Unless otherwise noted, chemicals and buffer components were purchased from Sigma-Aldrich.
Cell culture
PtK2 cells (ATCC, CCL-56) were cultured in MEM (Gibco) supplemented with 10% fetal bovine serum (FBS), 1 mM sodium pyruvate, 1× non-essential amino acids (Gibco), and 1× antibiotic–antimycotic (Gibco) at 37 °C in 5% CO2. All cell lines were obtained from ATCC, which validates lines by STR profiling. We did not further validate them after purchase. Cells were monitored for mycoplasma contamination with the ATCC universal mycoplasma detection kit (a PCR-based assay) and tested negative. No commonly misidentified cell lines were used in this study.
Cell line generation
For generation of a doxycycline-inducible F-tractin–mScarlet and zyxin–LDO–mNeongreen PtK2 cell line, HEK293T (ATCC, CRL-3216) cells were co-transfected with lentiviral plasmids and packing plasmids (PMD2.G and PsPAX2) using Lipofectamine 2000 (Thermo Fisher Scientific). Six hours post-transfection, the culture medium was refreshed. Forty-eight hours post-transfection, the culture medium containing lentivirus was collected and filtered. Lentivirus infection with polybrene (5 μg ml−1) followed immediately after collection. Three types of lentivirus were added to PtK2 cells: F-tractin–mScarlet, zyxin–LDO–mNeongreen, and TetOn. Twenty-four hours post-infection, cells that stably integrated the lentiviral plasmids were selected with puromycin and blasticidin for one week.
Cellular cryo-ET grid preparation
C-flat 1.2/1.3 holey carbon Au 200-mesh grids (Electron Microscopy Sciences) were sputter coated with 22 nm of carbon and baked in a dry oven at 60 °C overnight. Grids were plasma cleaned and coated with 10 μg ml−1 fibronectin (Sigma, FC010) in DPBS for 1–3 h at 37 °C in a tissue culture incubator. Fibronectin-coated grids were washed with DPBS and stored in fresh PtK2 culture medium in a 35 mm tissue culture dish for immediate use.
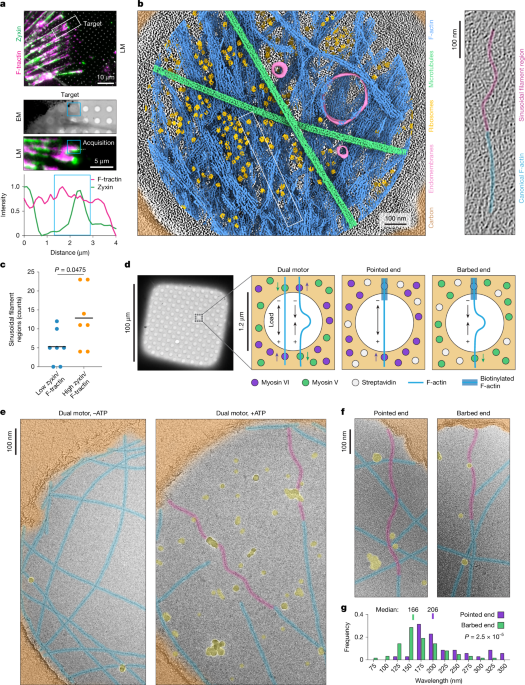
Doxycycline-inducible F-tractin–mScarlet and zyxin–LDO–mNeongreen PtK2 cells were grown to 90% confluency before passaging. Cells were thoroughly trypsinized and filtered through a 0.45-μm filter to remove clumps. Thirty thousand cells were added to each 35 mm tissue culture dish containing the fibronectin-coated grids. Twenty-four hours later, the medium was replaced with fresh PtK2 medium containing 100 ng ml−1 of doxycycline to induce expression of F-tractin–mScarlet and zyxin–LDO–mNeongreen. The next day, grids were prepared for plunge freezing. Grids for the majority of tomograms (presented in Fig. 1b and Extended Data Fig. 1c,d) were treated with 1 µg ml−1 Rho Activator II (Cytoskeleton, CN03) for 3.5 h, while the grid for the tomogram presented in Extended Data Fig. 1b (tomogram 11 in Extended Data Fig. 1d) was untreated. The grids were then washed with DPBS and loaded onto a Leica EM-GP plunge freezer operating at 25 °C. Three microlitres of DPBS was added to the front of the grid, followed by blotting from the back using a Whatman no. 5 filter paper for 7 s, then flash frozen in liquid ethane.
Cryo-fluorescence imaging
Grids were mounted on a Leica Cryo-CLEM microscope. Epifluorescence microscopy was performed using a 50× 0.9 NA air objective at −180 °C. z-Stacks of images were captured at a depth of 16 bit. Image acquisition was performed using LAS X software (Leica). Images were post-processed with THUNDER (Leica) to denoise, then maximum intensity projected.
Cellular cryo-ET data collection
Data were collected on a spherical-aberration (Cs) corrected Titan Krios TEM (Thermo Fisher Scientific) operating at 300 kV and equipped with a K3 direct electron detector (Gatan) and a BioQuantum energy filter (Gatan) using a slit width of 20 eV. Micrographs were collected at a nominal magnification of 15,000×, which corresponds to a calibrated pixel size of 5.05 Å (2.525 Å in super-resolution mode). The tilt-series were collected using the FastTomo acquisition scheme69 in SerialEM70 using a target defocus of −5 μm underfocus. Tilt angles ranged from −54° to +54° in 3° increments, grouped in a dose-symmetric manner71. Each tilt in the tilt-series had a total acquisition time of 2.4 s and total dose of 2.82 e− Å−2, fractionated across 16 frames (0.18 e− Å−2 per frame).
Cellular cryo-ET data processing
Individual tilts were motion-corrected and binned 2× (to a pixel size of 5.05 Å) using MotionCor2 (ref. 72), and contrast transfer function (CTF) parameters were estimated using CTFFIND4 (ref. 73). Tilt-series alignment was carried out using AreTomo2 (ref. 74), followed by reconstruction using IMOD75 with back-projection at a binning of 2 (voxel size of 10.1 Å). The reconstructed tomograms were denoised using cryoCARE76, then the missing wedge was predicted and restored with IsoNet77. The tomograms were then segmented using a U-Net convolutional neural network with the Dragonfly software78. For the tomogram displayed in Extended Data Fig. 1b, the plasma membrane was segmented using MemBrain v2 (ref. 79). All membrane segmentations were smoothed using mean curvature motion80. Segmentations were visualized using ChimeraX81.
Cellular cryo-ET quantification
Cellular tomogram quantification was performed in a blinded fashion. D.Y.Z.P. manually assigned tomograms into low zyxin/F-actin and high zyxin/F-actin bins based on the absence or presence of local peaks of zyxin signal intensity relative to F-actin intensity in line scans of cryo-fluorescence micrographs (Extended Data Fig. 1d). Tomograms were then randomized before quantification by K.H. and G.M.A. Sinusoidal filament regions were manually annotated using 3DMOD based on the appearance of continuous stretches of oscillating curvature, followed by unblinding for analysis. With the exception of tomograms 2 and 10, both annotators consistently found similar numbers of sinusoidal filament regions across tomograms (Extended Data Fig. 1f). However, relatively few overlapping sinusoidal filament regions were selected. Therefore, after removing overlaps, the results for both annotators were combined for final quantification (presented in Fig. 1c). The total volume occupied by F-actin in each tomogram was estimated by Dragonfly segmentation as described above.
Actin purification
Actin was prepared from chicken skeletal muscle as previously described82. All steps were performed at 4 °C unless otherwise indicated. In brief, 1 g of acetone powder was resuspended in 20 ml of G-Ca buffer (G buffer: 2 mM Tris-HCl pH 8, 0.5 mM dithiothreitol (DTT), 0.2 mM ATP, and 0.01% NaN3, supplemented with 0.1 mM CaCl2), then mixed by inversion for 30 min. The mixture was ultracentrifuged in a Beckman Ti70 rotor at 42,500 rpm (79,766 g) for 30 min. The supernatant, containing G-actin monomers, was collected, and 50 mM KCl and 2 mM MgCl2 were added to induce F-actin polymerization for 1 h. 0.8 M KCl was then added and the mixture incubated for 30 min to dissociate contaminants from F-actin, followed by ultracentrifugation in a Ti70 rotor at 42,500 rpm (79,766 g) for 3 h. The pellet containing F-actin was collected and resuspended in 2 ml G-Ca buffer, then incubated overnight to depolymerize filaments. The mixture was then Dounce homogenized for 10-15 passes, sequentially sheared with 26 G and 30 G needles, then dialysed against G-Ca buffer overnight in Spectra/Por 1 dialysis tubing (6-8 kDa MWCO). The solution was then collected, sheared again with a 30 G needle, then dialysed against G-Ca buffer for an additional day. The solution, containing dissociated G-actin monomers, was collected and ultracentrifuged in a Beckman Ti90 rotor at 70,000 rpm (187,354 g) for 3 h. The upper two-thirds of the supernatant was then loaded on to a Cytiva HiLoad 16/600 column for size-exclusion chromatography. Purified G-actin was maintained in G-Ca buffer at 4 °C.
