Brainwide fUSI measurements
Experiments were conducted in 5 C57/BL6 mice (3 male, 2 female), 13–34 weeks of age. All experimental procedures were conducted according to the UK Animals Scientific Procedures Act (1986). Experiments were performed at University College London, under a Project License released by the Home Office following appropriate ethics review.
Surgery
Mice were first implanted with a headplate and cranial window under surgical anaesthesia in sterile conditions. The cranial window replaced a dorsal section of the skull (∼8 mm in medial–lateral and ~5 mm in anterior–posterior) with 90-μm-thick ultrasound-permeable polymethylpentene (PMP) film (TPX DX845 PMP, Goodfellow Cambridge) attached to the skull with cyanoacrylate glue. The PMP film was then covered with Kwik-Cast (World Precision Instruments), except during imaging sessions. After surgery, mice were allowed to recover for seven days before starting habituation. Mice were then habituated to handling and head fixation in the rig across several days, as session duration progressively increased.
Histology
Mice were perfused with PBS, paraformaldehyde and a gel containing TRITC–Dextran (Sigma-Aldrich) to perform blood vessel labelling. We imaged full 3D stacks of the brains in a custom-made serial section62 two-photon63 tomography microscope. Images were acquired using ScanImage (Vidrio Technologies) and the hardware coordinated using BakingTray. We extracted the signal in green and red channels, for autofluorescence and labelled blood vessels.
fUSI recordings
Recordings were performed in 5 mice across different days for up to 10 weeks. On a given day, we sequentially recorded 2–3 sessions in different coronal planes, lasting ~45 min each. The final dataset consisted of 138 such sessions for a total of 101 h of recordings, ranging from 17 h to 24 h per mouse. Because each plane did not contain all regions, the exact number of recording sessions for each brain region differed (median by animal by region: 11 ± 7.7 s.d.; see Supplementary Fig. 1 for all numbers).
In each recording session, we head-fixed the mice by securing the headplate to a post placed 10 cm from three video displays (Adafruit, LP097QX1, 60 Hz refresh rate) arranged at right angles to span 270° in azimuth and ∼70° in elevation. The screens were uniform grey or black. In some sessions (13 out of 138), the screens switched from grey to black, or from black to grey halfway through the recording session. The mice were placed on a running wheel. Four infrared cameras and the wheel rotary encoder were used to monitor their spontaneous behaviour.
We covered the PMP film with ultrasound gel and positioned the ultrasound transducer above it (128-element linear array, 100 μm pitch, 8 mm focal length, 15 MHz central frequency, model L22-Xtech, Vermon). Doppler signals from the transducer were acquired using an ultrasound system (Vantage 128, Verasonics) controlled by a custom MATLAB-based user interface (Alan Urban Consulting) recording continuously at 500 Hz.
fUSI preprocessing
Coherent tissue motion was removed by performing SVD on the complex images (IQ data) in overlapping 600 ms windows (300 ms overlap) and discarding the first 50 principal components64. This number provided a trade-off between effectively removing tissue motion and preserving actual blood-related signal. Similar results were obtained when removing any number of components between 20 and 100 (Supplementary Fig. 5). Power Doppler images were then obtained by computing the average squared modulus of the filtered IQ data over non-overlapping 300 ms windows.
Compared to experiments in freely moving animals13,65 and with measurements through the cranium66, motion artefacts are less of a concern when fUSI is conducted through a cranial window in well-habituated head-fixed mice. In these conditions, the signals are strong, and the movements are small. Nonetheless, to minimize the possible impact of motion artefacts, we denoised the Power Doppler data using several steps:
-
1.
To correct for brain deformation, we performed motion registration. We used NoRMCorre, a piecewise-rigid registration algorithm from CaImAn67. This step corrects large in-plane movements of the brain in which displacements exceed the voxel size (>0.1 mm), which typically occur during running. We note that off-plane displacements are unlikely to affect the signal, as signals are integrated over a wider distance in the antero–posterior axis (>0.4 mm), much larger than the range of brain movements.
-
2.
To correct for time points with artefactually high values (outliers), we ran a procedure to identify those and replaced them through linear interpolation of neighbouring frames. The detection was based on: (1) identifying noisy voxels as those with kurtosis across time above 10; (2) computing the changes between successive frames of the average activity of these noisy voxels; and (3) identifying time points in which this signal was greater than the mean by 3 × s.d. This led to removing 0–3% of time points (see Supplementary Fig. 6 for the full distribution across sessions).
-
3.
