,000, transcription start site enrichment scores between 2 and 15, total RNA counts (nCount_RNA) of at least 2,000, detected gene counts (nFeature_RNA) ranging from 500 to 8,000, and mitochondrial gene percentages less than 10%. For exclusion of doublets, doublet probabilities were estimated using DoubletFinder (v.2.0.4, https://github.com/chris-mcginnis-ucsf/DoubletFinder), and nuclei with high doublet scores were removed. DropletQC59 was applied to further identify and filter potential empty droplets, ensuring that only high-quality nuclei were retained for downstream analysis.
snMultiome data integration
For snATAC data of snMultiome analysis, open chromatin region peaks were called on individual samples using MACS2 (v.2.2.7, https://github.com/macs3-project/MACS) with the CallPeaks function in Signac (v.1.14.0), with intervals overlapping ENCODE blacklist regions excluded. To integrate all snATAC-seq data, we first created a unified set of peaks for quantification across datasets using the GenomicRanges package, filtering out peaks shorter than 20 bp or longer than 10,000 bp. Fragments for each sample were then recalled on the basis of this unified peak set. Top features in each snATAC-seq dataset were identified using FindTopFeatures (min.cutoff = 10), followed by TF-IDF normalization with the RunTFIDF function and latent semantic indexing (LSI) dimensionality reduction using the RunSVD function in Signac. After merging the datasets, we repeated these steps on the combined data. The snATAC data were then integrated using the FindIntegrationAnchors and IntegrateEmbeddings functions. Finally, nonlinear dimensionality reduction was performed using RunUMAP on LSI components 2 to 40. For snRNA data, normalization and data scaling were performed on the merged snRNA dataset using SCTransform v.2, followed by principal component analysis (PCA) and uniform manifold approximation and projection (UMAP) dimensionality reduction using the RunPCA and RunUMAP function in Seurat (v.5.1.0). Weighted nearest-neighbour analysis was done using Seurat with 1–30 principal components from snRNA data and 2–30 integrated LSI components from snATAC data. The resulting nearest-neighbour graph was then used for UMAP embedding and clustering following the best practice described in Seurat and Signac for multiome data analysis.
Estimation of motif activity from snATAC-seq data
Motif/TF chromatin accessibility (motif activity) was computed for a set of 841 TFs (combined mouse and human TFs) from the JASPAR 2022 database using the RunChromVAR function in Signac (v.1.14.0), and differential motif activity was computed with the FindMarkers function. Motif enrichment in the differentially accessible regions was analysed with the FindMotif function.
Cell type annotation for snMultiome and scRNA
Cell type annotation of single-cell data was performed using CellTypeEstimate (v.0.3.1; https://github.com/combiozone/CellTypeEstimate), an in-house tool adapted from the published tool ScType (https://github.com/IanevskiAleksandr/sc-type), incorporating custom code modifications and an alternative reference database. Marker sets were derived from more than 40 published studies of human and mouse brain development. References for each marker set are recorded in the ‘source’ column of the marker database files (db/*.xlsx). Following automated annotation, all labels were manually reviewed and refined to ensure biological accuracy. For tumour samples, the suffix ‘-like’ was manually appended to each annotated tumour cell type.
Processing of TrackerSeq barcode reads
We processed TrackerSeq barcode reads following the methods described in ref. 19 (https://github.com/mayer-lab/Bandler-et-al_lineage). Reads from the R2 FASTQ files were pre-processed by trimming the flanking sequences on both sides of the LBs. Barcodes shorter than 37 bp were discarded. A whitelist of cell barcodes was generated using UMI-tools. These whitelisted cell barcodes, along with their corresponding unique molecular identifiers, were appended to the read names in the LB FASTQ files to produce new, modified FASTQ files. The modified files were then used for downstream processing, including matching of LBs to cells and further filtering of the LBs. To isolate high-confidence LBs, we retained only those containing the conserved nucleotide motif ‘…CTG…ACT…GAC…TGA…CTG…ACT…GAC…’ and excluded any barcodes with ambiguous bases (N). These high-confidence LBs were matched to clean cells from the Seurat object, resulting in a set of cells with confidently assigned LBs. Cells were classified as either single LB, containing one unique LB per cell, or multiple LB, containing more than one LB per cell, on the basis of these assignments. In the single-LB group, LBs were ranked by number of associated cells, with those having the highest counts designated as dominant single LBs (typically referred to as LB-1). Only cells with a single LB were retained for downstream analyses.