Protein expression and purification
Plasmids were propagated in NEB 5-alpha competent Escherichia coli cells and purified with either Qiaprep spin miniprep kits (Qiagen, for bacterial expression) or the PureYield plasmid maxiprep system (Promega, for transfection). Full-length human calmodulin was expressed in Rosetta2(DE3) E. coli cells and purified at 4 °C according to a published protocol83. The purified protein was collected in Storage Buffer (20 mM Tris-HCl pH 8.0, 100 mM NaCl, 2 mM β-mercaptoethanol, and 5% v/v glycerol), flash frozen in liquid nitrogen, and stored at −80 °C prior to use. For experiments presented in Fig. 1, Extended Data Fig. 2a–d, and Supplementary Videos 2–4, constructs for murine myosin Va (amino acids 1–1090) and human myosin VI (amino acids 1–1021) featuring C-terminal GFP and Flag tags were expressed and purified from Sf9 insect cells using the baculovirus system as previously described84. In the myosin VI construct, a GCN4 leucine zipper dimerization domain replaced the smooth muscle coiled-coil region originally used by Nishikawa et al.85. For all other experiments, equivalent constructs for human myosin Va (amino acids 1–1090) and myosin VI (amino acids 1–1021) were expressed from a modified pCAG vector featuring a C-terminal GFP and Flag tag in FreeStyle 293-F cells (ThermoFisher) and purified as described24.
In brief, cells were cultured in FreeStyle expression medium (ThermoFisher) at 37 °C on an orbital shaker in the presence of 8% CO2. Cells were transfected when the culture reached a density of 1.8 × 106 cells per ml. Per 400 ml of culture, 1.2 ml of 1 mg ml−1 PEI MAX (PolySciences) was pre-mixed with 400 µg of plasmid in 15 ml of FreeStyle expression medium and incubated for 20 min at room temperature prior to transfection. Myosin Va and myosin VI were co-transfected with a modified pCAG vector containing untagged, full-length human calmodulin (amino acids 1–149) at a mass ratio of 1:6. Cells were cultured for an additional 60 h, then collected and snap frozen in liquid nitrogen. Cell pellets were stored at −80 °C.
For myosin purification, all steps were performed at 4 °C. Cells were resuspended in myosin lysis buffer (50 mM Tris-HCl pH 8.0, 150 mM NaCl, 2 mM MgCl2, 0.2% 3-((3-cholamidopropyl) dimethylammonio)−1-propanesulfonate (CHAPS), 2 mM ATP, 1 mM phenylmethylsulfonyl fluoride (PMSF), 1 μg ml−1 aprotinin, leupeptin and pepstatin) and incubated with rocking for 40 min. The lysate was clarified by centrifugation in a Beckman JA-25.50 rotor at 20,000g for 30 min. The supernatant was collected and incubated with anti-Flag M2 affinity beads (Sigma-Aldrich) on a rocker for 1.5 h. The beads were collected and washed three times with myosin purification buffer (50 mM Tris-HCl pH 8.0, 150 mM NaCl, 2 mM MgCl2 and 2 mM ATP), then eluted with myosin purification buffer supplemented with 100 µg ml−1 Flag peptide (Sigma-Aldrich). The eluted protein was buffer exchanged into myosin storage buffer (10 mM Tris-HCl pH 8.0, 100 mM NaCl, 2 mM MgCl2 and 3 mM DTT) using an Amicon Ultra-4 concentrator (50 kDa MWCO), then snap frozen in liquid nitrogen. Purified myosins were stored at −80 °C prior to use.
The C-terminal ABD of human α-catenin (amino acids 664–906) was expressed from a pET vector encoding an N-terminal 6 × His tag, strep tag, and TEV protease cleavage site as previously described8. Transformed Rosetta2(DE3) E. coli were cultured in LB medium at 37 °C to an optical density of 0.8–1.0, then induced with 0.7 mM isopropyl β-D-1-thiogalactopyranoside (IPTG) and cultured at 16 °C for 16 h. Cells were then collected and flash frozen in liquid nitrogen. Cell pellets were stored at −80 °C.
For purification, all steps were performed at 4 °C. Cells were resuspended in lysis buffer (50 mM Tris-HCl pH 8.0, 150 mM NaCl, 2 mM β-mercaptoethanol, 20 mM imidazole) and disrupted in an Avestin Emulsiflex C5 homogenizer. The lysate was clarified at 15,000g in a Beckman JA-25.50 rotor for 30 min, then the supernatant was collected and incubated with Ni-NTA resin (Qiagen) with rocking for 1 h. The resin was collected, washed with 5 bed volumes of lysis buffer, then eluted with lysis buffer (50 mM Tris-HCl pH 8.0, 150 mM NaCl, 2 mM β-mercaptoethanol, 300 mM imidazole). Purified His-tagged TEV protease (prepared according to a published protocol86) was then added to reach a 0.05 mg ml−1 working concentration, and the solution was dialysed against dialysis buffer (20 mM Tris-HCl pH 8.0, 300 mM NaCl, 2 mM β-mercaptoethanol) for 16 h. The sample was then collected and re-applied to Ni-NTA resin to remove TEV protease. The flowthrough was collected, then sequentially purified by anion exchange chromatography using a HiTrapQ HP column (Cytiva), followed by size-exclusion chromatography on a Superdex 200 Increase column (Cytiva) in gel filtration buffer (20 mM Tris-HCl pH 8.0, 100 mM NaCl, 2 mM β-mercaptoethanol). The protein was snap frozen in liquid nitrogen and stored at −80 °C until use.
Force reconstitution on cryo-EM grids
All steps were performed at room temperature unless otherwise noted. F-actin was prepared from G-actin monomers in G-Mg (G buffer supplemented with 0.1 mM MgCl2) plus KMEI buffer (50 mM KCl, 1 mM MgCl2, 1 mM ethylene glycol-bis(β-aminoethyl ether)-N,N,N′,N′-tetraacetic acid (EGTA), 10 mM imidazole pH 7.0, 1 mM DTT) as previously described45. CF-1.2/1.3-3Au 200-mesh gold C-flat holey carbon cryo-TEM grids (EMS, CF213-50-Au) were used for initial cryo-EM imaging and tomography studies, while CF-1.2/1.3-3 Au 300-mesh grids (EMS, CF313-50-Au) were used for single-particle studies.
For the dual motor condition, the following solutions were prepared at 4 °C in motility buffer (MB, 20 mM 3-(N-morpholino)propanesulfonic acid (MOPS) pH 7.4, 5 mM MgCl2, 0.1 mM EGTA, 1 mM DTT): MB-PVP, 0.1% polyvinylpyrrolidone (Sigma-Aldrich, PVP10); MB-anchor, 0.1 mg ml−1 mouse monoclonal anti-GFP antibody (Sigma-Aldrich, G6539); MB-myosin, 0.02 μM myosin V and 0.08 μM myosin VI; MB-ATP, 1 mM ATP and 0.01% Nonidet P-40 Substitute (NP-40, Roche); MB-no-ATP, 0.01% NP-40. Solutions were clarified by ultracentrifugation at 50,000 rpm (108,726g) in a Beckman TLA-100 rotor, then brought to room temperature immediately prior to grid preparation.
Grids were untreated prior to sample preparation, as we found plasma cleaning and other treatments reduced the activity of myosins. First, 6 μl of MB-anchor was applied to each side of the grid and incubated for 2 min. Six microlitres of MB-myosin was then applied to each side of the grid and incubated for an additional 2 min. Twenty microlitres of MB-PVP was then applied to the top (shiny) side of the grid and incubated to block the surface for 1 min. Six microlitres of F-actin was then applied and incubated for 40 s. The grid was then washed in a 500 μl reservoir of MB in a microcentrifuge tube, then placed in the chamber of a Leica EM-GP plunge-freezing device operating at 25 °C and 100% humidity. Six microlitres of either MB-ATP (+force generation) or MB-no-ATP (no force generation) was then applied and incubated for 40 s. The grid was then blotted for 5 s and plunged into liquid ethane. For each grid preparation session, samples with F-actin concentrations ranging between 0.4–1.0 μM were prepared and screened, and specimens featuring optimal filament density were selected for further analysis.
