Cancer dependency map analysis
Genome-wide gene dependency scores were obtained from the Cancer Dependency Map (DepMap) 2024 Q2 CRISPRGeneEffect.csv public data release47.
Cell line generation, exogenous expression, siRNA and cell viability assays
Cell lines were grown in RPMI 1640 medium supplemented with 10% FCS with the exception of HEK293T cells, which were grown in Dulbecco’s modified Eagle’s medium supplemented with 10% FCS; CALU1 cells, which were grown in McCoy’s 5A, 10% FBS, 2 mM l-glutamine; and RERF-LC-AI cells, which were grown in MEM, 10% FBS, 2 mM l-glutamine. Cell line identity was verified by high-throughput single-nucleotide polymorphism (SNP) genotyping using Illumina Golden Gate multiplexed assays. SNP profiles were compared to SNP calls from internal and external databases to determine or confirm ancestry. All cell lines tested negative for mycoplasma contamination before storage/use at our institute.
siRNA KD experiments were performed using ON-TARGET-plus SMARTpools (siFBXO42 L-022191-00-0010, siCCDC6 L-010551-00-0010) or single SiGenome (siFBXO42 D-022191-02-0005) purchased from Horizon Discovery and were reverse transfected using Lipofectamine RNAiMax (Invitrogen) for 72 h according to the manufacturer’s instructions.
All constructs were generated through gene synthesis at Genscript with codon optimization for mammalian cell expression. Flag, HA or His tags were cloned at the N terminus of FBXO42 or CCDC6. FBXO42 deletion mutant amino acid boundaries were as follows: FBXO42(ΔF-box) (100–717), FBXO42(ΔKelch) (1–100), FBXO42(Δ1BC) (140–146), FBXO42(Δ)2BC (192–202) and FBXO42(ΔC-Ter) (2–295).
Generation of stable cell lines for expression of Flag–FBXO42 WT or mutants, sgRNAs, DOX-inducible CRISPR–cas9 or Flag–FBXO42 was carried out by the piggybac transposon system. Transfections were performed using Lipofectamine 3000 (Thermo Fisher Scientific) and cells selected for resistance to blasticidin (DOX-inducible expression) or puromycin (constitutive expression).
To assess clonogenic capacity, cells were seeded in 6- or 12-well plates at low confluency for 7 to 15 days. Cells were then fixed by 4% PFA and stained using 0.005% Crystal Violet.
Live-cell imaging was performed in a 96-well-plate format (×10 magnification) using the Incucyte Live-Cell Analysis System (Sartorius).
Cell lysates, IP, immunodepletion and WB
Whole-cell extracts were prepared in Pierce IP lysis buffer (87788) including protease inhibitor cocktail. Nuclear fractions were prepared using the Nuclear Extract Kit (Actif Motif 40010) according to the manufacturer’s instructions. For IPs, cells were pretreated with MG132 (10 µM, Cell Signaling, 2194S) or MLN4924 (5 µM, Cell Signaling, 85923S) overnight.
IPs of exogenous tagged-proteins were achieved using 1–5 mg of cell lysates incubated overnight at 4 °C in 1× IP High Buffer (Active Motif, 37510) with 50 µl of Magnetic Agarose beads corresponding to the respective bait tag (Thermo Fisher Scientific, Flag, A36797; HA, 88836).
IPs of recombinant proteins were done using 1 μg incubated for 4 h at 4 °C in 1× IP High Buffer (Active Motif, 37510) with 50 µl of magnetic agarose beads corresponding to baits tag. Beads were washed four times with the IP buffer and incubated overnight at 4 °C with 1–5 mg of cell lysates. The next day, IPs were washed four times in IP buffer, resuspended in a loading denaturing buffer and boiled for 15 min before WB analysis.
For immunodepletion experiments, 2 mg of lysates from HA-CCDC6 knock-in NCI-H1373 cells were incubated overnight in Active Motif High Buffer (37510) with either anti-HA or control IgG magnetic beads. The beads were then removed, replaced with fresh beads and incubation was continued for an additional 4 h. After bead removal, lysates were analysed by WB.
For routine WB, the samples were mixed in a denaturing buffer, boiled for 5 min and loaded to Novex Tris-Glycine protein gels at variable percentages. Gels were transferred onto an iBlot 2 Transfer Stacks nitrocellulose membrane by wet transfer (Thermo Fisher Scientific). Membranes were incubated with the following primary antibodies diluted in 5% whole milk and 0.01% Tween overnight at 4 °C: FBXO42 (Genentech, 96), CCDC6 (Sigma-Aldrich, HPA019051), PP2Ac (Cell Signaling, 2038), PP4c (Fortis, A300-835A), PP6c (Cell Signaling, 64221), HA (Cell Signaling, 3724), Flag (Sigma-Aldrich, M8823), HIS (Cell Signaling, 12698), UB (Cell Signaling, 43124), actin (Cell Signaling, 4970), HDAC1 (Cell Signaling, 2062), α-tubulin (Abcam, ab52866), vinculin (Cell Signaling, 4650) and RBPJ (Cell Signaling, 5442).
Membranes were incubated with 1/5,000 IRDye 800CW donkey anti-rabbit or anti-mouse for 1 h at room temperature and read by LI-COR ODYSSEY DLx. The signal was quantified using ImageStudioLite.
IF analysis
Cells were plated in 96-well plates and treated with the corresponding siRNAs for 48 h. Cells were fixed with PFA 4% for 10 min and incubated in a blocking and permeabilization buffer (10% FBS, 1% BSA, 0.1% Triton X-100 and 0.01% NaN3) for 30 min at room temperature. Cells were incubated with the following primary antibodies diluted in the same buffer overnight at 4 °C: anti-phospho-histone H3 (S10) (Cell Signaling, 3377), CENPB (Abcam ab25734) and α-tubulin (Proteintech, 66031-1-Ig). Cells were washed with PBS and incubated with 1/2,000 of ALEXA 488 or 647 secondary antibody for 1 h at room temperature. Cells were counterstained with 1/5,000 HOECHST 33342 solution. Images were taken either using a ×20 objective and analysed using the OPERA PHENIX confocal screening system or a ×60 objective on the Nikon Crest spinning-disc confocal microscope.
In vitro ubiquitination assay
Reactions (total, 40 µl) were assembled in assay buffer (1×, 30 mM Tris-HCl pH 7.5, 100 mM NaCl, 5 mM MgCl2, 5% glycerol) containing 2 mM ATP and 2 mM DTT. Final component concentrations were: E1 (UBA1), 0.1 µM; E2 (CDC34), 2 µM; ubiquitin, 20 µM; CUL1–RBX1, 0.15 µM; SKP1–FBXO42, 0.15 µM; CCDC6, 0.15 µM; and the substrate, 0.15 µM. After a 10 min incubation at room temperature, the reactions were incubated at 30 °C for 5–30 min, quenched in SDS sample buffer and analysed using immunoblotting.
Phosphatase assay
Phosphatase activity was measured using the EnzChek Phosphatase Assay Kit according to the manufacturer’s instructions. Then, 5 µg of cell lysates was incubated with the substrate in the assay buffer containing either DMSO or 10 nM of okadaic acid (Cell Signaling, 5934) for 30 min before measurement.
