SAMOSA from E14 mouse ES cells
We compiled a large compendium of SAMOSA data from wild-type E14 mouse ES cells, comprising both newly generated datasets and published datasets from our laboratory, including: (1) our study on the function of ISWI-family remodelling complexes3; (2) samples prepared with our SMRT-Tag protocol, which uses a transposase-based strategy for PacBio library preparation from lower input amounts of footprinted DNA38; and (3) mouse ES cells labelled with the halogenated thymidine analogue BrdU for varying time points in our study of single-fibre accessibility patterns in newly replicated chromatin52. Only samples with a mononucleosome–dinucleosome ratio (MDR; calculation detailed in ‘Data analysis and visualization’) greater than 10 were included for analysis. All cell lines were routinely tested for mycoplasma contamination. No statistical models were used to predetermine sample size. Neither blinding nor randomization was used.
Cell culture
For newly generated libraries, E14 mouse ES cells were grown on gelatin-coated (0.1% solution in 1× PBS) tissue culture plates. Cells were cultured in mouse ES cell medium (DMEM with GlutaMAX (Thermo Fisher Scientific, 10566-016)), supplemented with 15% fetal bovine serum (FBS; Thermo Fisher Scientific, SH30071.03), 14.2 mM 2-mercaptoethanol (Bio-Rad, 1610710), 1× NEAA (Thermo Fisher Scientific, 11140-50), 1 mM sodium pyruvate (Thermo Fisher Scientific, 11360-070) and 1× LIF (purified by the laboratory of B. Panning). Some samples were cultured with MEK (1 µM PD0325901 (Apex Bio/Fischer, A3013-25)) and GSK3β (3 µM CHIR99021 (Apex Bio/Fischer, A3011-100)) inhibitors (2i). The medium was changed daily and cells were passaged with 1× TrypLE (Thermo Fisher Scientific, 12605010) when confluent. To collect mouse ES cells, cells were rinsed with 1× PBS (Thermo Fisher Scientific, 10010023), dissociated with 1× TrypLE, quenched with mouse ES cell medium and centrifuged at 500g for 5 min. All mouse ES cell lines used in this study were regularly tested for mycoplasma contamination (Lonza LT07-318).
MTase footprinting and library preparation
Live cell counts were estimated (Countess 3) to ensure equal numbers of nuclei per unit MTase were used in each footprinting reaction. Nuclei were isolated by incubating 1 × 106 live mouse ES cells in 1 ml ice-cold NE1 buffer (20 mM HEPES, 10 mM KCl, 1 mM MgCl2, 0.1% Triton X-100, 20% glycerol and 1× protease inhibitor (Roche, 4693132001)) on ice for 5 min. Nuclei were pelleted by centrifugation at 500g for 5 min at 4 °C. Nuclei were rinsed in 1 ml buffer M (15 mM Tris-HCl pH 8.0, 15 mM NaCl, 60 mM KCl and 0.5 mM spermidine) and re-pelleted by centrifugation at 500g for 5 min at 4 °C. For footprinting, nuclei were resuspended in 200 µl buffer M with 1 mM SAM (NEB, B9003S) and 10 µl high-concentration EcoGII (custom order from NEB). Nuclei were incubated at 37 °C for 30 min, with a spike-in of 1 µl 32 mM SAM stock halfway through the footprinting reaction. For light MNase digest to liberate chromatin fragments, footprinted nuclei were pelleted by centrifugation at 500g for 5 min at 4 °C, resuspended in 200 µl buffer M with 1 mM CaCl2 and 0.02 U MNase (Sigma N3755) and incubated on ice for 30 min. To quench the MNase reaction, 2 mM EGTA was added.