For the barbed-end-directed and pointed-end-directed single-motor conditions, reagents were prepared as above, with the following modifications: MB-anchor was supplemented with 500 nM streptavidin (VWR, S000-01), and two separate myosin buffers were prepared, MB-myosin V (0.02 μM myosin V in MB) and MB-myosin VI (0.08 μM myosin VI in MB). To prepare 25% biotin–F-actin seeds biotinylated actin monomers (Cytoskeleton, AB07-C) were co-polymerized at a 1:4 molar ratio with unlabelled monomers with a total concentration of 10 μM for 1 h, then sheared through a 30G needle. F-actin was then extended by mixing seeds with unlabelled monomers at a 1:5 molar ratio and polymerizing for an additional 1 h. Samples were then prepared as above, using either MB-myosin V (barbed-end-directed) or MB-myosin VI (minus-end-directed).
Samples for the force-activated α-catenin–F-actin complex were prepared identically as for the dual motor condition, with the following modifications. After mounting in the plunge-freezing apparatus, 3 μl of MB-ATP supplemented with α-catenin ABD was applied and incubated for 30 s, then the grid was blotted for 5 s and plunged into liquid ethane. Grids were prepared with varying F-actin/α-catenin ABD concentrations, and 0.6 μM F-actin/1 μM α-catenin ABD was found to be optimal after screening. All grids were stored in liquid nitrogen prior to cryo-EM data collection.
Fluorescence imaging of myosin activity
For data presented in Supplementary Videos 2–4, samples were prepared as described above, except 10% rhodamine G-actin (Cytoskeleton, AR05-B) was included in all polymerization reactions (F-actin and biotinylated seeds). Instead of plunge freezing, grids were sandwiched between two No. 1.5 24 ×60 mm glass coverslips (Corning) in a drop of MB-ATP supplemented with oxygen scavengers (20 μg ml−1 catalase, 100 μg ml−1 glucose oxidase, 125 mg ml−1 glucose, 50 mM DTT). Epifluorescence image sequences (movies) were collected at room temperature using a 60× 1.40 NA PlanApo oil-immersion objective lens (Nikon) on an inverted TE2000-E microscope (Nikon) equipped with a solid-state white light illumination system (Lumencor SOLA SE) controlled through the Metamorph software package (Molecular Devices). Images were collected on a Photometrics Coolsnap HQ2 CCD camera with a frame rate of 1 exposure every 2–3 s, using filters for visualizing rhodamine (excitation 650/25 nm, emission 607/36 nm). A single image (either bright-field to directly visualize the cryo-EM grid, or fluorescence using filters to visualize anchored, GFP-tagged myosins: excitation 485/20 nm, emission 525/30 nm) was taken before and after each image sequence to visualize the grid substrate and verify the lack of substantial drift.
Low-magnification cryo-EM
For data and analysis presented in Fig. 1d–f and Extended Data Fig. 2a–d, samples were imaged on a Tecnai F20 transmission electron microscope operating at 120 kV using a Gatan 626 cryo-transfer holder, operated with the Leginon software package87. Exposures were collected at 29,000× magnification (corresponding to a calibrated pixel size of 3.2 Å per pixel) on a Gatan Ultrascan 4000 CCD camera with –4 µm underfocus. Images were low-pass filtered to 20 Å and binned by 4 for visualization and analysis. Initial quantification of sinusoidal filament region wavelengths (presented in Fig. 1f and Extended Data Fig. 2a) was performed manually with FIJI88 by measuring the distance between peaks along segments with clear oscillating curvature.
Filament annealing refractory condition
For data and analysis presented in Extended Data Fig. 2e,f, 50% PEG-biotin coated coverslips were prepared by adapting a previously described protocol30. No. 1.5H 24 × 60 mm glass coverslips (Marienfeld) were sequentially sonicated in 2% Hellmanex III (Hellma), acetone, and 1 M KOH. Coverslips were then sonicated in MilliQ water three times, followed by treatment with Nano-Strip 2X (Cyantek), then rinsed with MilliQ water. Coverslips were then coated in 0.5 mg ml−1 mPEG silane MW 5 K (Laysan Bio) and 0.5 mg ml−1 biotin–mPEG silane MW 5 K (Laysan Bio) in 96% ethanol and 9.6 mM HCl. PEG-coated coverslips were then washed in 100% ethanol, rinsed with MilliQ water, and dried before immediate use or storage at 4 °C.
For total internal reflection fluorescence (TIRF) microscopy experiments, 20% biotin-labelled F-actin seeds were prepared by co-polymerizing 0.6 μM of unlabelled purified G-actin with 0.2 μM biotin-labelled rabbit skeletal muscle G-actin (Cytoskeleton) and 0.2 μM ATTO-488-labelled rabbit skeletal muscle G-actin (Hypermol) in the presence of G-Mg + KMEI buffer at room temperature for 1 h. The biotinylated F-actin solution was sheared 5 times with a 26G needle, then mixed with G-Mg + KMEI buffer containing 2.4 μM unlabelled G-actin, 0.8 μM biotin-labelled rabbit skeletal muscle G-actin (for anchoring filament extensions to glass for imaging), 0.8 μM rhodamine-labelled rabbit skeletal muscle G-actin (Cytoskeleton) and 100 μM profilin (purified as previously described30,89) at a 1:19 volume:volume ratio, then incubated at room temperature for an additional 1 h. Polymerization was subsequently halted by the addition of 0.3 μM capping protein (CapZ, Hypermol), producing capped F-actin extended from biotinylated seeds.
Imaging chambers were prepared by attaching CultureWell reusable PDMS gaskets (Grace Bio-Labs) to PEG-biotin coated coverslips. Each well was treated with the following sequence: incubation with 500 nM of streptavidin (Rockland) in MB supplemented to 3 mM DTT for 4 min, blocking with Blocking Buffer (MB-PVP supplemented with 1 mg ml−1 κ-casein (Sigma) and 1 mg ml−1 bovine serum albumin (BSA, Gemini)) for 1 min, incubation with capped F-actin extended from biotinylated seeds for 2 min, washing with MB, then immersion in MB + oxygen scavenger mixture (described above) for TIRF imaging.
Images were acquired at room temperature on a Nikon H-TIRF system through a CFI Apo 100× TIRF oil-immersion objective (NA 1.49), a quad filter (Chroma), and an iXon EMCCD camera (Andor) with the Perfect Focus System (Nikon) engaged. Illumination was provided by 488 and 561 nm laser lines (Agilent) switched by an acousto-optic tunable filter. Images were captured at a depth of 16-bit using the NIS-Elements software (Nikon).
Optimized pointed-end force grids
To prepare cryo-EM specimens of the pointed-end-directed force condition using capped F-actin extended from biotinylated seeds (Extended Data Fig. 2g,h), we implemented an improved procedure by using an anti-GFP nanobody conjugated to fibrinogen (adapted from the protocol of Watson et al.90) to anchor myosin VI to the surface. This enabled us to maintain myosin activity after plasma cleaning the grids, improving sample quality.
To prepare the fibrinogen-coupled nanobody, we expressed and purified anti-GFP nanobody91 LaG 94-10 and chemically conjugated it to fibrinogen (MilliporeSigma, 341576). Periplasmic expression of the nanobody from the pET-21b vector was induced in BL21(DE3) E. coli cells at 0.6 OD using 0.1 mM IPTG, which were collected after overnight culture at 12 °C, flash frozen in liquid nitrogen, and stored at −80 °C. Cell pellets were resuspended in TES buffer (0.2 M Tris-HCl pH 7.5, 0.5 mM ethylenediaminetetraacetic acid (EDTA), 0.5 M sucrose), then lysed by osmotic shock on ice for 20 min through the addition of a 1:4 volume:volume ratio mixture of TES and MilliQ water in 1.5 fold excess over the original resuspension volume. Protoplasts were separated from the periplasmic fraction by centrifugation at 4,000g for 10 min at 4 °C. The supernatant was collected, then centrifuged at 48,000g for 30 min at 4 °C to remove insoluble material present in the periplasm lysate92.
The soluble fraction was then mixed with an equal volume of Nanobody Wash Buffer (20 mM sodium phosphate pH 8.0, 1 M NaCl, 20 mM imidazole) and incubated with Ni-NTA resin for 20 min at 4 °C. The resin was washed with 25 column volumes of Nanobody Wash Buffer before elution with Nanobody Elution Buffer (20 mM sodium phosphate pH 8.0, 0.5 M NaCl, 0.3 M imidazole). The eluted nanobody was buffer exchanged into fibrinogen buffer (0.1 M sodium bicarbonate pH 8.3, 0.5 mM EDTA) supplemented with 1 mM tris(2-carboxyethyl)phosphine (TCEP) and concentrated to 15 mg ml−1 using a 3 kDa MWCO centrifugal filter (Amicon).