To correct for residual contamination of motion artefacts, we removed any activity that could be linearly predicted from out-of-brain voxels (mostly located in the ultrasound gel, above the surface of the brain). We identified out-of-brain voxels on the basis of low average activity and not being assigned to any brain region. We computed SVD on out-of-brain voxel activity and used the first five components as regressors. We fitted linear weights to predict the full power doppler using ridge regression and subtracted this prediction from the data. We chose this number as a trade-off between capturing enough variance and not introducing noise through the regression. As shown in Supplementary Fig. 6, subsequent components only explained little additional variance.
Finally, we z-scored the denoised power Doppler signal of each voxel across time.
After these corrections, motion artefacts are unlikely to contribute significantly to the large haemodynamic signals we observe with arousal events. First, the increases in fUSI signals seen at times of arousal appear with the typical ~1 s lag of blood signals. If they were motion artefacts, they would occur at zero lag. Indeed, such zero lag effects appeared only when we relaxed our procedures for removing motion artefacts, for example by discarding fewer than ten components in the SVD filtering (Supplementary Fig. 5). Second, many of the arousal events involve only tiny twitches of the nose, which are unlikely to cause artefacts. Consistently, most of our denoising steps do not affect these signals (Supplementary Fig. 6).
Alignment to atlas and region extraction
To extract signals for different brain regions and combine data across sessions and animals, we used a custom, individual-based alignment to the Allen Common Coordinate Framework atlas (CCF)42.
For each mouse, we ran an anatomical scan using a motor interface. This scan consisted of average power Doppler images for short fUSI acquisitions in slices 0.1 mm apart, covering the whole craniotomy. This served as a reference to which we aligned each individual session, as well as the histology.
We perfused each mouse using a blood vessel labelling procedure and imaged the brain (see ‘Histology’), extracting signals in two colour channels: green for autofluorescence and red for labelled blood vessels. We aligned the volume to the Allen CCF atlas using the autofluorescence channel with brainreg68,69,70.
We then aligned the in-vivo fUSI anatomical scan of each mouse to its in-vitro volume using the stained blood vessels. To do so, we used an affine transformation fit to minimize the distance between several manually identified matched key points (~15–30) in both volumes. These were typically branching between big blood vessels visible in both the in-vivo, lower resolution fUSI scan, and the in vitro, higher resolution histology volume. We visually confirmed the proper alignment between the two volumes and then transposed the Allen regions labels to our fUSI volume, so that we obtained a region label for each voxel of the scan.
To select the target regions to keep for our analyses, we used the coarser Beryl mapping40 and set a threshold of volume for regions to be considered (~0.8 mm3), as smaller regions are less likely to be identified with confidence with the resolution of fUSI. We merged regions smaller than this threshold to their parent region and reiterated the procedure until all regions reached the threshold. This procedure resulted in the co-existence of regions at various stages of hierarchy, for example VPM, VP and VENT. To clarify that these excluded some regions, we used ‘-O’ to indicate ‘other’, in VP-O and VENT-O. A full list of region names and acronyms is shown in Supplementary Table 1.
The off-plane (AP) spatial resolution of fUSI images is ~0.4 mm at the focal depth71. Each pixel in the plane therefore corresponds to at least 40 points in the Common Coordinate Framework atlas, which has 10-μm resolution. We assigned a pixel to a brain region if at least two-thirds of those points had the same label. Thus, pixels in zones of transition between regions were not assigned to any region.
For each recording session, we then aligned the slice to our reference anatomical scan in two steps: (1) manual assignment to the closest AP slice in the reference scan; and (2) cross-correlation in space with the chosen slice of the reference scan. We then used the region labels from the aligned reference scan.
We discarded voxels with very little signal due to shadowing, for example, due to bone regrowth. To do so, we applied a Sobel filter to the anatomical image, which computes an approximation of the gradient of the intensity of the image, or edges. We identified shadowed areas by patches of the image (>30 voxels) with consistently low values of the Sobel filtered image. Indeed, in the brain, individual vessels create many edges, so that big patches of low values only occur outside of the brain, or when vessels are hidden by bone.
Once we obtained region labels for each recorded voxel, we computed the region signal for this session using the median activity across voxels. For each region, we only included sessions in which are least 20 voxels of the region were present in the slice. Region signals were extracted for each hemisphere separately and then averaged across the two hemispheres. For further analysis, we considered all regions for which we recorded at least 10 sessions in total over all mice.
Behaviour analysis and onset extraction
Locomotion velocity was obtained from the rotary encoder.
Whisking measurements were obtained through Facemap38. Whisking was obtained by taking the first component of a SVD decomposition of motion energy over the whisker pad. This was done on either one or two cameras (corresponding to both sides of the mouse face). If the signal was available for two cameras, we used the average across both cameras. To have a comparable signal across sessions, we ensured that higher values corresponded to higher motion, and normalized the signal between 0 and 1, based on the first percentile and the maximum.