To purify DNA, samples were treated with 10 µl RNaseA (Thermo Fisher Scientific, AM2270) for 10 min at 37 °C, followed by the addition of 2 µl 10% SDS and 2 µl 20 mg ml−1 proteinase K (Thermo Fisher Scientific, AM2548) for more than two hours (up to overnight) at 65 °C. To extract DNA, an equal volume (around 250 µl) phenol-chloroform-isoamyl alcohol was added, and samples were mixed by brief vortexing and centrifuged at maximum speed for 2 min. The aqueous phase was transferred to a fresh tube along with 0.1 volumes 3 M NaOAc, 2 volumes ice-cold 100% ethanol and 1 µl GlycoBlue co-precipitant (Thermo Fisher Scientific, AM9516). Samples were mixed by inverting tubes and incubated overnight at −20 °C, centrifuged at maximum speed for 30 min at 4 °C, washed in 500 µl ice-cold 70% ethanol, air dried and resuspended in 30 µl buffer EB. DNA concentrations were measured using the Qubit dsDNA High Sensitivity Quantification Kit (Thermo Fisher Scientific, Q32851). Up to 1 µg DNA was prepared into PacBio SMRT libraries with the SMRTbell Express Template Prep Kit 2.0 as per the manufacturer’s directions. Fragment size distributions were estimated by Femto Pulse (Agilent). Libraries were sequenced on PacBio Sequel II 8M SMRT cells.
SAMOSA from SOX2-FKBP mouse ES cells
Degron-tagged Sox2FKBP-F36V (hereafter referred to as SOX2-FKBP) mouse ES cells were provided by the laboratory of E. de Wit60. SOX2-FKBP mouse ES cells were cultured in the same mouse ES cell medium described above (including 2i) and were plated at a density of 5 × 105 per well in 6-well plates 24 h before collection. For degradation, SOX2-FKBP mouse ES cells were treated with 0.5 µM dTAGV-1 (Tocris 6914) for varying time points (30 min, 2 h or 6 h). As a control, SOX2-FKBP mouse ES cell samples were treated with 0.5 µM dTAGV-1-NEG (Tocris 6915) for six hours before collection. All experiments were performed in biological duplicate. Footprinting and downstream steps were performed as detailed in the section above, except that instead of the MNase digest step, we sheared purified genomic DNA with Covaris g-TUBEs (Covaris, 520079; 4,600g for six passes).
Western blot for SOX2 protein in nuclear extracts
To validate levels of SOX2 knockdown, mouse ES cell nuclei were isolated in NE1 buffer, as was done for the matched MTase-footprinted samples. Pelleted nuclei were resuspended in 100 µl ice-cold RIPA buffer (150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS, 50 mM Tris pH 7.4 and 1× protease inhibitor (Roche 4693132001)). Samples were incubated for 30 min on ice and sonicated briefly to solubilize chromatin. Protein extracts were clarified by centrifugation at 12,000g for 20 min at 4 °C. Soluble material was collected, protein concentrations were measured by bicinchoninic acid assay (Thermo Fisher Scientific, 23227), and samples were diluted in NuPAGE LDS sample buffer (Thermo Fisher Scientific, NP0007) with 5% β-mercaptoethanol. Samples were reduced by boiling for 10 min at 95 °C, resolved on 4–12% Bis-Tris gels (Thermo Fisher Scientific, NP0322), and transferred to PVDF membranes. Membranes were incubated overnight with the following primary antibodies: anti-SOX2 (CST 23064S; 1:1,000 dilution) and anti-vinculin (Sigma V9264; 1:1,000 dilution; used as loading control). After washing, membranes were incubated with secondary antibodies conjugated to IRDye 700 or IRDye 800 (1:10,000 dilution) and imaged with the LiCor Odyssey.
SAMOSA from CTCF-AID mouse ES cells
CTCF-AID mouse ES cells (EN52.9.1) were generated by the E.N. laboratory76. Raw data from CTCF-AID samples used in this study were previously published by our group52, and samples were required to have a MDR (calculation detailed in ‘Data analysis and visualization’) greater than 10 to be included in analyses. These criteria resulted in: (1) n = 10 biological replicates of auxin-treated (for six hours) CTCF-AID mouse ES cells; and (2) n = 6 biological replicates of ddH2O-treated CTCF-AID mouse ES cells (as controls).