Fibrinogen was resuspended in fibrinogen buffer, then reacted with 25-fold molar excess of maleimide-PEG8-succinimidyl ester (Millipore Sigma, 746207) while rocking at room temperature for 1 h. Maleimide coupled fibrinogen was then separated from unreacted maleimide-PEG8-succinimidyl ester by two rounds of precipitation with 25% mass/volume ammonium sulfate while rocking at room temperature for 30 min, centrifugation at 17,000g for 10 min, and resuspension in fibrinogen buffer. Maleimide coupled fibrinogen was then reacted with 5-fold excess of freshly purified anti-GFP nanobody overnight at 4 °C. The coupling reaction was quenched by the addition of 50 mM L-cysteine and incubation at room temperature for 30 min, followed by two additional rounds of ammonium sulfate precipitation to isolate the nanobody-fibrinogen conjugate. The conjugate was reconstituted to a final concentration of 5 mg ml−1 in fibrinogen buffer, flash frozen in liquid nitrogen, and stored at −80 °C.
For cryo-EM sample preparation, reagents were prepared as described above for reconstituting myosin activity on grids, with the following modifications: MB-anchor was replaced with 0.5 mg ml−1 anti-GFP nanobody-fibrinogen conjugate and 0.5 μM streptavidin prepared in MB, and the concentration of myosin VI in MB-myosin VI was increased to 0.5 μM. Capped F-actin extended from biotinylated seeds was prepared as described above while omitting both fluorescently labelled G-actins (which were substituted by equivalent amounts of unlabelled purified G-actin). Biotinylated actin was also omitted when filaments were extended from seeds. For sample preparation, grids were plasma cleaned using a Gatan Solarus in the presence of H2 and O2 for 10 s, then placed in CultureWell PDMS gaskets affixed to untreated coverslips. Grids were then treated sequentially with 30 μl of each solution as described above for preparing single-motor samples. For the no-ATP control, 200 μl of capped F-actin extended from biotinylated seeds was pre-mixed with 0.2 μl of apyrase (NEB M0398S), and specimens were incubated at room temperature for an additional 5 min upon addition of this F-actin to the grid. Prior to freezing, 3 μl of MB-ATP (+ATP condition) or MB-no-ATP (−ATP, +apyrase condition) was applied. Grids were incubated for 40 s, then blotted for 6 s and plunged into liquid ethane.
Single-particle cryo-EM data acquisition
For the dual motor condition and filament annealing refractory pointed-end-directed force condition, cryo-EM data were recorded on a Titan Krios (ThermoFisher/FEI) operated at 300 kV equipped with a Gatan K3 camera, BioQuantum energy filter and spherical-aberration (Cs) corrector. SerialEM70 was used for automated data collection. Movies were collected at a magnification of 64,000× in super-resolution mode resulting in a calibrated pixel size of 1.08 Å per pixel (super-resolution pixel size of 0.54 Å per pixel). For the dual motor condition, a defocus range of −1.5 to −3.5 µm underfocus was used. Sixty-three frames were recorded over 2.5 s of exposure at a dose rate of 27.6 e− pixel−1 s−1 (23.7 e− Å−2 s−1) for a cumulative dose of 59.2 e− Å−2. Three datasets were collected with these imaging conditions consisting of 12,453 movies, 10,842 movies and 8,504 movies, respectively. The first dataset was used for preliminary processing, and all three datasets were pooled to generate the final reconstruction. A single no-ATP control dataset was collected under the same imaging conditions, consisting of 3,749 movies. For the filament annealing refractory pointed-end-directed force condition, a defocus range of −0.8 to −2.5 µm underfocus was used. Forty frames were recorded over 2 s of exposure at a dose rate of 30 e− pixel−1 s−1 (25.7 e− Å−2 s−1) for a cumulative dose of 51.4 e− Å−2. 188 movies were analysed from the +ATP sample and 110 movies were analysed from the −ATP, +apyrase control.
For the force-activated α-catenin–F-actin complex, cryo-EM data were recorded on a Titan Krios (ThermoFisher/FEI) operated at 300 kV equipped with a Gatan K2 Summit camera. SerialEM70 was used for automated data collection. Movies were collected at a nominal magnification of 22,500× in super-resolution mode, resulting in a calibrated pixel size of 1.33 Å per pixel (super-resolution pixel size of 0.665 Å per pixel), over a defocus range of −1.5 to −3.5 µm underfocus. Fifty frames were recorded over 10 s of exposure at a dose rate of 10 e− pixel−1 s−1 (e− Å−2 s−1) for a cumulative dose of 56.5 e− Å−2. Three datasets were collected with these imaging conditions consisting of 1,909 movies, 1,792 movies and 754 movies, respectively. These datasets were pooled prior to processing.
Micrograph pre-processing
Movies were aligned with MotionCor2 using 5 × 5 patches72, and dose-weighting sums93 were generated from twofold binned frames with Fourier cropping, resulting in a pixel size of 1.08 Å in the images collected in the dual motor condition datasets (±ATP) and filament annealing refractory pointed-end force datasets (+ATP or −ATP, + apyrase) and 1.33 Å for the force-activated α-catenin–F-actin complex datasets. Non-dose-weighted sums were used for CTF parameter estimation using CTFFIND4 (ref. 73).
Two-dimensional quantification of cryo-EM images
To manually annotate micrographs, we implemented a custom Napari94 plugin. For analysis of the dual motor condition (presented in Extended Data Fig. 2b), data were binned to a pixel size of 12.82 Å per pixel. For analysis of the pointed-end-directed force condition prepared with the filament annealing refractory protocol, data were binned to a pixel size of 4.32 Å per pixel. Micrographs were manually annotated by tracing individual filaments within holes using a stylus; each annotated micrograph was saved as an mrcs stack with a separate channel for each filament. This enabled each filament trace to span the entire observed filament length without discontinuities at intersections with other filament traces.
Filament traces were converted to spline representations by thresholding and skeletonizing each mrcs slice. Filament traces with a contour length of at least 325 nm were retained for analysis. To minimize the effects of long-range filament curvature and to quantify filament properties at the same length scale as the in vitro tomographic analysis (Fig. 2c,d), filament traces were analysed using a 300 nm sliding windows sampled every 25 nm. For each 300 nm-long window, principal component decomposition was performed: PC1 established the filament region’s central axis, and PC2 corresponded to deviation of the filament’s centre from this central axis. Peaks and troughs along the PC1/2 decomposition were identified, and the largest difference in the PC2 value between neighbouring peaks and troughs was recorded as the filament region’s peak-to-peak amplitude. The highest amplitude score along a filament was retained as well as the average absolute curvature of the filament within that window. For the dual motor data, 912 filaments from 100 micrographs were traced for the +ATP condition, of which 857 filaments were longer than 325 nm. For the no-ATP condition, 745 filaments from 46 micrographs were traced, of which 712 filaments were longer than 325 nm. For the filament annealing refractory pointed-end-directed force data, 396 filaments from 188 micrographs were traced for the +ATP condition, of which 312 filaments were longer than 325 nm. For the −ATP, +apyrase condition, 414 filaments from 110 micrographs were traced, of which 320 filaments were longer than 325 nm.
Cryo-ET of reconstituted specimens
Grids were clipped and transferred to Talos Arctica TEM (ThermoFisher) operating at 200 kV with a Gatan K2 Summit camera. Tomograms were collected using a dose-symmetric scheme from –60° to +60° with 3° angular increments. Each tilt image was fractionated into 10 frames with a total exposure dose of 2.4 e− Å−2 (0.24 e− Å−2 per frame) and a total exposure time of 1 s. The defocus range was between −1.5 μm and −5 μm underfocus. The acquisition magnification was 17,000× with a pixel size of 2.4 Å. Frames were aligned with MotionCor2 (ref. 72).
Tomograms were reconstructed using the Appion-Protomo software95,96. Images were aligned and dose-weighted, then reconstructed using the simultaneous iterative reconstruction technique (SIRT) as implemented in Tomo3D97. Reconstructions were binned by 4 resulting in a final pixel size of 9.6 Å.
Synthetic cryo-ET data generation
In vitro tomograms were processed using a denoising autoencoder (DAE) approach to enable direct, confident tracing of F-actin filaments in 3D. A DAE with a similar architecture to that reported in our previous work53 was trained. In brief, synthetic datasets approximating subtomograms were first generated. Twenty-four thousand synthetic noiseless subtomograms were prepared which contained between zero and five in silico actin filament models (PDB: 7R8V) of varying curvature. All of these filament models were bent in a single plane (to prevent supercoil model bias), then converted to volumetric data using the pdb2mrc procedure in EMAN2 (ref. 98). Each actin filament was rotated about the rot and psi angles by random, uniformly sampled values between 0° and 359°, while the tilt rotation angle was sampled from a Gaussian probability distribution centred at 90° with a standard deviation of 20°. Each actin filament was translated in the box along each dimension by random values uniformly sampled within the range of ±196 Å.