All behavioural metrics were binned in the same 300 ms bins as the fUSI data.
Whisking bouts were detected when the signal crossed a given threshold (0.15) and lasted until the signal reached another threshold (0.05) under the condition that the average whisking in the next 4 s remained below this threshold (this is to exclude very short interruptions in longer whisking bouts and allow better separation of whisking bouts).
Brief bouts were defined as bouts with duration between 1.3 and 3.5 s, while longer bouts lasted longer than 3.5 s. Whisking bouts were considered accompanied by locomotion when wheel velocity above 1 cm s−1 was detected at any point in the 6 s following whisking onset.
Whisking-evoked activity
To compute whisking-evoked activity for all regions, we first computed results for each mouse and then averaged across mice. For each region and each mouse, we aligned data to the detected onsets (or offsets) and averaged across all recorded whisking events of a given category (for example, short or longer bouts, with or without locomotion). Before averaging, for each bout we excluded times when another, following whisking bout had started. We then averaged results across mice, only keeping regions with more than ten recording sessions in total. Finally, we recentred by the baseline activity (0.5–2 s before whisking onset). Because not all regions were recorded in all sessions, whisking-evoked activity for each region reflects a variable number of sessions, and thus of whisking events. We report the number of sessions and events considered for each region and mouse in Supplementary Fig. 1.
Statistical testing
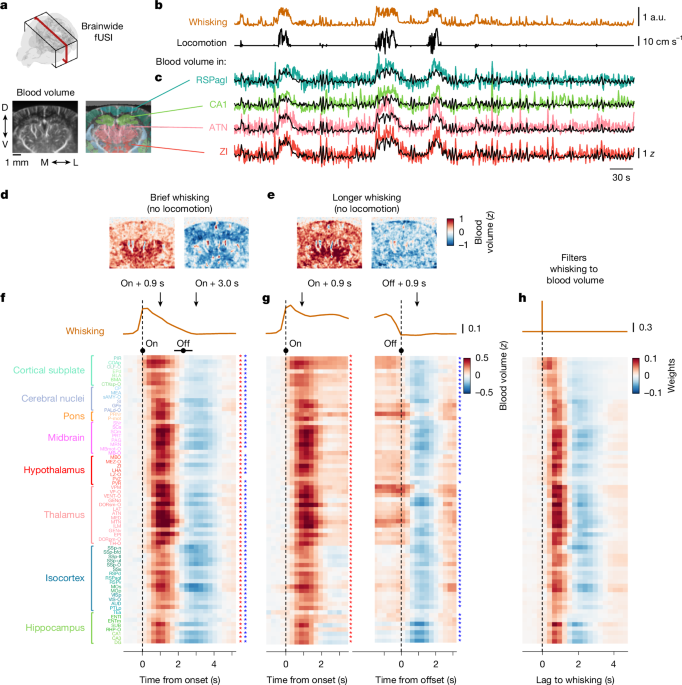
To assess the statistical significance of the changes in blood volume associated with whisking events (Fig. 1f–h and Extended Data Fig. 2), we used a permutation test. For each animal, we randomly shuffled the mapping between sessions and whisking and locomotion velocity, so that we aligned blood volume to the whisking events that occurred in another session. We repeated this procedure 100 times, thus obtaining a null distribution of event-evoked changes in blood volume. For each region and time point, we compared the ‘true’ value of blood volume change to the null distribution and defined the P value as the proportion of samples from the null distribution above (or below) the ‘true’ value. We corrected for multiple comparisons with the Benjamini–Hochberg procedure. For select time points (0.9 s and 3.0 s) relative to whisking events, we mark the regions that show significant (P < 0.05) increases or decreases in activity with an asterisk (Fig. 1f,g).
Predictions from whisking
To predict blood volume from whisking, we used cross-validated ridge regression. The lambda hyperparameter used for regularization was fitted through nested cross-validation. A feature matrix was constructed using lagged versions of the whisking trace, with 25 lags ranging from −1.8 s to 5.7 s in 0.3 s intervals, where positive lags mean that whisking precedes blood volume. This matrix was used to predict activity in each brain region, for each session. For each session, we allowed for a different non-linear scaling of whisking (the same across all regions). We ran predictions from whisking raised to the power of different values (from 0.1 to 1.2 with steps of 0.1) and selected the value providing the best prediction accuracy (averaged across regions). The median exponent across sessions was 0.4, so that this procedure gave more relative weight to small whisking events relative to the strong whisking associated for example with locomotion (Supplementary Fig. 2).