CRISPR–Cas9 survival screen
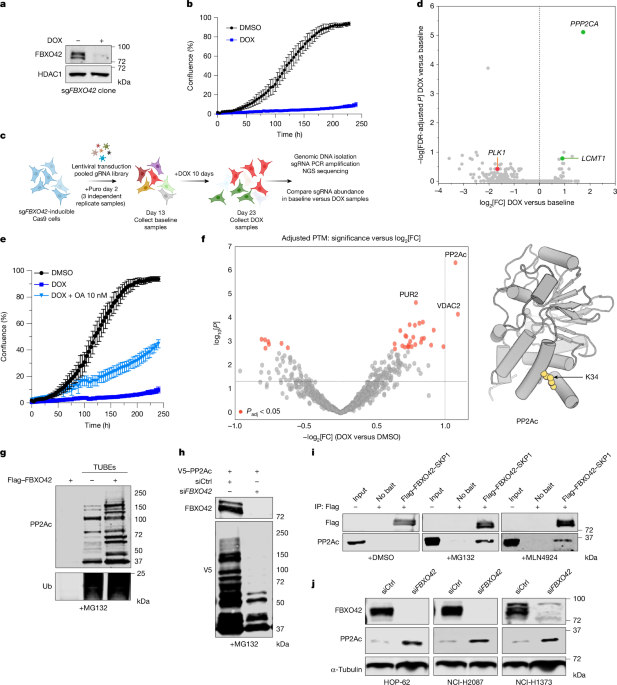
HOP-62 cells were engineered by transfection of a piggyBac transposon vector to express S. pyogenes Cas9 driven by the Tet-On 3 G DOX-inducible system and FBXO42 sgRNA from the U6 promoter. A single-cell clone was isolated that demonstrated DOX-inducible growth arrest, as well as reduced CD81 protein expression after transduction with lentiviral CD81 sgRNA. An sgRNA library targeting all protein-coding genes in the human genome at a complexity of 8 sgRNAs per gene was designed and produced in 8 plasmid pools, as described previously48, for an analogous mouse sgRNA library. The sgRNA sequences are provided in Supplementary Table 1. Library sgRNA plasmid pools were packaged into lentivirus and titred in the screening cell line as described previously48. For the screen, lentiviral stocks of the 8 sublibrary pools were combined in proportion to their viral titres. HOP-62 DOX–Cas9 sgFBXO42 cells were transduced in independent triplicate samples of 9.88 × 107 cells each with the lentiviral whole-genome sgRNA library at a multiplicity of infection of 0.46–0.59, as measured by puromycin resistance, to achieve a representation of 242–292 cells per sgRNA. All cell culture and passaging was performed on the CompacT SelecT (Sartorius Stedim Biotech) automation system in barcoded, triple-layer T500 flasks (132925, Thermo Fisher Scientific). Then, 2 days after transduction (day 2), 9.00 × 107cells were collected for a reference timepoint from each replicate, and 1.07 × 108 cells per replicate were passaged for selection with 2 μg ml−1 puromycin starting on day 3. Next, 5.20 × 107 cells per replicate were passaged without puromycin on days 6 and 9. For each replicate on day 13, 9.00 × 107 cells were collected for a baseline timepoint, and 4.00 × 107 cells of each replicate were plated and treated with 1 μg ml−1 DOX (treatment day 0). At this time, cells from each replicate were also seeded at 250 cells per well into 96-well cell culture plates and treated with and without 1 μg ml−1 DOX, and a mean of 21.5-fold reduction in cell number after 10 days of DOX treatment was measured using CellTiter-Glo (Promega) according to the manufacturer’s instructions. Medium with fresh 1 μg ml−1 DOX was replenished in all flasks on treatment day 2, 4.00 × 107 cells per replicate were passaged into DOX medium on treatment day 4 and DOX medium was replenished again on treatment day 7. On treatment day 10, 3.80 × 107 cells from each replicate were collected.
For all cell collections, genomic DNA isolation, PCR amplification of the sgRNA sequences, amplicon purification and next-generation paired-end sequencing were performed as described previously48. Sufficient genomic DNA was used for PCR amplification to maintain a representation of 250 cells per sgRNA for all baseline and DOX-treated cell samples. For each read pair, both read sequences were searched for an exact match of any one of the sgRNA barcode sequences, using the ScreenCounter R package available from the CRISPRVerse49. The number of reads matched to each guide in each sample was used to obtain a guide-by-sample count matrix for further analysis. The raw counts data were stored in a standard Bioconductor SummarizedExperiment object50. Normalization is needed to adjust for the difference in sequencing depth between samples and to account for potential compositional biases due to the competitive nature of pooled genetic screening. We estimated normalization factors for each sample by applying the TMM method51 on the count data for sgRNAs targeting a gold-standard set of non-essential genes52. The sequence reads across all samples achieved a mean sgRNA count of 350–459, with 0.04–0.12% of sgRNAs showing complete depletion (Supplementary Table 1).
We performed a differential abundance analysis at the sgRNA level using the popular limma-voom approach53. Specifically, we fitted a linear model to the log-transformed counts per million values for each sgRNA, using voom-derived observation and quality weights. We performed robust empirical Bayes shrinkage to obtain shrunken variance estimates for each sgRNA, and we used moderated F-tests to compute P values for each of the two-group comparisons of interest. To control the FDR in each comparison, we applied the standard Benjamini–Hochberg method to obtain an adjusted P value for each sgRNA (Supplementary Table 1).
We used sgRNA-level statistics to obtain gene-level summaries using two complementary statistical approaches. The first approach was to aggregate sgRNA statistics using the minimum Holm approach54, implemented in the metapod R package55, to obtain a combined P value for each gene based on the P values for all associated sgRNAs (Supplementary Table 2). Gene-level P values were corrected for multiple comparisons using the Benjamini–Hochberg method. In the second approach, DrugZ (v.1.1.0.2)56 was run using the default parameters on the raw sgRNA read counts for each comparison of interest across the three available replicates. DrugZ computes z scores of each sgRNA for the fold change between the replicate-matched baseline and DOX-treated samples. z scores are then aggregated at the gene level into a normalized z score56 (normZ; Supplementary Table 2). Both the Holm-min method P values and normZ scores identify PPP2CA as the top screen hit for sgRNA enrichment, with considerably greater statistical significance than any other gene. LCMT1 is the fourth most significant enrichment hit by both methods, with only 3 genes other than PPP2CA with slightly greater significance scores, each mapping near PPP2CA on chromosome 5.
Proteomics
IP–MS
Protein lysate (3 mg) in denaturing buffer (8 M urea, 20 mM HEPES, pH 8.0) was reduced (5 mM dithiothreitol (DTT), 45 min at 37 °C), alkylated (15 mM iodoacetamide (IAA), 20 min at room temperature in the dark) and quenched (5 mM DTT, 15 min at room temperature in the dark). Proteins were pelleted by chloroform–methanol precipitation. The resulting pellet was resuspended in denaturing buffer, diluted to 4 M urea and digested for 4 h at 37 °C with lysyl-endopeptidase (Wako) at an enzyme to protein ratio of 1:100. The sample was further diluted to 1.3 M urea and subjected to overnight enzymatic digestion at 37 °C with sequencing-grade trypsin (Promega; enzyme:protein ratio, 1:50). The resultant peptides were acidified with 20% trifluoroacetic acid (TFA, 1% final concentration), centrifuged at 18,000g for 15 min and desalted using the Sep-Pak C18 column (Waters). The eluted peptides from each treatment group were lyophilized and reserved for global proteome abundance.
For global proteome samples, 100 μg of peptides from each sample was dissolved in 100 mM HEPES pH 8.0 (1 mg ml−1). Isobaric labelling was performed using TMTPro16-plex reagents (Thermo Fisher Scientific). Each unit (0.5 mg) of TMT reagent was allowed to reach room temperature immediately before use, spun down on a benchtop centrifuge and resuspended with occasional vortexing in 20 μl anhydrous acetonitrile (ACN) before mixing with peptides (18% final ACN concentration). After incubation at room temperature for 1 h, the reaction was quenched for 15 min with 20 μl of 5% hydroxylamine. Labelled peptides were combined in equimolar ratios and dried. The TMTpro-labelled sample was redissolved in 80 μl 0.1% TFA, centrifuged at 16,000g and the supernatant was processed further. Offline high-pH reversed-phase fractionation was performed on the 1100 HPLC system (Agilent) using an ammonium-formate-based buffer system. Peptides (400 μg) were loaded onto a 2.1 × 150 mm 3.5 μm 300 Extend-C18 Zorbax column (Agilent) and separated over a 75-min gradient from 5% to 85% ACN into 96 fractions (flowrate = 200 μl min−1). The fractions were concatenated into 24 fractions, mixing different parts of the gradient to produce samples that would be orthogonal to downstream low-pH reversed phase LC–MS/MS. The fractions were dried and desalted using C18 stage-tips as previously described57. Peptides were lyophilized and resuspended in 10 μl buffer A (2% ACN, 0.1% formic acid) for LC–MS/MS analysis.