SAMOSA from NIPBL-FKBP mouse ES cells
NIPBL-FKBP mouse ES cells (EA18.1) were cultured in the same conditions as E14 mouse ES cells and NIPBL was depleted using 500 nM PROTAC dTAG-13 (Sigma-Aldrich, SML2601), as described previously55,56. EA18.1 cells were treated with dimethyl sulfoxide (DMSO) for 24 h, dTAG-13 for 6 h (preceded by DMSO) or dTAG-13 for 24 h, with n = 2 biological replicates per condition. Cells were also pulsed with BrdU for one hour before collection. SAMOSA was done as for wild-type E14 mouse ES cells, with no MNase treatment; instead, DNA was sheared using g-TUBEs (Covaris 520079) to a target length of 7–8 kb. Replicates were pooled and the 6-h and 24-h time points were combined for treatment conditions to cover the full range of phenotypes that are relevant to perturbing loop extrusion.
SAMOSA from primary mouse hepatocytes
Mouse husbandry and genotyping
All animal experiments were performed under the supervision and approval of the Institutional Animal Care and Use Committee (IACUC) at the University of California, San Francisco (UCSF) (protocol AN179718-03F). Six-week-old, C57BL/6J mice (Jackson Laboratory, 000664) were used as wild-type hepatocyte samples in our dataset (n = 3 biological replicates). Foxa2∆Hx-TagRFP-expressing mice (hereafter referred to as FOXA2-ΔHx) were provided by the laboratory of K. Zaret63. Samples in this study included heterozygous FOXA2-ΔHx mice (n = 3; one male and two females) and a littermate wild-type control male mouse.
Genotyping for the presence (around 230 bp) or absence (around 200 bp) of the associated Tag-RFP fluorophore was performed with the following primer pair:
lox2272_F: AGTGTTGTCTTCTGCCTTTGAG
lox2272_R: GCTTACCTTAGTCTCGGTCTTGG
Genotyping for the presence (around 300 bp) or absence (around 270 bp) of the helical domain in FOXA2 was performed with the following primer pair:
TagRFP_F: GCTCTTCGCCCTTAGACACC
TagRFP_R: ATCAGCCCCACAAAATGGAC
Isolation of primary mouse hepatocytes
Hepatocyte isolation was performed by the Liver Cell Isolation, Analysis and Immunology Core at the UCSF Liver Center. After flushing the liver free of blood, the organ was perfused for 3 min at 4 ml per min with Liver Perfusion Medium (Thermo Fisher Scientific, 17701038), followed by an additional 7 min at 4 ml per min with Liver Digest Medium (Thermo Fisher Scientific, 17703034). The liver was then removed, diced with scissors and suspended in DME-H21 with insulin-transferrin-selenium (Thermo Fisher Scientific, 41400045), penicillin–streptomycin and 5% FBS. The crude suspension was strained through a 100-µm filter and pelleted twice at 60g with interval washing. Hepatocytes were purified by suspension of the pellet and centrifugation through 45% Percoll (Cytiva 17089102) at 130g for 15 min. Purified hepatocytes, which pelleted through the Percoll, were suspended in DME-H21 as above. Final hepatocyte suspensions had a viability higher than 90%.
MTase footprinting for hepatocyte samples
Footprinting reaction for samples denoted HepWT_Rep1 and HepWT_Rep2 was performed in 1 × 106 purified hepatocyte nuclei per library. Footprinting and downstream steps were performed as described above, except that primary hepatocytes were first dounced in NE1 buffer (10× with loose pestle and 10× with tight pestle) to facilitate nuclear isolation.
The footprinting reaction for all remaining hepatocyte samples was performed with 1 × 106 digitonin-permeabilized hepatocytes. Footprinting and downstream steps were performed as described above, with two notable exceptions. First, instead of isolating nuclei with NE1 buffer, we directly permeabilized cells by adding 0.05% digitonin (Thermo Fisher Scientific, BN2006) to buffer M when performing the EcoGII footprinting reaction. Second, instead of treating nuclei with MNase to liberate chromatin fragments, we sonicated purified genomic DNA with the Megaruptor 3 (concentration, 10 ng µl−1; volume, 100 µl; speed, 031) to achieve more consistently sized and longer molecules (>10 kb).