TrackerSeq data analysis from quality control to cell type annotation
Raw TrackerSeq data were processed using Cell Ranger (v.9.0.0, https://www.10xgenomics.com) with mm10 (10x Genomics, refdata-cellranger-mm10-2020-A-2.0.0) as the reference genome. Then, filtered_feature_bc_matrix data were used to filter pooled cells and normalized in Seurat. We selected high-quality cells on the basis of cutoffs such as more than 500 genes, more than 2,000 unique molecular identifiers (nCount_RNA), less than 10% of mitochondrial genes, and less than the customized maximum quantile for nFeature_RNA (less than quantile(probs = 0.9) and no more than 8,000) and nCount_RNA (less than quantile(probs = 0.99)). Next, we used DoubletFinder (v.2.0.4) to estimate doublets, retaining only cells classified as ‘Singlet’, and removed empty droplets on the basis of DropletQC filtering. Afterwards, data were normalized using the SCTransform v.2 method in Seurat (v.5.1.0), regressing out the percentage of mitochondrial gene expression (percent.mt) to reduce its confounding effects. Last, high-quality cells were used to perform PCA, UMAP and clustering analyses in Seurat. Cell type annotation was then carried out using CellTypeEstimate (v.0.3.1) on the basis of the identified clusters. In downstream analyses, this information was integrated with LB-matched cells to enable more in-depth analysis. For the integration analysis, we merged all high-quality cells from independent TrackerSeq datasets using Seurat (v.5.1.0), performed SCTransform v.2 normalization (regressing out mitochondrial percentage) and PCA, applied Harmony (v.1.2.3; https://github.com/immunogenomics/harmony) with the IntegrateLayers function, and used the top 30 Harmony embeddings for UMAP visualization and clustering.
Classification of malignant cells
To classify cells as malignant or non-malignant, we inferred genome-wide CNVs using InferCNV (v.1.22.0, https://github.com/broadinstitute/infercnv) with default parameters. InferCNV was run at the sample level with integrated count matrices of snRNA-seq and snATAC-seq data, separately. The reference used immune cells and microglial cells. We first defined the CNVs as loss (InferCNV value ≤ 0.8) or gain (InferCNV value ≥ 1.2) and filtered CNV regions (hidden Markov model outputs) with fewer than 20 genes to detect large-region (≥5 Mb) chromosomal CNVs. Next, we calculated the copy number alteration ratio per CNV region. Last, we estimated malignant cells as those with large CNV regions (at least 100 CNV genes in the region) or a CNV ratio greater than 0.5 per sample.
ZR fusion signal signatures
The single-cell ZR fusion signature was calculated on the basis of 93 ZR driver genes18,20,45 using the AddModuleScore function in Seurat (v.5.1.0). We considered a score on our in-house benchmark test greater than 0.2 to indicate confidence, whereas scores of 0.1–0.2 indicated uncertainty, and scores less than 0.1 indicated distrust. Scores greater than 0.1 corresponded to the ZR target gene signal, whereas scores less than 0.1 indicated non-signal.
Cell cycling and non-cycling signatures
The CellCycleScoring function in Seurat (v.5.1.0) was used to score cells according to their cell cycle phase (G1, G2M or S). To define cycling and non-cycling cells, we recalculated the cell cycle score on the basis of cell cycle markers (cc.genes.updated.2019) using the AddModuleScore function in Seurat. The recalculated score was named cc.score. On the basis of internal testing, we defined cycling cells as those with cc.score, S.score or G2M.score greater than 0.2, and non-cycling cells as those with all scores less than 0.1.
Construction of pseudotime trajectories
Pseudotime trajectories were constructed to model cellular differentiation dynamics using two R packages, Monocle3 (v.1.3.7, https://cole-trapnell-lab.github.io/monocle3) and Slingshot (v.2.14.0). For Monocle3, the Seurat object was converted into a Monocle3 cell_data_set object. Root nodes were selected on the basis of known progenitor cell types identified in the UMAP, and cells were subsequently ordered along the trajectory for assignment of pseudotime values. For Slingshot, clustering results and dimensionality reduction coordinates from the Seurat object were used as input. Lineages were inferred on the basis of Seurat clusters or cell types, and pseudotime values were assigned to cells. The pseudotime from lineage 1 was used for downstream analyses.
Progenitor and lineage score of malignant cells
The progenitor and lineage score were calculated following the method in ref. 29, using only malignant cells identified by CNV (as described in the Methods). RGC-like or cycling progenitor-like cell types were designated as progenitors to serve as the starting point at the top. Progenitor score was estimated as the expression of the progenitor shared program minus the maximal expression of the two differentiation programs, and differentiated cells were further classified on the basis of average gene expression differences to differentiate between astrocyte- or ependymal-like and neuron-like lineages.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.