For global proteome peptides, LC–MS/MS analysis was performed by injecting 1 μl of each fraction on an Orbitrap Eclipse mass spectrometer (Thermo Fisher Scientific) coupled to the Dionex Ultimate 3000 RSLC (Thermo Fisher Scientific) system using the 25 cm IonOpticks Aurora Series column (IonOpticks) with a gradient of 2% to 30% buffer B (98% ACN, 2% H2O with 0.1% formic acid; flow rate, 300 nl min−1). Global proteome samples were analysed with a total run time of 95 min. For all samples, the Orbitrap Eclipse with FAIMS Pro DUO of −40, −60CV (proteome) collected FTMS1 scans at 120,000 resolution with an AGC target of 1 × 106 and a maximum injection time of 50 ms. FTMS2 scans on precursors with charge states of 3–6 were collected at 15,000 resolution with collision-induced dissociation (CID) fragmentation at a normalized collision energy of 30%, an AGC target of 2 × 104 and a maximum injection time of 100 ms. Real-time database search was performed before acquisition of MS3 spectra using ThermoRTS. The following parameters were used for global proteome analysis: UniProt human database August 2021 version, including 218,136 SwissProt sequences of canonical and protein isoforms, plus common contaminants and decoys; static modifications included Cys carbamidomethylation (+57.0215), Lys and N-term TMTPro (+304.207146); variable modifications included Met oxidation (+15.9949) and Tyr TMTPro (+304.207146). Offline search was performed using comet v.2019.01 with parameters matched to the RTS search. The peptide FDR was filtered to <1% using the linear discriminator algorithm. TMT reporter ions produced by the TMT tags were quantified with Mojave in-house software package by calculating the highest peak within 20 ppm of theoretical reporter mass windows and correcting for isotope purities.
Quantification and statistical testing of global proteome proteomics data were performed by MSstatsTMT_2.0.1 R package58. Multiple fractions from the same TMT mixture were combined in MSstatsTMT (v.2.0.1)58. In particular, if the same peptide ion was identified in multiple fractions, only the single fraction with the highest maximal reporter ion intensity was retained. Global median normalization was performed to reduce the systematic bias between channels.
KGG-MS: TMTPro sample preparation
Protein lysate (700 μg) in denaturing buffer (8 M urea, 20 mM HEPES, pH 8.0) was reduced (5 mM DTT, 45 min at 37 °C), alkylated (15 mM IAA, 20 min at room temperature in the dark), and quenched (5 mM DTT, 15 min at room temperature in the dark). Proteins were pelleted by chloroform–methanol precipitation. The resulting pellet was resuspended in denaturing buffer, diluted to 4 M urea and digested for 4 h at 37 °C with lysyl-endopeptidase (Wako) at an enzyme to protein ratio of 1:100. The sample was further diluted to 1.3 M urea and subjected to overnight enzymatic digestion at 37 °C with sequencing grade trypsin (Promega; enzyme:protein ratio, 1:50). The resultant peptides were acidified with 20% TFA (1% final concentration), centrifuged at 18,000g for 15 min and desalted using a Sep-Pak C18 column (Waters). Around 200 μg of eluted peptides from each treatment group was lyophilized and reserved for global proteome abundance. The remaining eluted peptides were lyophilized and used for di-glycine (KGG) analysis.
For global proteome samples, 100 μg of peptides from each sample was dissolved in 100 mM HEPES pH 8.0 (1 mg ml−1). Isobaric labelling was performed using TMTPro16-plex reagents (Thermo Fisher Scientific). Each unit (0.5 mg) of TMT reagent was allowed to reach room temperature immediately before use, spun down on a benchtop centrifuge and resuspended with occasional vortexing in 20 μl anhydrous ACN before mixing with peptides (18% final ACN concentration). After incubation at room temperature for 1 h, the reaction was quenched for 15 min with 20 μl of 5% hydroxylamine. Labelled peptides were combined in equimolar ratios and dried. The TMTpro-labelled sample was redissolved in 80 μl 0.1% TFA, centrifuged at 16,000g, and the supernatant was processed further. Offline high-pH reversed-phase fractionation was performed on the 1100 HPLC system (Agilent) using an ammonium-formate-based buffer system. Peptides (400 μg) were loaded onto a 2.1 × 150 mm 3.5 μm 300 Extend-C18 Zorbax column (Agilent) and separated over a 75-min gradient from 5% to 85% ACN into 96 fractions (flowrate, 200 μl min−1). The fractions were concatenated into 24 fractions, mixing different parts of the gradient to produce samples that would be orthogonal to downstream low-pH reversed-phase LC–MS/MS. The fractions were dried and desalted using C18 stage-tips as previously described59. Peptides were lyophilized and resuspended in 10 μl buffer A (2% ACN, 0.1% formic acid) for LC–MS/MS analysis.
For ubiquitylome quantitation of KGG peptides, lyophilized peptides were reconstituted in 1× detergent containing IAP buffer (Cell Signaling Technology) for immunoaffinity enrichment. KGG peptide enrichment was performed at 4 °C on the MEA2 automated purification system (Phynexus) using 1 ml Phytips (Phynexus) packed with 20 μl ProPlus resin coupled to 200 μg of anti-KGG (Cell Signaling Technology) antibody. Phytip columns were equilibrated for 2 cycles (1 cycle, aspiration and dispensing, 0.9 ml, 0.5 ml min−1) with 1 ml 1× IAP buffer before contact with peptides. Phytip columns were incubated with peptides for 16 cycles of capture, followed by 6 cycles of washing, twice with 1 ml 1× IAP buffer and 4× with 1 ml water. Captured peptides were eluted with 60 μl 0.15% TFA in 8 cycles in which the volume aspirated/dispensed was adjusted to 60 μl. Enriched ubiquitinated peptides were prepared as previously described60. Labelled peptides were combined, dried and resolubilized in 0.15% TFA for high-pH reversed-phase fractionation using a commercially available kit (Thermo Fisher Scientific). Fractionation was performed according to the manufacturer’s protocol with a modified elution scheme where 11 fractions were collected (F1, 13.5% ACN; F2, 15% ACN; F3, 16.25% ACN; F4, 17.5 ACN; F5, 20% ACN; F6, 21.5% ACN; F7, 22.5% ACN; F8, 23.75% ACN; F9, 25% ACN; F10, 27.5% ACN; and F11, 30% ACN) and then combined into 6 fractions (F1 + F6, F2 + F7, F8, F3 + F9, F4 + F10, F5 + F11). Peptides were lyophilized and resuspended in 10 μl buffer A for LC–MS/MS analysis.
For global proteome and ubiquitylome quantitation of KGG peptides, LC–MS/MS analysis was performed by injecting 1 μl (Proteome) or 2.5 μl (KGG) of each fraction on an Orbitrap Eclipse mass spectrometer (Thermo Fisher Scientific) coupled to the Dionex Ultimate 3000 RSLC (Thermo Fisher Scientific) system using a 25 cm IonOpticks Aurora Series column (IonOpticks) with a gradient of 2% to 30% buffer B (98% ACN, 2% H2O with 0.1% formic acid; flow rate, 300 nl min−1). Global proteome samples were analysed with a total run time of 95 min and ubiquitylome samples were analysed with a total run time of 180 min. For all samples, the Orbitrap Eclipse with FAIMS Pro DUO of −40, −60CV (proteome) or −40, −60 and −75CV (KGG) collected FTMS1 scans at a resolution of 120,000 with an AGC target of 1 × 106 and a maximum injection time of 50 ms. FTMS2 scans on precursors with charge states of 3–6 were collected at 15,000 resolution with CID fragmentation at a normalized collision energy of 30%, an AGC target of 2 × 104 (proteome) or 2 × 105 (KGG), and a maximum injection time of 100 ms (proteome) or 200 ms (KGG).