SAMOSA from H1 TKO mouse ES cells
ES cell lines derived from H1c/H1d/H1e TKO and wild-type littermates were expanded on mitotically inactivated mouse embryonic fibroblasts (MEF) feeder layers as we described previously77. Before extraction of nuclei, MEF feeder cells were depleted from ES cell culture and H1 TKO and wild-type ES cells were subsequently cultured feeder-free on 0.1% gelatin-coated tissue culture plates with 2i/LIF medium for three passages following protocols as described78. SAMOSA was performed on 1 × 106 nuclei per replicate, as with E14 mouse ES cells, without MNase treatment. DNA was then purified and used directly for PacBio library preparation and sequencing.
SAMOSA from iPS cells and differentiated endoderm
The normal human male iPS cell line GM25256 (WTC, hPSCreg: UCSFi001-A, generated at Gladstone Institutes, distributed by the Coriell Institute for Medical Research) was used for endoderm differentiation79. Undifferentiated iPS cells were cultured in mTeSR Plus medium (100-0276, STEMCELL Technologies) in six-well plates coated with 0.25 mg ml−1 Matrigel GFR basement membrane matrix (356231, Corning) at 37 °C in a humidified incubator with 5% CO2 and 5% O2. Cells were passaged every three days using Versene (A4239101, Gibco) and replated in mTeSR Plus medium supplemented with 10 µM Y-27632 (101763-964, Selleck Chemicals). Mycoplasma testing was performed every six months during routine culture using the MycoAlert Mycoplasma Detection Kit (LT07-218, Lonza).
Endoderm was differentiated from iPS cells using a modified version of a previously described protocol80. In brief, iPS cells were dissociated with Accutase (07920, STEMCELL Technologies) at 70–80% confluency and replated at a density of 75,000 cells per cm2 on Matrigel-coated six-well plates in mTeSR Plus medium including 10 µM Y-27632. After 18 h, cells were exposed to endoderm-induction medium, consisting of RPMI 1640 (11875093, Gibco) supplemented with 2% B-27 supplement minus insulin (A1895601, Gibco), 1% Glutamax (35050061, Gibco), 1% non-essential amino acid solution (NEAA; 11140050, Gibco), 0.5 mM sodium butyrate (B5887, Sigma-Aldrich) and 100 ng ml−1 activin A (120-14E, PeproTech). Cells were cultured at 20% O2 for six days. During the first three days of differentiation, the following compounds were added to endoderm-induction medium: 3 µM CHIR99021 (501742210, Sigma-Aldrich) on day 1; 10 ng ml−1 BMP4 (120-05ET, PeproTech) and 20 ng ml−1 basic fibroblast growth factor (FGFb; 100-18B, PeproTech) on day 1 and day 2; 50 nM PI103 (501932056, Thermo Fisher Scientific) from day 1 to day 3; and knockout serum replacement (KSR; 10828010, Gibco) at the following concentrations: 2% on day 1, 1% on day 2 and 0.5% on day 3. The medium was changed every day during differentiation and maintenance.
SAMOSA was performed on approximately 1 × 106 digitonin-permeabilized nuclei per replicate, for both iPS cells and endoderm. DNA was purified then sheared using the Megaruptor 3 to a target length of 15 kb before PacBio library preparation and sequencing.
SAMOSA-ChAAT on reconstituted Widom arrays
Data from HMGB1 and linker histone H1 Widom 601 array SAMOSA-ChAAT experiments performed in and described in detail in a previous study46 were reprocessed using the IDLI pipeline as below.