Matched synthetic noisy volumes were generated by first projecting the sum of the oriented filaments at integer angular increments (held constant for a particular subtomogram) uniformly sampled between 1° and 5°, inclusive. The maximum tilt was sampled from a random integer between 45° and 60°, yielding tilt angle ranges from ±45° to ±60°. These projections were corrupted by a CTF, using parameters matching those of the microscope and a defocus range of −3 μm to −11 μm underfocus, then reconstructed back into a 3D volume using reconstructor class functions as implemented in the EMAN2 python package98. Empirical noise was modelled by extracting 1,000 subtomograms of 96 × 96 ×96 voxels from each experimental tomogram and computing the per tomogram average Fourier transform. For each synthetic volume, one of these empirical noise boxes was randomly selected, multiplied element-wise by a white noise box of the same dimensions, normalized, and scaled by a random scale factor that modulated the signal-to-noise ratio of each synthetic subtomogram. The synthetic volume was then summed with its noise volume in Fourier space. To account for interpolation artefacts from CTF application or noise addition in Fourier space, the synthetic volumes were then cropped to 48 voxels in real space. After these operations were complete, each noisy particle was paired with its corresponding noiseless ground truth particle.
Denoising in vitro tomograms
A DAE featuring adversarial training was constructed and trained using an approach we have recently reported53. Pre-training of this DAE consisted of 3D convolutional layers in a U-net architecture, and it was performed using a single NVIDIA A100 GPU with 80 GB of VRAM, using a learning rate of 0.00001. Training was run on the 24,000 pairs of synthetic noisy and noiseless volumes, with a 90:10 training:validation split, for 30 epochs. This model had a cross-correlation coefficient validation loss of 0.9147. After pre-training, an additional network head was added for domain classification by forking the network output after the feature extraction layers. The domain classification head consists of a gradient reversal layer and additional 3D convolutional layers, which are followed by flattening to a dense layer, then a binary classification layer with a sigmoid activation function. 1,000 synthetic volumes and 1,000 volumes extracted from the experimental tomograms were used as the training set for the domain classifier. Adversarial training was performed by alternatively passing these data through the domain classifier head, followed by re-training the feature extractors with the denoising head using only the 1,000 synthetic volumes. This adversarial training was run for ten iterations, and the final iteration of adversarial training was used for denoising.
The adversarial-trained neural network was then used to denoise empirical tomograms, using A100 GPUs. 48-voxel tiles sampled every 16 voxels in each dimension were extracted, normalized, and passed as inputs to the neural network. The network outputs were stitched together via maximum intensity projection.
Quantification of in vitro tomograms
Denoising the in vitro tomograms enabled 3D contour tracing of F-actin. Denoised tomograms were visualized in UCSF Chimera99, and each filament’s central axis was manually traced using Chimera’s marker set tool. Univariate splines were fit through the data with a fixed smoothing factor of 150 to minimize jitter from manual picking. The spline fits were resampled evenly with 9.6 Å steps and visually inspected to ensure trace quality.
The cartesian coordinates of these traces were subsequently analysed. The instantaneous curvature and torsion along each filament trace was inspected to identify filament segments with potential oscillatory character. Filament segments with high, oscillating curvature and torsion that were between 350 nm and 500 nm in length were identified and selected. PCA was performed on these selected filament segments, and the decompositions were aligned such that the maximum amplitude along the second principal component was at the centre point and the second derivative of the second principal component at this point was positive. If a filament’s principal component decomposition was inverted from the other traces, the starting end of the filament spline was flipped. One limitation of this analysis is that filament polarity could not be assigned in the denoised tomograms. The aligned traces were analysed within a common 350 nm window to produce average traces, as well as peak-to-peak amplitude and wavelength measurements (Fig. 2c,d).
To assess whether the supercoil filament segments exhibited non-uniform orientation relative to the ice layer, the second principal component’s angle relative to the ice layer was analysed. The tomograms contained protein aggregates at the air–water interface, which appeared as amorphous density in the denoised tomograms. Chimera’s marker set tool was used to position centroids within these densities. PCA was then performed on these points, where the first and second principal components defined the ice plane. The angle between the second principal component of the oscillatory region (that is, the major axis of each supercoil’s elliptical cross-section) and the ice plane was measured by projecting the ice plane’s normal vector along the first principal component of the oscillatory domain and computing the angle between these vectors. A Kolmogorov–Smirnov test as implemented in Scipy100 was performed to assess whether the observed major axis angle distribution was significantly different from a normal distribution, and no significant difference was found (P = 0.27).
Coarse-grained molecular dynamics
Parameterization
Coarse-grained molecular dynamics simulations of individual actin filaments under force were performed using the software package ESPResSO101 and custom Python scripts. Each subunit was mapped to its centroid based on the structure of canonical F-actin48, which consists of two right-handed strands intertwining to form a left-handed helix with a rise of 27.8 Å and a twist of –166.67°. The filament was modelled as a 3D spring network, with each subunit (positioned 1.6 nm radially from the filament axis) connected to its four nearest neighbours (Extended Data Fig. 3a). Implicit solvent effects were simulated using the Langevin thermostat. The filament’s energetics were described by a harmonic bond potential (equation 1), harmonic angle potential (equation 2), and dihedral potential (equation 3):
$$V(l)=\frac12k_l(l-l_)^2$$
(1)
Where \(k_l\) is the harmonic bond stiffness, and \(l_\) is the diagonal or longitudinal bond length in equilibrium configuration.
$$V(\theta )=\frac12k_\theta (\theta -\theta _)^2$$
(2)
Where \(k_\theta \) is the harmonic angle bending constant, and \(\theta _\) is the angle formed by neighbouring triplets in equilibrium configuration. Each particle is associated with six different harmonic angles.
$$V(\phi )=\,k_\phi [1-\cos (n\phi -\,\phi _)]$$
(3)
Where \(k_\phi \) is the dihedral angle bending constant, \(\phi _\) is the dihedral angle formed by neighbouring quadruplets in equilibrium configuration, and \(n\) is the multiplicity of the potential (number of minima). Each particle is associated with two different dihedral angles; \(n\) was set to 1 for simplicity.
The geometric parameters of the equilibrium configuration, \(l_\), \(\theta _\) and \(\phi _\) were derived from the canonical actin filament structure48. For simplicity, deformations of individual subunits were ignored. The effects of actin nucleotide state transitions on filament mechanics were also neglected.
The stiffness and bending constants, \(k_l\), \(k_\theta \) and \(k_\phi \), have not been experimentally measured; nonetheless, they can be calibrated through matching to previously reported data, as demonstrated by Yogurtcu et al.102. Moreover, Schramm et al. previously developed a mesoscale model to investigate actin and cofilactin filament buckling and fragmentation under mechanical strain103. Their approach began with all-atom molecular dynamics simulations, from which they constructed elastic network models by mapping each actin subunit to its centre of mass and connecting it to four neighbours via elastic bonds. Bond stiffnesses were iteratively optimized to reproduce molecular dynamics-derived fluctuations and then incorporated into a mesoscale model, where each subunit was approximated as an ellipsoid. Harmonic bonds were randomly distributed across subunit interfaces based on buried solvent-accessible surface area.
By contrast, our model describes filament energetics using harmonic bond, angle, and dihedral potentials. Each interaction interface is represented by a single bond. Interaction potentials were parameterized sequentially and iteratively until convergence, as detailed below.
The harmonic bond potential captures F-actin’s tensile rigidity and was calibrated to match a force-extension relationship previously reported by Chu and Voth104. Using coarse-grained molecular dynamics simulations, they probed the force-extension behaviour of actin filaments, reporting stretch stiffness values of 37 pN nm−1 for ATP F-actin and 31 pN nm−1 for ADP F-actin104. To calibrate \(k_l\), parameter scans were performed by simulating an actin filament composed of 39 subunits (the same number of subunits modelled by Chu and Voth) under tension. Simulations were conducted with a low friction coefficient at extremely low temperature (0.298 K) to minimize stochastic deviations. The stretch stiffness was extracted by plotting the pulling force against the filament extension at equilibrium (Extended Data Fig. 3b). The target value for calibration was 31 pN nm−1, corresponding to ADP F-actin to match our cryo-EM sample conditions.