RTS was performed before acquisition of MS3 spectra using InSeqAPI software61, which operated similarly to previously published approaches62,63. The following RTS parameters were used for global proteome analysis: UniProt human database August 2021 version, including 218,136 SwissProt sequences of canonical and protein isoforms, plus common contaminants and decoys; static modifications included Cys carbamidomethylation (+57.0215), Lys and N-term TMTPro (+304.207146); variable modifications included Met oxidation (+15.9949) and Tyr TMTPro (+304.207146). Offline search was performed using comet v.2019.01 with parameters matched to the RTS search. The peptide FDR was filtered to <1% using the linear discriminator algorithm. TMT reporter ions produced by the TMT tags were quantified using Mojave custom software package by calculating the highest peak within 20 ppm of theoretical reporter mass windows and correcting for isotope purities.
Quantification and statistical testing of global proteome proteomics data were performed by MSstatsTMT_2.0.1 R package58. Multiple fractions from the same TMT mixture were combined in MSstatsTMT (v.2.0.1)58. In particular, if the same peptide ion was identified in multiple fractions, only the single fraction with the highest maximal reporter ion intensity was retained. Global median normalization was carried out to reduce the systematic bias between channels.
RAW spectra files can be found on MassIVE with the dataset identifier: MassIVE MSV000094889.
Molecular biology and protein expression
All constructs were generated through gene synthesis at Genscript with codon optimization for insect cell expression (except for LCMT-1, which was expressed in Escherichia coli). DNA encoding the genes of interest was cloned into modified versions of the baculovirus transfer vector pAcGP67A or pAcSG2 (BD Biosciences), adapted for high-throughput cloning. The expression cassette is flanked by segments of the baculovirus genome to facilitate transfer into the linearized baculovirus DNA by homologous recombination in insect cells64. Baculovirus was generated using homologous recombination between BEVS transfer vector and BestBac 2.0 Linearized Bacmid (Expression Systems, 91-002) in a high-throughput manner as described earlier for generating P2 virus65. P3 stocks of baculovirus for large-scale production were amplified for 4 days in a flask at 200 ml followed by collection and used for infecting 8–10 l of Sf9 or Tni cells for protein production. Cells were seeded at a density of 2 × 106 cells per ml and infected at multiplicity of infection of 0.5 for Sf9 cells and multiplicity of infection of 1 for Tni cells. Sf9 infected cells were collected after 3 days and Tni cells were collected after 2 days. Pellets were frozen, followed by purification.
Protein purification
Flag–FBXO42–SKP1
Human FBXO42 (UniProt: Q6P3S6) with an N-terminal truncation (Δ2–44) and an N-terminal Flag tag, and Human SKP1 (UniProt: P63208) with deletions of residues 37–42 and 70–81 (70-81 were replaced with a GGSG linker), were separately cloned (pAcGP67A) and co-expressed in Sf9 cells using the protocol described above. The frozen cell pellet of expressed Flag–FBXO42–SKP1 was resuspended and thawed in CytoBuster Protein Extraction Reagent (EMD Millipore, 50 ml reagent per 1 l biomass), 1 mM TCEP pH 7.5, benzonase (1:1,000 dilution of 0.5 mg ml−1 benzonase stock:lysate), 5 mM MgCl2, 5 mM ATP pH 7.3 and cOmplete EDTA-free Protease Inhibitor Cocktail (Roche, 1 tablet per 50 ml of lysate). The thawed lysate was homogenized with a sonicator until the cell mixture was evenly distributed. The homogenized cell lysates were incubated at room temperature for 5 min and then incubated with anti-DYKDDDDK magnetic beads (GenScript, 0.8–1 ml settled beads per 1 l biomass) on a shaker (Stuart Orbital Shaker SSL1) at 170 rpm for 1 h at 4 °C. The lysates were separated from the magnetic beads using an AmMag Block (GenScript). Batch purification methods were used to purify Flag–FBXO42–SKP1. Anti-DYKDDDDK Magnetic Beads underwent two washes with 5× settled bead volume of wash buffer (50 mM Tris pH 7.5, 150 mM NaCl, 5 mM MgCl2, 5 mM ATP pH 7.3, 10% glycerol, 1 mM TCEP pH 7.5), followed by 3 elutions with 3× settled bead volume of elution buffer (50 mM Tris pH 7.5, 150 mM NaCl, 5 mM MgCl2, 10% glycerol, 1 mM TCEP pH 7.5, 0.3 mg ml−1 3×Flag peptide). The AmMag MR magnetic rack (GenScript) was used to bind beads, decant washes and decant eluates. For each wash, the mixture of buffer and beads was gently inverted 5–6 times before decanting. For each elution, the mixture of buffer and beads was incubated for 1 h at 4 °C before decanting eluates. Incubation of Anti-DYKDDDDK magnetic beads and batch purification methods were carried out twice on the same lysate. All washes and eluate fractions that contained Flag–FBXO42–SKP1 were combined, concentrated and passed over the HiLoad 16/600 Superdex 200 pg (Cytiva) column equilibrated in the following SEC buffer: 20 mM HEPES pH 7.5, 200 mM NaCl, 10% glycerol and 1 mM TCEP pH 7.5. The protein concentration was measured using the absorbance at 280 nm (NanoDrop). The final SEC fractions containing Flag–FBXO42–SKP1 were combined, concentrated, flash-frozen in liquid nitrogen and stored at −80 °C until use.
SCFFBXO42
Human CUL1 (UniProt: Q13616) with an N-terminal His-tag and human RBX1 (UniProt: P62877) were cloned into a bicistronic pAcGP67A vector in which each target was downstream of a polyhedrin promoter. Human CUL1, human RBX1, human FBXO42 and human SKP1 were co-expressed in Sf9 cells using the protocol described above. Human FBXO42 and human SKP1 were the same constructs as those used in the Flag–FBXO42–SKP1 purification method above. The frozen cell pellet of expressed SCFFBXO42 was resuspended and thawed in lysis buffer (40 mM HEPES pH 7.5, 200 mM NaCl, 10% glycerol; 50 ml buffer per l of biomass), benzonase (1:1,000 dilution of 0.5 mg ml−1 benzonase stock:lysate), 5 mM MgCl2, 5 mM ATP pH 7.3 and cOmplete EDTA-free protease inhibitor cocktail (Roche, 1 tablet per 50 ml of lysate). The thawed cell lysates were passed through a microfluidizer three times. Whole-cell lysates were clarified by ultracentrifugation at 186,000g and filtered by a 0.8-µm filter followed by a 0.45-µm filter. The clarified lysates were gently rocked on a nutator for 1 h at 4 °C with anti-Flag M2 affinity gel resin (Sigma-Aldrich, 1.25 ml settled resin per 1 l of biomass). Resin was separated from lysate by centrifugation (700g). Resin was applied to a gravity column (Bio-Rad) and the sample was washed with 20 column volumes of lysis buffer. SCFFBXO42 was eluted with lysis buffer containing 0.3 mg ml−1 3× Flag peptide. Eluates were supplemented with 3 mM TCEP pH 7.5. Incubation of anti-Flag M2 affinity gel resin and gravity purification methods were performed twice on the same lysate. The eluate fractions that contained SCFFBXO42 were combined, concentrated and passed over the HiLoad 16/600 Superdex 200 pg (Cytiva) column in the following SEC buffer: 20 mM HEPES pH 7.5, 200 mM NaCl, 10% glycerol, 3 mM TCEP pH 7.5. The protein concentration was measured using the absorbance at 280 nm (NanoDrop). The final SEC fractions containing SCFFBXO42 were combined, concentrated, flash-frozen in liquid nitrogen and stored at −80 °C until use.