Preprocessing PacBio data for single-molecule accessibility
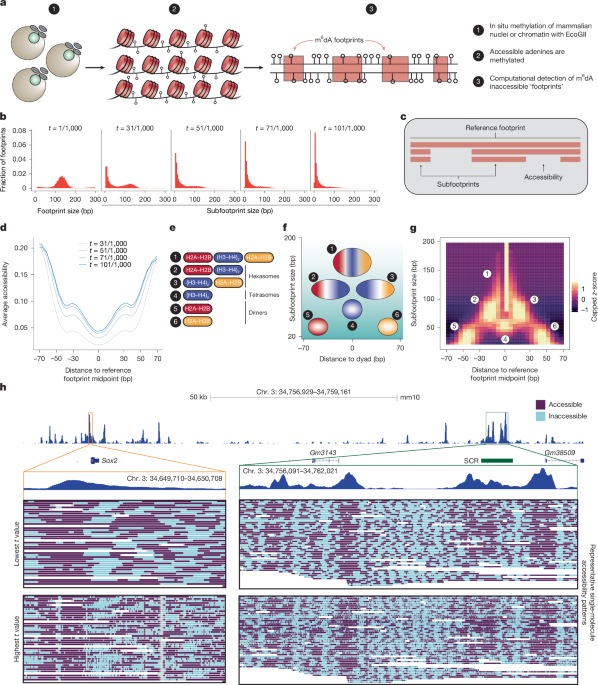
To preprocess PacBio data, we used software from Pacific Biosciences and a custom script (hmm_output_t_values.py) to successively run the NN-HMM across a range of user-specified transition probabilities. These resulted in the following output per sample: (1) an alignment of CCS reads to the mouse reference genome (mm10); (2) an accessibility prediction per CCS molecule (Viterbi path of HMM component of model); and (3) an atlas of footprint locations (start/end positions) and sizes on a per-molecule basis. Files for (2) and (3) were generated for t = 1/1,000, 31/1,000, 51/1,000, 71/1,000 and 101/1,000 for each sequencing library.
Data analysis and visualization
Quantifying the extent of MTase footprinting per sequencing library
We used the ratio of mononucleosome-sized to dinucleosome-sized footprints (MDR) as a proxy for measuring the extent of EcoGII footprinting on a per-sample basis. Poorly methylated samples exhibit a higher fraction of footprints longer than the expected length of DNA protected by a mononucleosome (lower MDR values), which can be ascribed to the failure of EcoGII to efficiently methylate in linker DNA. We used a custom script (compute_mdr_per_sample.py), which uses the find_peaks function from scipy and computes the ratio of maximum peak heights in footprint length histograms for mononucleosome- and dinucleosome-sized footprints per sequencing library.
Visualizing single-molecule data across different t values at single genomic loci
To visualize SAMOSA data at loci of interest, data were processed with a custom script (generate_bam_for_igv.py) to encode single-molecule accessibility patterns (for any given t value) as an additional flag in aligned BAM files. These BAM files were concatenated across E14 mouse ES cell samples with MDR > 10 and visualized at the Sox2 locus (chr. 3: 34,756,929–34,759,161; including the Sox2 promoter, gene body and downstream SCR) using a custom version of IGV (https://github.com/RamaniLab/SMRT-Tag/tree/main/igv-vis). Bulk ATAC-seq data from mouse ES cells were obtained as processed BW files (GSE98390) and visualized at identical coordinates on the UCSC Genome Browser.
Identifying fibres that overlap annotated epigenomic domains in mouse ES cells
A custom script from our group (https://github.com/RamaniLab/SAMOSA-ChAAT/blob/main/scripts/zmw_selector.py) was adapted to identify sequencing reads in which a portion of the read falls within ±1 kb of midpoints of histone post-translational modification-defined epigenomic domains in mouse ES cells (H3K4me1, H3K4me3, H3K9me3, H3K27me3, and H3K36me3 ChIP–seq peaks; derived from ENCODE data).
Identifying fibres that overlap with repeat elements in the mouse genome
A custom script from our group (https://github.com/RamaniLab/SAMOSA-ChAAT/blob/main/scripts/blast_bam.sh) was used to identify sequencing reads with one or more matches to repeat elements of interest. In brief, we ran BLAST on CCS reads from a database consisting of mouse major satellite sequence (M32564.1), minor satellite sequence (X14462.1) and telomeric DNA sequence (pentameric repeats of TTAGGG). For fibres with multiple repeat matches, only the repeat sequence with the lowest E-value was considered for downstream analyses.