The harmonic angle potential reflects F-actin’s bending rigidity and was optimized to reproduce the 9 μm persistence length of ADP F-actin57. To calibrate \(k_\theta \), parameter scans were performed by simulating thermal fluctuations of an actin filament composed of 100 subunits using the friction coefficient of water at room temperature. To compute the persistence length, a centre axis approximating a linear polymer was constructed as follows: for each subunit \(i\), an anchor point on the opposite strand was calculated as the midpoint between subunits \(i-1\) and \(i+1\). The midpoint between subunit \(i\) and this anchor, denoted \(c_i\), was used to trace the filament’s centre axis. The persistence length \(P\) was determined using the relation:
$$\mathrmln\langle \cos \alpha _i,i+n\rangle =\,-\fracL_nP$$
(4)
where \(\alpha _i,i+n\) is the angle between the segments \((c_i-\,c_i-1)\) and \((c_i+n-\,c_i+n-1)\), and \(L_n\) is the contour length defined as the sum of segment lengths \(|c_i+1-\,c_i|\) from \(c_i\) to \(c_i+n\). The persistence length was extracted from the slope of the linear fit (Extended Data Fig. 3c).
The dihedral potential captures F-actin’s twisting rigidity and was optimized to reproduce the dependence of cumulative twist variance on subunit index, as established from cryo-EM data by Bibeau et al.105. To calibrate \(k_\phi \), parameter scans were performed by simulating thermal fluctuations of an actin filament under the same conditions used for calibrating \(k_\theta \). Subunit twist was calculated using the centre axis defined as described above. For each pair of consecutive subunits \(i-1\) and \(i\), their positions were projected onto the segment \((c_i-\,c_i-1)\) along the centre axis, and the twist angle was determined from the resulting projection vectors. At each time point, the deviation of each subunit’s twist from the canonical F-actin value was computed. A zero-mean normal distribution was then fitted to the distribution of these deviations across the simulation for each subunit, and the variance was extracted. The twist variance was plotted against subunit index (Extended Fig. 3d). To avoid edge effects, the first and last 20 subunits were excluded, resulting in the same number of analysed subunits (n = 60) as in the study by Bibeau et al.105.
Unit derivations
ESPResSO does not predefine units. Instead, the user defines the fundamental units of length and energy, from which all other units can be derived.
In our simulations, we set the length unit as:
We simulate the system at room temperature (298 K), so the energy unit is defined as:
$$[E]=\,k_B\cdot 298\,\rmK=4.12\times 10^-21\,\rmJ$$
By default, ESPResSO also has no specification of particle mass. We thus define the mass unit using the molecular mass of G-actin (M = 42 kg mol−1) and Avogadro’s constant (NA = 6.02 × 1023 mol−1):
$$[m]=\fracMN_\rmA=\frac42\,\mathrmkg\,\mathrmmol^-16.02\times 10^23\,\mathrmmol^-1=6.97\times 10^-23\,\mathrmkg$$
Using the kinetic energy relation \(E\,=\,\frac12mv^2\,=\,\frac12m\left(\fraclt\right)^2\), the time unit is derived as:
$$[t]=[l]\cdot \sqrt\frac[m]2[E]=1\times 10^-9\,\rmm\times \sqrt\frac6.97\times 10^-23\,\mathrmkg2\times 4.12\times 10^-21\,\rmJ=1\times 10^-10\,\rms=0.1\,\mathrmns$$
Based on the relation \(F=\fracEl\), the force unit is derived as:
$$[F]\,=\,\frac4.12\times 10^-21\,\rmJ1\times 10^-9\,\rmm=4.12\times 10^-12\,\rmN\approx 4\,\mathrmpN$$
Therefore, in our simulations, a force value of 1.5 corresponds to a physical force of approximately 6 pN. All final parameters in both simulation units and physical units are reported in Extended Data Table 2.
Force perturbations
The dynamics of a 400-subunit actin filament were then simulated. Thermal fluctuations were simulated at room temperature with the friction coefficient of water. The filament force response was then simulated by fixing the initial five subunits \((i=1:5)\) and applying compressive or tensile force on the terminal five subunits \((i=396:400)\), with their orientations constrained to prevent rotation.
To mimic the mechanical fragmentation behaviour of a physical filament, filament ruptures were implemented by enabling bond breaking when the length of a diagonal or longitudinal bond exceeded defined critical values. These critical lengths were estimated based on the helical parameters measured from the cryo-EM density map of supercoil F-actin (Fig. 3a,b), using the following equations:
$$l_\mathrmdiag=\sqrt2r^2(1-\cos $$
(5)
$$l_\mathrmlong=\,2\sqrt\beta ^2+h^2$$
(6)
Where \(l_\mathrmdiag\) and \(l_\mathrmlong\) are the diagonal and longitudinal bond length, respectively; \(h\) is the helical rise, \(\beta \) is the helical twist, and \(r\) is the distance from the subunit to the filament’s centre axis.
Based on the measured ranges \(h\in [2.5\,\mathrmnm,\,3.2\,\mathrmnm]\) and \(\beta \in [-177^\circ ,\) \(-154^\circ ]\) (Fig. 3b), the maximum diagonal bond length is 4.5247 nm (at \(h=3.2\,\mathrmnm\) and \(\beta =-177^\circ \)), corresponding to a 7.2% increase over the canonical value of 4.2226 nm. The maximum longitudinal bond length is 6.5519 nm (at \(h=3.2\,\mathrmnm\) and \(\beta \,=\,-154^\circ \)), corresponding to a 16.8% increase over the canonical value of 5.6087 nm.
Bond breaking was treated as irreversible; thus, subunit reincorporation and re-annealing of filament fragments were not permitted.
Actin filaments under tension or compression were simulated at a range of forces (10 pN, 20 pN, 30 pN and 40 pN total force) to investigate the effect of force magnitude on the resulting supercoil structures. For each replicate (n = 5) of each condition, a random seed was selected, then an initial simulation was performed under constant force with bond breaking enabled (Extended Data Fig. 4b), and the simulation step at which the filament ruptured was recorded. To maintain filament integrity for supercoil parameter analysis, a second simulation (with bond breaking disabled) was performed where force was applied until the previously observed rupture simulation step, after which force was released, followed by 5,000 additional simulation steps. Analyses for the ‘constant force’ (Fig. 2e) conditions were performed on simulation frame 0 through the frame preceding force release (Fig. 2g and Extended Data Fig. 3e,f), and analyses for the ‘apply force and release’ conditions (Extended Data Fig. 4c) were performed on all frames following force release (Fig. 2h and Extended Data Fig. 3e,f).
To mimic the asynchronous activity of myosin motors, a stochastic motor-firing scheme was implemented by randomly selecting five subunits from the lower half of the filament (\(i\in [200,400]\)), with each subjected to a 6 pN pulling or pushing force for a randomly assigned duration (Extended Data Fig. 4d). Bond breaking was disabled in these simulations.
Actin filaments under torque were simulated by fixing the initial two subunits (\(i=\mathrm1,2\)) and applying either under-twisting or over-twisting torque to the terminal two subunits \((i=\mathrm399,400)\). The rotation axis was defined by two midpoints: one between the last \((i=400)\) and second-to-last \((i=399)\) subunits, and the other between the second-to-last \((i=399)\) and third-to-last \((i=398)\) subunits. Vectors for torque application (levers) were calculated by projecting the positions of the last two subunits onto the rotation axis, and the direction of the applied force was determined by the cross product of the lever and the rotation axis. Forces of 80 pN and 800 pN were applied to each of the terminal two subunits \((i=\mathrm399,400)\) to probe the effects of torque magnitude. Simulations were performed both with and without bond breaking enabled (Extended Data Fig. 4e).
To investigate the effects of helical parameters on supercoil architecture, simulations were performed with a 200-subunit filament (to maintain reasonable computation times) where F-actin’s helical rise and twist were systematically varied around their canonical values. Separately, to examine the influence of flexural rigidity, the harmonic angle bending constants were adjusted to produce a range of persistence lengths, calculated using the following equation:
$$P=\fracB_sk_\rmBT$$
(7)
Where \(P\) is persistence length, \(B_s\) is flexural rigidity, \(k_\rmB\) is the Boltzmann constant, and \(T\) is absolute temperature (n.b. persistence length scales linearly with flexural rigidity, rendering it a suitable proxy). In both cases, simulations were performed by fixing the initial five subunits \((i=1:5)\) and applying a 6 pN compressive force to each of the terminal five subunits \((i=196:200)\), with their orientations constrained to prevent rotation. Bond breaking was disabled in these simulations.
All mechanical perturbations and thermal fluctuations were simulated using the friction coefficient of water at room temperature.