HisAvi(biotin)–FBXO42–SKP1
Full-length human FBXO42 (UniProt: Q6P3S6), with N-terminal His and Avi tags, and SKP1 (the same construct used for Flag–FBXO42–SKP1 above) were separately cloned (pAcGP67A) and co-expressed in Sf9 cells using the protocol described above. The frozen cell pellet of HisAvi(biotin)–FBXO42–SKP1 was resuspended and thawed in 50 mM Tris pH 8.0, 150 mM NaCl, 1 mM TCEP and 0.05% Triton X-100 with protease inhibitor tablets. The cell lysates were homogenized and stirred on ice for 30 min, then centrifuged (186,000g) for 1 h. Next, 5 ml of NiNTA agarose resin was added to the supernatant, and rotated at 4 °C for 90 min. Resin was centrifuged (700g) for 5 min, then washed twice with 50 mM Tris pH 8.0, 150 mM NaCl and 1 mM TCEP. The resin was further washed with 25 ml of 25 mM Tris pH 8.0, 150 mM NaCl, 1 mM TCEP and 7.5 mM imidazole, then 25 ml of 25 mM Tris pH 8.0, 150 mM NaCl, 1 mM TCEP and 12.5 mM imidazole. Protein was eluted with 25 mM Tris pH 8.0, 150 mM NaCl, 1 mM TCEP and 500 mM imidazole. The fractions were analysed using an SDS–PAGE gel. The fractions containing FBXO42–SKP1 were pooled together and diluted 3× in 25 mM Tris pH 8.0 and 1 mM TCEP. The sample was loaded onto the 5 ml Hitrap Q HP (Cytiva) column, then eluted with a 0–500 mM NaCl gradient over 20 column volumes (100 ml, 0–50% B). The fractions containing FBXO42–SKP1 were pooled together and concentrated to 80–100 μM, then diluted to 40–50 μM with 25 mM Tris pH 8.0 and 1 mM TCEP; the NaCl concentration was 150 mM. Avidity’s biotin reaction kit was used, and the sample was incubated at room temperature overnight. Details of the protocol are available online (https://www.avidity.com/commerce/product.asp?NUMBER=1). After biotinylation, the protein sample was concentrated and loaded onto the HiLoad 16/600 Superdex 200 pg (Cytiva) column pre-equilibrated in 25 mM Tris pH 8.0, 150 mM NaCl, 1 mM TCEP. The protein concentration was measured using the absorbance at 280 nm (NanoDrop). The final SEC fractions containing HisAvi(biotin)–FBXO42–SKP1 were combined, concentrated, flash-frozen in liquid nitrogen and stored at −80 °C until use.
PP2Ac, PP4c and PP6c
Human PP2Ac (UniProt: P67775-1), human PP4c (UniProt: P60510) and human PP6c (UniProt: O00743-1), each with both N-terminal Flag and HA (YPYDVPDYA) tags, were individually cloned (pAcGP67A) and expressed in Tni cells using the protocol described above.
Purification was carried out using the same protocol as Flag–FBXO42–SKP1 above, with the following exceptions: 5 mM MgCl2 was replaced with 1 mM MnCl2; 0.1–0.7 ml of settled anti-DYKDDDDK magnetic beads (Genscript) were used per 1 l biomass; the HiLoad 16/600 Superdex 75 pg (Cytiva) column was used instead of the Superdex 200. The final SEC buffer was as follows: 20 mM HEPES pH 7.5, 200 mM NaCl, 1 mM MnCl2, 10% glycerol, and 1 or 3 mM TCEP pH 7.5. The protein concentration was measured using the absorbance at 280 nm (NanoDrop). The final SEC fractions containing the PP2Ac, PP4c or PP6c were separately combined, concentrated, flash-frozen in liquid nitrogen and stored at −80 °C until use. Human PP2Ac constructs with a N-terminal His tag, both with and without the C-terminal tail (Δ296-309), were purified using the same method with the following exceptions: Ni-charged Magbeads (GensScript, 1 ml settled resin per 1 l biomass) were used instead of anti-DYKDDDK magnetic beads, and the samples were further processed by a 5 ml HisTrap HP (Cytiva) column using a gradient elution (similar to PP2Aa+c dimer described below) and a 1 ml HiTrap Q (Cytiva) column (similar to His-CCDC6 described below) before being passed over the Superdex 75 column.
PP2Aa+c dimer
Human PP2Ac (UniProt: P67775-1) with an N-terminal Flag tag and human PP2Aa (UniProt: P30153) with an N-terminal His tag were individually cloned (pAcGP67A) and co-expressed in Tni cells using the protocol described above. Cell lysis and Flag affinity purification were performed using the same methods as for Flag–FBXO42–SKP1 above, with the following exceptions: 2 ml of settled anti-DYKDDDDK magnetic beads (GenScript) were used per 1 l biomass; incubation of anti-DYKDDDDK magnetic beads and batch purification methods were carried out three times on the same lysate. Flag eluates were combined and imidazole was added to 20 mM concentration. The sample was further processed by a 5 ml HisTrap HP (Cytiva) column using a gradient elution (starting buffer: 25 mM Tris pH 7.5, 150 mM NaCl, 20 mM imidazole, 10% glycerol, 5 mM ATP pH 7.3, 5 mM MgCl2, 1 mM TCEP pH 7.5; final elution buffer: starting buffer with 300 mM imidazole and without ATP). Eluates containing dimer were passed over the HiLoad 16/600 Superdex 200 pg (Cytiva) column equilibrated in the following SEC buffer: 20 mM HEPES pH 7.5, 300 mM NaCl, 10% glycerol, 1 mM TCEP pH 7.5. The protein concentration was measured using absorbance at 280 nm (NanoDrop). The final SEC fractions containing the PP2a+c dimer were combined, concentrated, flash-frozen in liquid nitrogen and stored at −80 °C until use.
His–CCDC6
Human CCDC6 (UniProt: Q16204) with an N-terminal His tag was cloned (pAcGP67A) and expressed in Sf9 cells using the protocol described above. The frozen cell pellet of expressed His–CCDC6 was lysed using the same method as for Flag–FBXO42–SKP1 with the following exceptions: Ni-charged MagBeads (GenScript, 1 ml settled beads per 1 l biomass) were used instead of anti-DYKDDDDK magnetic beads. Batch purification methods were used to purify His-CCDC6. Ni-charged MagBeads underwent three washes with 5× settled bead volume of wash buffer (25 mM Tris pH 7.5, 150 mM NaCl, 20 mM imidazole, 10% glycerol, 2 mM MgCl2, 1 mM TCEP pH 7.5), followed by 5–8 washes with 1.5× settled bead volume of elution buffer (wash buffer with 300 mM imidazole pH 8.0). The AmMag MR magnetic rack (GenScript) was used to bind beads, decant washes and decant eluates. For each wash and elution, the mixture of buffer and beads was gently inverted 5–6 times before decanting. Incubation of Ni-charged MagBeads and batch purification methods were carried out four times on the same lysate. All washes and eluate fractions that contained His–CCDC6 were combined, diluted to a final NaCl concentration of 20–50 mM, and processed by a 5 ml HiTrap Q HP (Cytiva) column using a gradient elution (starting buffer: 25 mM Tris pH 8.5, 20 mM NaCl, 1 mM TCEP pH 7.5; elution buffer: starting buffer with 1 M NaCl). Fractions containing His–CCDC6 were combined, concentrated and passed over the HiLoad 16/600 Superdex 200 pg (Cytiva) column equilibrated in the following SEC buffer: 20 mM HEPES pH 7.5, 200 mM NaCl, 10% glycerol, 1 mM TCEP pH 7.5. A portion of His-tagged CCDC6 was incubated overnight at 4 °C with TEV protease (in-house) to cleave the N-terminal His tag before injection onto the SEC column. Intact LC–MS was used to confirm His tag removal. The protein concentration was measured using absorbance at 280 nm (NanoDrop). The final SEC fractions containing CCDC6 were combined, concentrated, flash-frozen in liquid nitrogen and stored at −80 °C until use.