Identifying fibres that overlap with bound and unbound TF motif matches
To define instances of TF-bound motifs, we referenced the following datasets: ChIP-nexus data for SOX2 in mouse ES cells (GSE137193)59, ChIP–seq data for CTCF in mouse ES cells (ENCFF508CKL) and ChIP–seq data for FOXA2 in mouse hepatocytes (GSE157452)64. For SOX2 motifs, we retained only the subset of peaks that contained a SOX2-OCT4 composite motif match as defined by this dataset81. For CTCF data, we used a curated set of ChIP–seq backed motifs first processed in this study82. For FOXA2 data, we retained only the subset of peaks that contained a position weight matrix (PWM) match (MA0047.2) for the FOXA2 motif83. For control sites, we randomly sampled an equal number of motif matches per factor of interest from a previously published list of motifs across the mouse genome82, requiring that these motifs fall at least 1 kb away from the nearest ChIP-annotated peak. As above with epigenomic domains, we used a custom script to identify sequencing reads in which a portion of the read falls within ±1 kb of motif midpoints.
Clustering single-molecule accessibility patterns with respect to footprint midpoints
To examine patterns of nucleosomal distortion, we clustered accessibility patterns across a range of t values with respect to footprint midpoints. Footprints less than 200 nt in length (intended to capture both nucleosomal and subnucleosomal species) within ±500 bp of midpoints of epigenomic domains, repeat sequences or TF-binding motifs were selected. On a per-footprint basis, accessibility data (from t = 31/1,000, 51/1,000 and 71/1,000) for a 140-bp window centred on each footprint midpoint (420 bp in total per footprint) were used as input for Leiden clustering (res = 0.5, 0.6, 0.6 and 0.6 for domains/repeats, SOX2, CTCF and FOXA2 motifs, respectively). Clusters were filtered such that the least abundant clusters that collectively summed up to 10% of each dataset were removed. It should be noted that: (1) accessibility data were oriented accounting for sequence feature strand before clustering to preserve any potential directional effects associated with non-palindromic TF-binding motifs; (2) this approach was intended to enable clustering on accessibility information across multiple t values per footprint; and (3) this window size was selected to prioritize signal from SHL −7 to SHL +7 for nucleosomal footprints. Representative code for performing this analysis is provided as a custom script (process_footprints_nucleosomal_distortion.py).
Plotting footprint size distributions, mean accessibility patterns and horizon plots per cluster
To assign putative biological functions to clusters of nucleosomal distortion patterns or translational positions, we visualized the following features. First, we plotted the distribution of footprint sizes on a per-cluster basis for data from the lowest t value examined (t = 1/1,000). Second, we plotted mean accessibility as a function of distance to respective footprint/sequence feature midpoints from the highest t value used for Leiden clustering (t = 71/1,000). Third, we generated horizon plots (analogous to commonly used V-plots in MNase-seq data) by computing z-scores for the enrichment of footprints of a particular size (20–200 nt) at specific positions (±70 bp from footprint/sequence feature midpoints) for each individual cluster. z-scores are summed per size–position combination across the range of t values previously used for clustering (t = 31/1,000, 51/1,000 and 71/1,000). Colour scale limits for the resulting heat map are capped by the 0th (lower) and 95th (upper) percentile z-scores for positions within ±70 bp of the footprint/sequence feature midpoint. Clusters of entirely unmethylated footprints were manually filtered out before downstream quantification. Representative code for performing this analysis is provided as a custom script (plot_footlen_meanacc_horizondata.py).