Supercoil analysis
To compute the amplitude of a supercoil, the filament’s centre axis was traced at each time point as described above. Analogous to our quantification of sinusoidal filament regions in cryo-EM images, a sliding window with a contour length of 300 nm was applied along the axis using a 25 nm step size. Within each window, PCA was performed on the segment, and PC2 was plotted against PC1. Peaks and troughs were identified, and the maximum difference along PC2 between neighbouring peaks and troughs was recorded for each window. The largest such difference across all windows of a given simulation frame was used to determine the instantaneous supercoil peak-to-peak amplitude for that frame.
To identify actin filaments that adopted supercoil configurations, a peak-to-peak amplitude cut-off of 16 nm was applied. For filaments meeting this criterion, the supercoil pitch was determined using the distance along PC1 between the neighbouring peak and trough corresponding to the recorded amplitude; the pitch was calculated as twice this distance.
To enable comparison with experimental measurements (Fig. 1f and Extended Data Fig. 2a), lower and upper pitch cut-offs of 75 nm and 350 nm, respectively, were applied when analysing thermal fluctuations, and constant force and force and release conditions (Fig. 2e–h and Extended Data Fig. 3e,f). For simulations exploring the effects of helical parameters and flexural rigidity (Extended Data Fig. 3g,h), pitch analysis was conducted on all frames until the filament reached a 50% reduction in end-to-end length, with no cut-offs applied.
Eccentricity (Extended Data Fig. 3i) was calculated from ellipses fitted to the PC3 versus PC2 projections of the maximal peak-to-peak amplitude window of each frame as described above.
All analyses were performed using trajectories from five independent simulations per condition (n = 5).
Neural network particle picker training
To analyse single-particle data, a DAE featuring a previously described architecture48 was trained to learn features of actin filaments in cryo-EM projection images (Extended Data Fig. 5a). In brief, the DAE consisted of an encoder and decoder. The encoder was composed of nine convolutional layers followed by three dense, fully connected layers of decreasing size. The decoder was composed of two dense, fully connected layers of increasing size, which then connected to nine convolutional layers, the last of which produced the denoised image.
Synthetic projection images to train the DAE and an accompanying semantic segmentation network were generated as outlined previously48. To improve network performance for semantic segmentation of the experimental data presented in this study, two modifications were made: incorporation of known background picks, and improvement of the noise model. Our data collection scheme resulted in a substantial amount of carbon in the micrographs. To prevent the semantic segmentation network from picking the edges of holes or actin filaments over carbon, 15,208 manual picks of hole edges and thick carbon areas were selected from micrographs in the dataset using RELION106. These picks were extracted at a box size of 512 pixels, then binned by 4 to a box size of 128 pixels and a pixel size of 4.32 Å per pixel. These picks were then integrated into the training dataset of the fully convolutional neural network for semantic segmentation, paired with targets composed entirely of background with no signal.
In our previously reported neural network training schemes, EMAN2’s pink noise generation function98 was used to produce realistic-looking projection images. However, this noise model proved insufficient to pick on the current datasets, possibly due to ice thickness or different microscope or detector parameters. To improve the approximation of the synthetic data to real micrographs, an empirical pink noise model was used. The synthetic 2D particle image generation model can be summarized by the following equation:
$$X_i=C_i\mathop\sum \limits_k=0^KP(\phi _k)V_k+\eta _i$$
(8)
The following procedure was used to generate a particle image \(X_i\): first, a synthetic volume was randomly chosen from a library of 135 bent actin filaments \(V_k\), and the pose of the filament was transformed by a 3D rotation and translation before it was projected \(P(\phi _k)\). This process was repeated for each filament in the image (\(K\), up to four filaments per particle image), and the results were summed in real space to produce a noiseless projection image. Then, the image was corrupted by a CTF with the sampled defocus range, Cs, amplitude contrast, and voltage matching those used for CTF estimation. Finally, noise was added in Fourier space \(\eta _i\).
The noise model, \(\eta \), was empirically determined by using RELION to manually pick and extract 2,177 empty regions within holes in the micrographs to yield empty noise boxes with pixel sizes of 4.32 Å per pixel and box sizes of 128 pixels. The average and standard deviation of the Fourier transform of each empty pick was computed to model the empirical noise distribution. For each synthetic particle image, a white noise box was generated, and the Fourier transform of the white noise box was multiplied element-wise by the sum of the empirical noise average and a Gaussian sampling of the empirical noise standard deviation. This pink noise box, \(\eta _i\), was added to the Fourier transform of the image after CTF corruption, and the inverse Fourier transform was computed to yield the final noisy projection image.
To produce the semantic segmentation target, the noiseless projection image was low-pass filtered to 40 Å, binarized, and eroded by 8 pixels.
To train the DAE, 800,000 image pairs with a 90:10 training:validation split and a learning rate of 0.00005 were used to train the network. Cross-correlation coefficient was used as the loss function. The network was trained until validation loss did not improve for three epochs; then the best network weights were restored and saved. The validation cross-correlation coefficient of the trained DAE was 0.9887.
The trained convolutional layers of the DAE were then used to initialize the weights of the initial layers of a fully convolutional neural network for semantic segmentation, featuring a previously described architecture48. In brief, the trained convolutional encoding layers of the DAE were copied as a separate neural network, and additional convolutional layers were added to form a fully convolutional neural network for semantic segmentation107. The final layer of this network consisted of two channels with sigmoid activation and default initializations. To train the network for semantic segmentation, 75,000 image pairs (60,000 of which contained synthetic projections and 15,000 of which were carbon picks, as described below) with a 90:10 training/validation split and a learning rate of 0.001 were used to train the network. Binary cross-entropy (BCE) was used as the loss function. The network was trained until validation loss did not improve for three epochs; then the best network weights were restored and saved. The validation BCE of the trained DAE was 0.0444. The architectures described above have since been superseded by a U-net architecture, which we found produces better segmentation loss with a smaller training set in a shorter time108.
Particle picking with trained network
A custom Python script was used to pass images to a fully convolutional neural network for semantic segmentation and execute curvature-sensitive filament picking, modified from our previously described method48. In brief, each micrograph was binned by 4 to a pixel size of 4.32 Å per pixel, then 128-pixel tiles featuring 32 pixels of overlap were extracted and passed as inputs to the network. The outputs were stitched together by maximum intensity projection at the overlaps, producing a semantic segmentation map of the micrograph. These maps were then binarized using a fixed, empirically determined threshold of 0.3 and skeletonized. An additional step of dilation by 8 pixels after the skeletonization was performed to link filament ends, which we found were often disjointed in these datasets. Another round of skeletonization was then performed. Branches shorter than 8 pixels were pruned, and pixels within a radius of 16 pixels from filament intersections were removed. Continuous filaments were then identified by matching tracks with common end points, and 2D splines were fit through the filaments for curvature estimation. To prevent spuriously high curvature values due to edge effects, the terminal 50 pixels of the spline were omitted from picking. From the remaining filament sections, the instantaneous curvature was measured along the spline at three-pixel intervals and used for segment selection. This resulted in substantial filament segment overlap during initial picking; these overlapping segments were retained for initial alignment before subsequent duplicate removal. Segments from the same filament were flagged in the output metadata (a RELION-formatted STAR file).
To be assigned as a supercoil segment, substantial curvature in both positive and negative directions along the same filament were required. An identified filament segment was considered a supercoil if it contained a segment at least 200 Å in length (approximately the length of 7 subunit rises) where the instantaneous curvature was always greater than or equal to 1.5 μm−1, as well as another segment of the same length where the instantaneous curvature was always less than or equal to −1.5 μm−1 (Extended Data Fig. 5b). These criteria exclude straight filaments and filaments with substantial uniplanar curving in one direction. All filament segments assigned as supercoils were retained for subsequent image processing.
Modelling of curvature distributions
The curvature distribution of free, unloaded F-actin would be expected to follow a Boltzmann distribution set by the F-actin’s persistence length and the filament segment length, as we modelled in our previous study48. In this scheme, the energy to bend F-actin is defined as:
$$E\,=\,\frac12k_\rmBTL_\rmpL\kappa ^2$$
(9)
Where is the \(k_\rmB\) is the Boltzmann constant, \(T\) is absolute temperature, \(L_\rmp\) is persistence length in microns, \(L\) is the filament segment length in micrometres, and \(\kappa \) is curvature in μm−1 (refs. 48,58). This corresponds to the probability function defined by:
$$P_\mathrmadjusted(\kappa )\,=\,\frac1Z_\mathrmadjustede^\frac\alpha L_\rmpL\kappa ^22$$
(10)
$$Z_\mathrmadjusted\,=\,\mathop\sum \limits_k=0^\infty e^\frac\alpha L_\rmpL\kappa ^22$$
(11)
Where \(\alpha \) is a multiplicative adjustment factor that serves as a proxy for differing persistence length in this simple approximation. \(\alpha \) was both fixed at a value of one for a basic modelling (Extended Data Fig. 5c, grey curves) and fit to the experimental curvature distributions (Extended Data Fig. 5c, pink and blue curves).