PP2Aa/B56
Human PP2Aa (UniProt: P30153) and residues Ile30-Ala437 of B56 (UniProt: Q13362-1), each with an N-terminal His tag, were individually cloned (pAcGP67A) and expressed in Tni cells using the protocol described above. Purification was carried out using the same method as in His–CCDC6 above, with the following exception: samples purified by Ni-charged MagBeads were further processed by a 5 ml HisTrap HP (Cytiva) column using gradient elution (buffers were the same as the batch purification) before being processed by SEC. To obtain a sample without His tag, purified protein was incubated overnight at 4 °C with hyperTEV6066 protease (in-house). His tag removal was confirmed by intact LC–MS and the sample was further processed by reverse nickel using the 1 ml HisTrap HP (Cytiva) column. The flow through from reverse nickel was passed over a HiLoad 16/600 Superdex 200 pg (Cytiva) column equilibrated in the following SEC buffer: 25 mM Tris pH 7.5, 150 mM NaCl, 10% glycerol, 1 mM TCEP pH 7.5. The protein concentration was measured using absorbance at 280 nm (NanoDrop). The final SEC fractions containing PP2Aa or B56 were separately combined, concentrated, flash-frozen in liquid nitrogen and stored at −80 °C until use.
Mutant proteins
All mutant FBXO42 and CCDC6 proteins were expressed at a 200 ml scale in Sf9 cells and purified using a semi-automated parallel affinity purification method, as described previously65. In brief, the proteins were lysed and purified in a buffer of 25 mM HEPES pH 7.5, 150 mM NaCl, 1 mM TCEP using Integrated Micro Chromatography Systems tips on the Hamilton STAR liquid handler. The elutions were performed using 150 μg ml−1 of 3×Flag peptide for Flag-tagged proteins and 250 mM imidazole for His-tagged proteins. The protein concentration was measured using absorbance at 280 nm (NanoDrop). Proteins were flash-frozen in liquid nitrogen and stored at −80 °C until use.
LCMT-1
LCMT-1 (UniProt: Q9UIC8-1) with an N-terminal His tag was cloned (pAcSG2) and expressed in BL21 DE3 cells at 16 °C overnight after induction with 0.4 mM IPTG. Cells were lysed in 20 mM HEPES pH 7.5, 200 mM NaCl, 20 mM imidazole, 0.5 mM TCEP (lysis buffer), and Roche EDTA-free protease inhibitors using a microfluidizer. Whole-cell lysates were clarified by ultracentrifugation (186,000g), and then filtered by a 0.8 μm filter. The sample was then applied to a 5 ml HisTrap column (Cytiva) pre-equilibrated in the lysis buffer. Protein was eluted using lysis buffer with 300 mM imidazole. The elution fractions containing LCMT-1 were processed by anion exchange using a 5 ml Hi Trap Q (Cytiva) column with a gradient elution (starting buffer: 25 mM Tris pH 8.0, 0.5 mM TCEP; elution buffer: starting buffer with 0.5 M NaCl). The fractions containing LCMT-1 were combined, incubated with TEV protease (in-house) to cleave the N-terminal His tag and processed again by anion exchange. The fractions containing LCMT-1 were passed over a HiLoad 16/600 Superdex 200 pg (Cytiva) column equilibrated in 50 mM HEPES pH 7.5, 200 mM NaCl, 1 mM TCEP. The protein concentration was measured using the absorbance at 280 nm (NanoDrop). The final SEC fractions containing LCMT-1 were concentrated, flash-frozen in liquid nitrogen and stored at −80C until use.
PP2Ac methylation
A 20:2 µg ratio of PP2Ac:LCMT-1 was combined with 200 μM SAM (New England Biolabs) and the mixture was incubated at room temperature. The reaction was monitored by intact LC–MS at the following timepoints: 1 h, 2 h, 5 h and 24 h. Analysis was performed on the Agilent 6230 LC/TOF mass spectrometer coupled to an Agilent 1260 Infinity HPLC system with an AdvanceBio RP-mAb Diphenyl column (2.1 × 50 mm). The sample was eluted by gradient elution with 0.1% trifluoroacetic acid in ACN. Data were analysed using Agilent MassHunter Qualitative Analysis software (Agilent v.B.07.00) and plotted using GraphPad Prism v.10.0.3 for macOS, GraphPad Software (www.graphpad.com). Intact LC–MS spectra were evaluated for a 14 Da increase from the molecular mass of PP2Ac (38,841 Da). At the 24 h timepoint, nearly all PP2Ac had been methylated (Extended Data Fig. 4f).
Cryo-EM sample preparation, data collection, image processing and model building
Cryo-EM sample preparation
Cryo-EM samples were prepared by incubating 1.5 nmol each of mPP2Ac and CCDC6 for 2 h at 4 °C (mPP2Ac was prepared using the PP2Ac methylation method described above with a 88:13 µg ratio of PP2Ac:LCMT1; the reaction was done at a final concentration of 2 mg ml−1 PP2Ac from which 1.5 nmol was used to prepare the cryo-EM sample). Then, 0.8 nmol of SCFFBXO42 was added to mPP2Ac–CCDC6 and the entire sample was allowed to incubate overnight at 4 °C. The entire sample was passed over the Superose 6 Increase 3.2/300 (Cytiva) column equilibrated in the following SEC buffer: 20 mM HEPES pH 7.5, 200 mM NaCl, 1 mM TCEP pH 7.5. The two centre fractions on the peak indicated in Extended Data Fig. 4g (asterisks) were combined (around 0.2 mg ml−1 as measured by NanoDrop, absorbance at 280 nm) and vitrified on grids for cryo-EM analysis. A similar protocol was used for the 3×FBXO42-CCDC6–mPP2Ac structure, with the exception that the sample was mixed with in-house developed cryoprotective peptides immediately before vitrification.
Data for all structures were collected from Ultraufoil 300 mesh 1.2/1.3 (Quantifoil) grids coated in self-assembled monolayers (4 mM monothiolalkane(C11)PEG6-OH (11-mercaptoundecyl) hexaethyleneglycol (SPT-0011P6, SensoPath Technologies))67 and blotted using the Vitrobot MARK IV (Thermo Fisher Scientific) for 3 s at 7 force. Additional data collection for the 3×FBXO42–SKP1-bound CCDC6–mPP2Ac structure was obtained from Ultraufoil 300 mesh 1.2/1.3 (Quantifoil) grids subjected to glow discharge using a Solarus Model 950 Advanced Plasma System (Gatan). All grids were clipped for use with a Thermo Fisher Scientific autoloader (Nanosoft).
Cryo-EM data collection
For the high-resolution FBXO42 degradasome, 21,534 videos were collected on the Krios G3i microscope (Thermo Fisher Scientific) with a BioQuantum energy filter and K3 camera (Gatan). The K3 detector was operated in non-CDS counting mode and the energy filter slit width was set to 20 eV. Videos were collected at a nominal magnification of ×105,000, physical pixel size 0.838Å px−1, with a 50 µm C2 aperture at a dose rate of 15 e− px−1 s−1. A total dose of 64 e− Å−2 was collected as a 59-frame video, resulting in a 3 s video with 1.08 e− per frame. Data were collected semi-automatically using SerialEM68 image shift pattern of three images per hole in a 3 × 3 pattern. The nominal defocus was set to −0.8 to −2.4 µm. Beam tilt was corrected for using SerialEM’s image shift vs coma calibration.
For the low-resolution 3×FBXO42 complex, 4,412 videos were collected on the Glacios1 microscope (Thermo Fisher Scientific) with a K3 camera (Gatan). The K3 detector was operated in non-CDS counting mode. Videos were collected at a nominal magnification of ×36,000, physical pixel size 1.114Å px−1, with a 50 µm C2 aperture and 100 µm at a dose rate of 15 e− px−1 s−1. A total dose of 36 e− Å−2 was collected as a 30-frame video, resulting in a 3 s video with 1.2 e− per frame. Data were collected automatically using Smartscope69 and SerialEM68 image shift pattern of three images per hole in a 3 × 3 pattern. Nominal defocus was set to −1.0 to −2.5 µm. Beam tilt was corrected for using SerialEM’s image shift vs coma calibration.