Identifying groups of clusters with shared nucleosomal distortion patterns or translational positions
To group Leiden assignments for nucleosomal distortion patterns or translational positions in an unbiased manner, we performed hierarchical clustering of horizon plot data as follows. z-scores per cluster were filtered such that only those corresponding to footprints less than 100 nt and less than 50 nt in length were retained for defining nucleosomal distortion and translational position groups, respectively. This choice was intended to prioritize the clustering of horizon plot data specifically within subfootprint size ranges that capture subnucleosomal organization. Data were manually inspected and z-scores flipped with respect to footprint/sequence feature midpoints to account for the nucleosomal dyad as an axis of symmetry. The distance matrix was computed with the dist function in R, hierarchical clustering was performed with hclust (from the factoextra package; method = ward.D2) and output was visualized as a dendrogram (with fviz_dend). In most cases, the group definitions are driven by the dendrogram. In a subset of cases, we found that footprint size drove the clustering, but patterns were visually most similar to alternative groups. In those cases, we manually assigned a label.
UMAP visualization of footprint types within epigenomic domains and at repeat sequences
From accessibility data for footprints within histone-modification-defined domains and at mouse repeat elements, we used Scanpy (v.1.9.3) for principal component analysis (PCA)-based dimensionality reduction, construction of a k-nearest neighbours graph (metric = correlation, n_neighbours = 15) and UMAP visualization (min_dist = 0.5). Clusters were coloured according to Leiden assignments and proportions of each Leiden-defined cluster were represented as a pie chart surrounding the UMAP visualization.
Structural visualization of nucleosomal accessibility patterns
For structural visualization of mean accessibility patterns from defined clusters, we used a custom script (accessibility_to_pdb_structure.py) to convert our mean single-molecule accessibility data per cluster of interest from the highest t value used for Leiden clustering (t = 71/1,000) into a format compatible with visualization via ChimeraX (v.1.7.1; attribute = percentAcc, match mode = 1-to-1 and recipient = residues)84. Accessibility values were overlaid on Protein Data Bank (PDB) structure 7KBD (ref. 44).
Computing the enrichment of nucleosomal distortion patterns and translational positions across distinct genomic loci
To compute ORs for Leiden-defined clusters across epigenomic domains and repeat elements, or for bound versus randomly sampled TF motifs, Fisher’s exact tests were done with scipy (in Python). All computed P values were corrected with a Storey q-value correction, using the qvalue package in R.
Computing effect sizes associated with TF knockdown or across genotypes
To compute effect sizes after degron-mediated TF knockdown or across mouse genotypes, Fisher’s exact tests were performed for footprints specifically at TF-bound motifs (for example, footprints from dTAG-V1 versus DMSO-treated SOX2-FKBP mouse ES cells) with scipy (in Python). All computed P values were corrected with a Storey q-value correction, using the qvalue package in R.
Reproducibility of nucleosomal distortion patterns or translational positions across replicates
To assess reproducibility in our large-scale E14 mouse ES cell dataset, we asked whether the rank order of clusters of nucleosomal distortion patterns or translational positions is preserved across sequencing libraries. To do this, the relative proportion of each Leiden-defined cluster at domains/repeats or at TF-bound motifs was computed on a per-sample basis and analysed using Spearman’s correlation (with spearmanr from scipy). Footprints from all sequencing libraries with MDR > 10 were included in Leiden clustering of accessibility patterns; this choice was intended to include as many footprints as possible to increase statistical power. However, for these correlation analyses, we imposed a minimum sequencing depth for a biological replicate to be included, with the goal of assessing reproducibility in sufficiently sequenced samples (minimum number of footprints indicated on each respective plot).
To evaluate reproducibility in SOX2-FKBP mouse ES cells, CTCF-AID mouse ES cells and FOXA2-ΔHx hepatocytes, we calculated the Pearson’s correlation (with pearsonr from scipy) for effect sizes computed on a per-sample basis. This choice was intended to capture potential quantitative changes in cluster proportions between degron conditions and genotypes across replicates.