Supercoil F-actin reconstruction
For initial model generation, data from a single imaging session were used. Selected supercoil segments were extracted in RELION (v3.1.2) with a box size of 512 × 512 pixels and pixel size of 2.16 Å per pixel (bin 2; Extended Data Fig. 5d,e). To avoid reference bias, these segments were imported to cryoSPARC109 (v3.2.0) for successive rounds of ab initio initial model generation and subsequent 3D classification. The initial 148k picked segments were used for ab initio model generation with three classes as implemented in cryoSPARC. The subset of filament segments contributing to the best class were then selected and used for three more rounds of ab initio model generation with two classes; after each round only the segments belonging to the best class were retained. The reconstructions from the final round of ab initio model generation were then used as references for a two-class heterogeneous refinement in cryoSPARC, with the initial, unaligned 148k picked segments used as inputs. This heterogeneous refinement produced one class with clear protomer definition and one class with apparently mixed filament polarity. The segments from the good class were retained for subsequent homogeneous refinement in cryoSPARC and 3D auto-refinement in RELION. Focused 3D classification without segment alignment was then performed using three classes. Segments from a single class with poor protomer definition were rejected, and the remaining segments were refined using local 3D auto-refinement in RELION to produce the initial model for processing the full dataset.
For processing of the full dataset of 276k filament segments from three datasets, selected supercoil segments were extracted as described above. These segments were imported to cryoSPARC for 2D classification. After removing junk classes and exceptionally straight filament classes, 173k segments were retained. These segments were passed to a cryoSPARC homogeneous refinement for initial alignment. The previously obtained 3D reconstruction was low-pass filtered to 30 Å and used as the initial model. After exporting a RELION STAR file containing the cryoSPARC alignment parameters, the original filament tube IDs were restored using a custom python script. A second custom python script implementing per-filament non-maximum suppression was used to remove overlapping segments from the same filament that were closer than 768 pixels at bin 1 (approximately 830 Å), yielding 13,146 filament segments. These segments were re-imported to cryoSPARC and a non-uniform refinement110 was performed to improve local resolution near the filament edges using a mask encompassing the filament segment without overlap (75% z-length). 3DVA111 was also performed in cryoSPARC using default parameters except for the filter resolution, which was set to 8 Å.
Two independent asymmetric reconstructions from the no-ATP control dataset were generated using a similar approach (Extended Data Fig. 6). To obtain these reconstructions, 345k segments from the no-ATP dataset that did not meet the supercoil criteria were extracted in RELION (v3.1.2) with a box size of 256 × 256 pixels and pixel size of 4.32 Å per pixel (bin 4). These segments were imported to cryoSPARC for 2D classification. After removing junk, 315k segments remained, and they were then split into two equally sized subsets. Each subset was then subjected to the following processing steps. First, ab initio reconstruction of one class was performed, followed by homogeneous refinement. The aligned segments were then re-extracted in RELION at a box and pixel size matching that of the supercoil reconstruction. Overlapping segments within 830 Å were then removed, resulting in 3,436 segments in the first subset and 3,137 segments in the second subset. 3D auto-refinement with local angular searches was performed in RELION. These aligned segments were then imported to cryoSPARC for a final non-uniform refinement using a 75% z-length soft mask.
Force-activated α-catenin complex
For the three pooled α-catenin datasets, particle picking was largely similar to the supercoil dataset (Extended Data Fig. 8). In brief, the micrographs were binned to a pixel size of 4.32 Å per pixel (bin 3.25) and picked using the same selection criteria to identify filaments that contained long stretches of high curvature in both directions. In total, 133,677 segments with no overlap within 3 Å were picked, then extracted in RELION with a box size of 384 × 384 pixels and pixel size of 2.66 Å (bin 2). These extracted segments were subjected to initial 2D classification in RELION to remove junk picks. In order to align the psi (in-plane rotation) angle to a common reference, the remaining 110k segments were subjected to 3D classification in RELION using the initial supercoil model low-pass filtered to 30 Å as a reference and an angular sampling interval of 3.7°. The resulting reconstruction deviated substantially from the reference map. Therefore, the Euler angles were reset such that the rot and tilt angles were removed, the tiltPrior was set to 90°, and the psi angle was set as a prior, while translations were retained. 3D classification was performed with global rot search, local tilt and psi searches with a 12° angular search range, and bimodal priors on the psi angle.
After segments were aligned into this one class, subsequent 3D classification was performed with three classes using a fine 1.8° angular sampling interval, a global search for rot, and a 9° angular search range for tilt and psi. This yielded one bare class, which was rejected from further processing, as well as two classes with α-catenin decorating one side of the filament. To retain the maximum number of segments, the aligned segments from both decorated classes were subjected to focused 3D classification with no alignment, using a mask for α-catenin binding sites along one side of the filament. This resulted in one decorated class and one undecorated class. The decorated class was passed to a 3D auto-refinement job, and the undecorated class was subjected to another round of focused 3D classification with no alignment using a mask for α-catenin binding sites along the other side of the filament. This resulted in one undecorated class and one decorated class. The decorated class was retained and subjected to 3D auto-refinement. Upon inspection, the two refined maps were indistinguishable after a shift and rotation of one short-pitch helical step. The constituent segments of one of the classes were therefore rotated and translated by one short-pitch helical step, then combined with the segments from the other class. Overlapping segments within 768 Å were removed and refined in cryoSPARC using non-uniform refinement.
The aligned segments from the refinement were then subjected to 3DVA as implemented in cryoSPARC111. In order to detect large changes, a filter resolution of 20 Å was used. Large changes in filament curvature and α-catenin occupancy were observed along the first variability component. Filament segments from the 0th to the 25th percentile, featuring high α-catenin occupancy, were selected and refined in RELION to produce an asymmetric reconstruction.
Helical parameter measurements
Measurements were performed using our previously described approach48. In brief, canonical F-actin protomers (PDB: 8D13) were rigid-body fitted into each reconstructed map and combined into a single model, and three copies of the model were stitched together to minimize edge artefacts. Only the central protomers from a single model were reported. A 3D spline is fit to the filament axis of each model. Rise is measured by computing the path length along the central axis between neighbouring protomers. Twist is determined using the Frenet-Serret frame of reference defined by the orthonormal basis of the unit tangent, unit normal, and unit binormal vectors along the length of the spline. The frame is then rotated along the normal-binormal plane. Twist is measured along the short-pitch helix between consecutive protomers. The models analysed in this work exhibited substantial lattice deformations that required minor adjustments to reliably measure the filaments’ helical parameters. The measurement approach was updated to account for a variable radius of the filaments. During fitting of the filament axis spline, the radius was allowed to vary along the filament length and was fit using a separate univariate spline; this tended to suppress spurious twist deviations. Additionally, the modest resolution of the reconstructed maps, particularly at the filament edges, limited our confidence in protomer fitting. Consequently, for the consensus supercoil F-actin reconstruction, the no-ATP control reconstructions, and the 3DVA-sorted α-catenin-bound F-actin map, three independent manual rigid-body fittings were performed to provide confidence intervals for the helical parameter measurements.
Actin subdomain rearrangements
First, 25 copies of a canonical protomer model (PDB: 8D13) were rigid-body fit into each map. These 25 protomers were combined into a single PDB model for each map, and these combined models were subjected to Phenix geometry minimization with default parameters of 500 maximum iterations and 5 macro cycles in order to remove clashes. These minimized PDB models were then input to ISOLDE112 and hydrogen atoms were added. Secondary structure distance restraints were imposed on each actin subunit for the following residue ranges: 7–35, 35–72, 72–147, 340–377, 147–183, 272–340 and 183–272, based on previously defined subdomain boundaries113. Torsional and distance restrains were then imposed on the entire secondary structure of each protomer. ISOLDE simulations were run with a weight of 30 × 1,000 kJ mol−1 (map units)−1 Å3. The temperature was set to 100 K for 10 min of simulation, then slowly decreased to 0 K. Subdomain movement magnitudes and directions were determined as previously described48. In brief, protomers were aligned to a reference protomer model (PDB: 8D13) and the displacement vectors between Cαs of the flexibly fit protomers and the reference protomer were computed. Finally, the average of the displacement vectors was calculated for each subdomain defined above.
Molecular graphics and data analysis
Molecular graphics were prepared with ChimeraX81. Unless otherwise noted, plotting and statistical analysis was performed with GraphPad Prism. Python codes were prepared with the assistance of ChatGPT 4.0.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.