Cryo-EM image processing
For the high-resolution FBXO42 degradasome, cryo-EM data were transferred, patch motion-corrected, patch CTF-estimated, template picked and extracted in a 450 px box binned to 128 px. In total, 7,261,626 putative particles were picked and after two rounds of 2D classification and selection, 2,309,685 particles were further classified in 3D. In total, 551,049 of particles from the classes with the largest assemblies were used to generate 5 ab initio references. The 2.3 million particles were subjected to four rounds of subsequent heterogeneous refinement using these five classes as input references. Then, 578,587 particles from two large assembly classes were subjected to a new round of ab initio reconstruction to generate five new 3D references. The same 578,587 particles were run through a round of heterogenous refinement in parallel, the best FBXO42-bound class from each selected, and the duplicates were removed before performing a NU-refinement with dynamic masking. Global CTF refinement and another round of heterogenous refinement were performed on the remaining 147,335 particles. After the last round of heterogeneous refinement, 98,992 particles were extracted from the original micrographs at a pixel size of 1.24 Å px−1. Concurrently the original 2.3 million particles were subjected to three rounds of heterogenous refinement with a good FBXO42-bound class from the original ab initio references and three junk classes from a previous, unrelated dataset. Then, 408,123 particles were recovered in the good class and subjected to a new round of ab initio reconstruction to generate four new volumes. After a final round of heterogenous refinement, 135,171 particles were extracted at 1.24Å px−1, combined with the original track particles, and duplicates within 50 Å of each other were removed. The remaining 155,244 particles were run through reference-based motion correction at pixel size 0.838 Å px−1, NU refinement and multiple local refinements to generate maps that ranged in resolution from 3.0 to 4.2 Å. From the second track of classification, two classes were selected from the final heterogeneous reconstruction representing 8× occupied PP2Ac. These particles were extracted at 1.24 Å px−1, duplicates within 50 Å were removed and a NU-refinement yielded a map at 3.5 Å. Local refinements were run with multiple masks to generate maps ranging from 3.2 to 4.5 Å resolution. All data were processed in cryoSPARC (v.4.4.0)70.
For the low-resolution 3×FBXO42 complex, cryo-EM data were transferred, patch motion-corrected and patch CTF-estimated in cryoSPARC live. Images with CTF fits higher than 8 Å were processed. Initial 2D averages for template picking were generated from 8,889 particle blobs picked with a diameter of 140–170 Å. Better templates were generated by template picking 510,891 particles with a diameter of 180 Å. These templates were used to pick particles from 2609 micrographs. Then, 32,536 particles from the best 2D averages were used to generate ab initio models and 2,672,451 particles were subjected to iterative heterogeneous refinement. After a final stack of 209,666 particles was refined using NU-refinement, these particles were subjected to 3D classification without alignment at different resolution filters. Classes with any indication of more than two FBXO42 densities bound were subjected to another round of 3D classification and selection. The final particle stack of 57,743 particles was used to train a Topaz model for picking 3,111 micrographs. These 2,468,634 particles were subjected to heterogeneous refinement using the original ab initio models. A final stack of 158,066 particles was subjected to focused 3D classification and the classes with strongest density for the third FBXO42 density were selected. Two classes from the initial 2D classification that also showed extra FBXO42 density around the central core were also selected. These stacks of particles were merged, and duplicates were removed to give a final stack of 28,195 particles. These particles yielded a 6.9 Å map after local refinement around the central PP2A and FBXO42 densities. The local resolution in the map ranged from 5 to 15 Å.
Model building and refinement
Model building for the FBXO42-bound degradosome (FCP) and CCDC6–PP2Ac (CP) complexes was done by combining all the local refinements in a single composite map using the Combine Focused Maps job in Phenix71. A PP2Ac model was derived from PDB 3FGA and rigid-body fit into the maps using ChimeraX72. We generated a model of FBXO42–SKP1 using AlphaFold229 and rigid body fit into the maps using ChimeraX. The resolution in the map was sufficiently high enough for us to manually build CCDC6 in Coot73. Models were then subjected to local and global real-space refinement in Coot and Phenix. For the 3×FBXO42 structure, we first docked in the high-resolution FCP structure, and used the coordinates of FBXO24–SKP1 derived from the FCP structure to dock in the third copy using ChimeraX. Coordinates of this low-resolution model have been uploaded to Zenodo (https://doi.org/10.5281/zenodo.17260372). Final statistics of the maps and models are reported in Extended Data Table 1.
NanoBit
Construct generation
The NanoBiT constructs and the respective amino acid boundaries of the LgBiT-tagged proteins are annotated as follows: LgBiT-PP2Ac (aa. 2-309); LgBiT-PP4c (aa. 2-307); LgBiT-PP6c (aa. 2-305); FBXO42-SmBiT-IRES-mNeon (aa. 95-717). The cDNAs were synthesized, codon optimized for mammalian expressions, and then cloned into a pRK5 vector, which is adapted for high-throughput cloning. Recombinant genes are expressed as fusion proteins when cloned into one of the available restriction enzyme sites under the control of the strong CMV promoter.
Stable cell line generation and maintenance
The FBXO42∆F-box-SmBiT_IRES_mNeon construct was used for stable cell line generation. A549 FBXO42 KO cells were seeded at 2.5 × 106 per 15 cm2 TC-treated dish in RPMI medium 24 h before transfection. For transfection, 7 μg of the DNA construct and 3 μg of the PiggyBac Transposase Expression Vector were diluted in Opti-MEM (Gibco, 31985-062). FuGene HD Transfection Reagent (Promega, E2312) was used according to the manufacturer’s instructions at a ratio of 3 μg (FuGene):1 μg (DNA). After a brief incubation, diluted DNA and transfection reagents were mixed and incubated for 5 min. The mixture was added to the cells drop by drop. Cells were incubated at 37 °C TC incubator with 5% CO2 for 48 h before medium exchange containing 200 μg ml−1 of Zeocin (Thermo Fisher Scientific, R25001) for 2 weeks.
Transfection, cell lysis and NanoBiT assay
The protocol was adopted from a previous study74. A549 FBXO42 KO cells stably expressing FBXO42∆F-box-SmBiT fusion were maintained in a tissue culture incubator at 37 °C and 5% CO2 in RPMI medium supplemented with FBS (v/v, 10%), l-glutamine (3 mM) and penicillin–streptomycin antibiotics (100 U ml−1). Before transfection, cells were seeded at 2.5 × 106 into 15 cm2 TC-treated dishes for 24 h. Individual transfection of LgBiT constructs was carried out according to the manufacturer’s instructions using FuGene HD Transfection Reagent (Promega, E2311) diluted with Opti-MEM (Gibco, 31985-062) at a ratio of 3 μg (FuGene):1 μg (DNA). For individual transfection, 7 μg DNA was used per 15 cm2 dish. After 48 h of transfection, the medium was removed, and the cells were washed once with PBS buffer. The cells were then collected and lysed with 1× passive buffer (Promega, E1941) supplemented with protease (Roche, 05892970001) and phosphatase (Roche, 04906837001) inhibitors according to the manufacturer’s instructions. The cell lysates were sonicated for 10 s and centrifuged in a tabletop centrifuge at 4 °C for 10 min at maximum speed. The cell lysates were then quantified using a BCA protein assay kit (Thermo Fisher Scientific, 23227), snap-frozen in liquid nitrogen and then stored at −80 °C. Thawed cell lysates were diluted using titration buffer (25 mM Tris pH8, 100 mM NaCl) to 1 μg μl−1 and dispensed to white Proxiplates (PerkinElmer, 6008280) (7 μl per well). The Nano-Glo Live Cell Substrate (Promega, N2012) was prepared according to the manufacturer’s instructions, and 3 μl of the diluted substrate was added per well. Plates were then centrifuged at 360 rpm for 1 min and incubated for 10 min on a shaker at 300 rpm, and the luminescence signal was recorded using the Envision plate reader.