Identifying single-molecule co-occupancy patterns for footprint groups
Individual fibres with at least three Leiden-classified footprints within ±500 bp of repeat elements or TF motifs of interest were selected. Fibres with footprints greater than 200 nt and/or with footprints in the least abundant 10% of clusters were excluded from further analysis. Consecutive ‘triplet’ footprints were tabulated by computing midpoints and lengths for: (1) the central-most footprint nearest the repeat element or TF-binding motif; (2) the most proximal footprint upstream of the central-most footprint; and (3) the most proximal footprint downstream of the central-most footprint. Per-molecule accessibility data were computed for a 2-kb window centred on the repeat element or TF-binding motif of interest for visualization alongside footprint positions, accounting for sequence feature strand appropriately. Representative code for performing this analysis is provided (process_triplet_footprints.py).
Computing the enrichment of consecutive triplets on single chromatin fibres
Individual clusters were combined on the basis of groups defined by shared subfootprint organization from hierarchical clustering, as described above. Expected frequencies of footprint ‘triplets’ were computed as the product of the independent probabilities of observing each group within a given dataset. These values were used to calculate log2(observed (O)/expected (E)) ratios for each triplet across epigenomic domains, repeat elements and TF-binding motifs. Enrichment was assessed using Fisher’s exact test (fisher.test function in R), with false discovery rate (FDR) correction applied for multiple comparisons. The results were visualized as a ranked-order scatter plot, reflecting the effect size for each triplet.
Visualizing mean accessibility patterns at bound versus random TF motifs
Previously published scripts from our group were used to compute and cluster mean accessibility observed within ±750 bp of TF-binding motifs across different conditions (before and after TF knockdown; across genotypes)3. Euclidian distances for the resultant ORs for bound versus unbound motifs per sample are plotted to visualize their similarity across conditions and replicates.
Motif enrichment and classification
For each nucleosome, ±75 bp from the footprint centre was used as the input sequence for motif analyses. For each class of nucleosome, SEA51 was used to find enriched motifs within that nucleosome class, using the sequences from the total other nucleosomes as background. SEA was performed with default parameters using the JASPAR 2024 core non-redundant vertebrate motif database85. The resulting motifs were then assigned a class based on their JASPAR classification; classes with no representation across all nucleosome types were omitted. CentriMo analysis was performed using default parameters and significant and representative motifs were chosen to display86.
Quantitative reproducibility and motivation for the architecture of the IDLI pipeline
Quantitative reproducibility information for all SAMOSA experiments and IDLI analyses presented in this paper is provided in Supplementary Figs. 1–8. Supplementary Fig. 1 visualizes the reproducibility of IDLI-based horizon plots from our various E14 SAMOSA replicate experiments, and quantitative estimates of reproducibility across replicates and within samples through bootstrapping. Supplementary Fig. 2 describes benchmarking analyses to ascertain the quality of CTCF-AID degron experiments, including visualizations of average single-molecule accessibility at CTCF sites, replicate reproducibility and examples of correlations between degradation and changes in the abundance of distinct single-molecule states, as previously described by our group3,38,52. Supplementary Fig. 3 is akin to Supplementary Fig. 2 but for SOX2-FKBP line experiments. Supplementary Fig. 4 is akin to Supplementary Fig. 2 but for FOXA2 perturbation experiments in hepatocytes. Supplementary Fig. 5 collates all dendrograms resulting from hierarchical clustering of nucleosome types to define groups. All dendrograms are labelled according to where nucleosome footprints were sampled from across the genome, and which sample was used for IDLI analysis. Supplementary Figs. 6 and 7 describe correlations between single-molecule accessibility patterns and nucleosome types, using CTCF (Supplementary Fig. 6) and SOX2 (Supplementary Fig. 7) as examples. This is provided as a reference to the reader to clarify that there are quantitative links between the overall accessibility state of the molecule (nucleosome-occluded motif versus open) and the abundance of distinct nucleosome types on those molecules. Supplementary Fig. 8 captures the quantitative reproducibility of all nucleosome-type definitions across each experiment presented in the manuscript.
We also include a Supplementary Note with the manuscript. This describes the usability of the IDLI approach and also motivates the parameter choices and computational design of the NN-HMM that underlies the IDLI pipeline.
Finally, all library metadata for sequencing experiments performed and analysed for this study are provided in Supplementary Table 1.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.