HTRF assays
FBXO42, CCDC6 and phosphatase binding HTRF
First, CCDC6 was prepared at concentrations of 120 nM, 60 nM, 30 nM and 0 nM (control) in the assay buffer (20 mM HEPES pH 7.5, 200 mM NaCl, 1 mM TCEP pH 7.5, 0.1% BGG). This was done in the presence of 2 nM Europium-anti-Flag donor dye (PerkinElmer, AD0274). Next, twofold serial dilutions of 240 nM Flag–HA–PP2Ac, Flag–HA–PP4c or Flag–HA–PP6c were performed at the fixed CCDC6 concentrations described above. These mixtures were then incubated for 30 min on a rotor with mild agitation. Third, the acceptor mix was prepared by labelling either 120 nM, 60 nM or 0 nM (control) biotinylated Avi–FBXO42–SKP1 with 0.01 mg ml−1 Streptavidin-XL665 acceptor (PerkinElmer, 610SAXLB). This solution was incubated for 30 min. Finally, an equal amount (4 µl) of each donor and acceptor mix was pipetted into individual wells of assay plates (ProxiPlate-384 F Plus, Revvity 6008260) and incubated on a shaker with mild agitation for 1 h. Data were subsequently collected using the PheraStar plate reader (BMG Labtech).
FBXO42, CCDC6 and mPP2Ac binding HTRF
Methylated or non-methylated Flag–HA–PP2Ac, diluted to a 2× concentration (120 nM) in assay buffer (20 mM HEPES pH 7.5, 200 mM NaCl, 1 mM TCEP pH 7.5, 0.1% BGG), was incubated with 2 nM Europium-anti-Flag donor fluorophore (PerkinElmer, AD0274) and 2× CCDC6 (120 nM) for 30 min at room temperature. Concurrently, 2× biotinylated Avi–FBXO42–SKP1 (120 nM) was incubated with 0.01 mg ml−1 Streptavidin-XL665 acceptor (PerkinElmer, 610SAXLB) for 30 min. An equal volume (4 µl) of both the donor and acceptor mixes was then pipetted into each well of ProxiPlate-384 F Plus assay plates (Revvity 6008260), resulting in equimolar final protein concentrations (60 nM). The plates were incubated on a shaker with mild agitation for 1 h at room temperature, after which data were collected using the PheraStar plate reader (BMG Labtech).
For peptide competition assays, the procedure was amended as follows. Proteins were labelled with either donor or acceptor fluorophores at 4x concentrations (240 nM), combined in equal volume and incubated for 1 h at room temperature. Subsequently, 4 µl of this combined mix was pipetted into each well of ProxiPlate-384 F Plus assay plates (Revvity 6008260), followed by the addition of 4 µl of serially diluted peptides at 2× concentrations in assay buffer. The HTRF ratio obtained at the 3 h timepoint is presented on the graph.
PP2Ac and CCDC6 binding HTRF assays
Non-methylated Flag–HA–PP2Ac was serially diluted in assay buffer (20 mM HEPES pH 7.5, 200 mM NaCl, 1 mM TCEP pH 7.5, 0.1% BGG, 1 mM DTT) containing 2 nM Europium-anti-Flag donor fluorophore (PerkinElmer, AD0274) at concentrations of 480 nM, 240 nM, 120 nM, 60 nM and 0 nM (control). Biotinylated CCDC6 was diluted in the assay buffer in the presence of 0.01 mg ml−1 of SA-d2 acceptor fluorophore (Revvity, 610SADLB). The proteins were then labelled for 30 min at room temperature. An equal amount (4 µl) of each donor and acceptor mix was pipetted into individual wells of assay plates (ProxiPlate-384 F Plus, Revvity 6008260). The HTRF ratio was subsequently quantified using the PheraStar plate reader (BMG Labtech) in a time-course manner. The 3 h timepoint is presented on the graph.
FBXO42 competition HTRF assays
To generate the donor mix, methylated Flag–HA–PP2Ac was diluted to a 4× concentration (240 nM) in the assay buffer (20 mM HEPES pH 7.5, 200 mM NaCl, 1 mM TCEP pH 7.5, 0.1% BGG) containing 4 nM Europium-anti-Flag donor fluorophore (PerkinElmer, AD0274), and incubated for 30 min at room temperature. CCDC6 was then serially diluted at 4× concentration using labelled PP2Ac mix to achieve various PP2Ac:CCDC6 ratios (0.5:1, 1:1, 2:1 and 4:1), as well as a sample without CCDC6. The acceptor mix was prepared by incubating biotinylated Avi–FBXO42–SKP1 at 4× concentration (240 nM) with 0.02 mg ml−1 Streptavidin-XL665 fluorophore (PerkinElmer, 610SAXLB) for 30 min at room temperature. Next, the donor and acceptor mixes were then combined in equal volumes and incubated for one hour. Following this, 4 µl of the combined mix was added to each well of the assay plate (ProxiPlate-384 F Plus, Revvity, 6008260). Finally, the competitor unlabelled non-biotinylated Avi–FBXO42–SKP1 was serially diluted in the assay buffer and 4 µl of the competitor was added to wells. A time-course quantification was then performed using the PheraStar plate reader (BMG Labtech); the 3 h timepoint was used for quantification of the apparent binding affinity. The apparent binding affinity (KD) was calculated using the nonlinear fit of the HTRF competition assay results. The HTRF signal was normalized to the total FBXO42 (normalized HTRF signal = (observed HTRF signal/[FBXO42labelled]) × [FBXO42total]).
Holoenzyme HTRF assays
To assess whether CCDC6 can induce the disassembly of the PP2A holoenzyme complex, 4× concentrated non-methylated Flag–HA–PP2Ac (20 nM) was first labelled in the assay buffer (20 mM HEPES pH 7.5, 200 mM NaCl, 1 mM TCEP pH 7.5, 0.1% BGG) at 4 nM Europium-anti-Flag donor fluorophore (PerkinElmer, AD0274). Concurrently, 4× concentrated His-tagged PP2Aa was labelled at 200 nM anti-His-d2 acceptor fluorophore in the presence of 200 nM untagged B56. Both donor and acceptor mixes were combined in equal volumes and allowed to incubate for an additional 2 h. CCDC6 was serially diluted at 2× concentrations (concentration range, 1,000–0 nM), and 4 µl of the assembled holoenzyme and CCDC6 was added to wells in the assay plate ProxiPlate−384 F Plus assay plates (Revvity, 6008260). Data were collected using the PheraStar plate reader (BMG Labtech). The 3 h timepoint is presented in the figure.
SPR
SPR experiments in this paper were recorded on a Biacore 8K instrument. Protein was captured at 20 °C with data collection at 5 °C. A Xantec SAHC 200M (Streptavidin) chip was inserted into the instrument and primed into running buffer (50 mM HEPES pH 7.4, 150 mM NaCl, 0.002% Tween-20, 0.25 mM TCEP and 2% DMSO). Avi-tag-biotinylated FBXO42 protein in a complex with SKP1 was prepared at 400 nM in capture buffer (50 mM HEPES pH 6.5, 150 mM NaCl, 0.002% Tween-20, 0.25 mM TCEP and 2% DMSO) and captured on FC2 to 3,500–4,000 RU. Affinity was measured using a titration series, injected from low to high with a six-point, twofold dilution dose–response with a top concentration of 50 µM in multicycle kinetics format with a flow rate of 50 μl min−1, and a contact and dissociation time of 30 s and 300 s, respectively. All data were analysed using the Insight software (Cytiva) using a 1-to-1 binding affinity model.
Statistics and reproducibility
Representative WB experiments
The numbers (n) of independent repeats for Figs. 1g,h, 2c,d, 3g, 4d,e,i and 5g,i,j and Extended Data Figs. 3d, 8e and 10h were n ≧ 3. The numbers (n) of independent repeats for Figs. 1a,i,j, 2b,e and 4h,j and Extended Data Figs. 2e, 3a,b,e, 4e and 10b were n ≧ 5.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.