Combined fUSI and Neuropixels recordings
Imaging and recordings
For combined fUSI and neuronal recordings, we used our published dataset8 which combines Neuropixels recordings in visual cortex and hippocampus with fUSI in 5 C57/BL6 mice (4 male, 1 female) 9–12 weeks of age (see ref. 8 for details). The dataset comprises ten independent recording sessions, each corresponding to an acute Neuropixels insertion, and thus sampling different neurons. In each recording session, three to five different fUSI planes were sampled sequentially, each time repeating the same protocol, including spontaneous activity (grey screen) and visual stimulation (flickering checkerboards).
Power Doppler was recomputed from IQ data to match the pre-processing steps explained above. Blood volume was averaged within two regions of interest, corresponding to voxels immediately around the probe track and in either visual cortex or hippocampus (~50 voxels in each region). In five out of ten recordings, two probes were inserted bilaterally. In these recordings, for each region we averaged the corresponding fUSI signal across the two hemispheres and considered neurons across the two probes. Units selected were ‘good’ and ‘multi-unit’ clusters from Kilosort as in ref. 8. Blood volume, behavioural metrics and single-unit firing rate were all binned in 300 ms bins.
For display, the firing rate for each neuron was divided by the 95th percentile of firing rate across time.
Behaviour analysis
Mice were not on a running wheel so that there is no locomotion in this dataset. An infrared camera over the right side of their face was used to monitor behaviour. Whisking was measured as the motion energy in a region of interest focused on the whisker pad, binned in 300 ms windows, and normalized as previously (between first percentile and maximum). We also tracked the pupil using DeepLabCut72 and extracted pupil size (diameter), eye position and eye movements on the basis of markers on the top, bottom, left and right extremes of the pupil.
Definition of Arousal+ and Arousal– populations
We computed the Pearson correlation with whisking in two splits of the session. We defined neurons as Arousal+ if this correlation was above 0.05 on both splits of the data, and Arousal– if it was below −0.05 on both splits. We first z-scored the firing rate of each neuron throughout the session and then averaged these across neurons of each group (Arousal+, Arousal–, or all).
In the cortex, 3.4% of Arousal+ neurons were putative inhibitory, significantly less than Arousal– neurons (5.8%, P = 0.03, t-test across sessions). In addition, 44% of neurons in the infragranular layers were Arousal+ neurons, significantly more than in the supragranular layers (23%, P = 0.027, t-test across sessions).
To relate Arousal+ and Arousal– neurons to previously described ‘positive-delay’ and ‘negative-delay’ neurons48,49 (Fig. 3 and Extended Data Fig. 3), we aligned the z-scored average firing rate of all neurons to global events, which we operationalized as corresponding to whisking events We computed the centre of mass of each neuron relative to these events by summing the dot product of each neuron’s whisking-related activity (normalized between 0 and 1) and the corresponding lags and dividing it by the sum of its normalized whisking-related activity. We then computed principal component analysis on the z-scored (neurons × lags) matrix and looked at the first component (PC1), which mapped onto the correlation with whisking and thus well segregated Arousal+ and Arousal– neurons.
Predictions from firing rate
We fit filters to predict blood volume from the average firing rate(s) of either all neurons, or Arousal+ and Arousal– neurons using linear regression with Lasso regularization. We did so separately for each region and each fUSI plane within a recording session, using all corresponding time points (not only whisking bouts). We used lags ranging from −2.1 s to 6.6 s in 0.3 s intervals (30 values in total), where positive lags mean that firing rate precedes blood volume.
Since the resolution of our HRFs is 3.33 Hz, we could only characterize variations in blood volume that are slower than 1.67 Hz. This is not a limitation, because most of the power of changes in blood volume occur at lower frequencies. In our experiments, for instance, 84% of the power was below 1 Hz.
Cross-validation was done using folds of ~8 min, with nested cross-validation for the regularization hyperparameter. In each fold, a chunk of ~8 min of data was left out, and the rest of the session (including different states, not only whisking epochs) was used to fit the HRF weights. These weights were then used to predict the left-out data. This procedure was repeated to obtain cross-validated predictions for the whole session. Filters were then computed by averaging the weights obtained across cross-validation folds. The definition of Arousal+ and Arousal– neurons was done on the whole recording session and not included in the cross-validation loop. We report our results by first averaging the metrics (for example, coherence and filters) obtained in different fUSI planes within the same recording session (so, with the same neurons). We then report the mean, s.e.m. and statistical testing across the different, independent recording sessions (n = 10 for each region).
As shown in Extended Data Fig. 7, we also tested alternative models by fitting filters to predict blood volume from: (1) two random populations of neurons; (2) whisking; (3) Arousal+ or Arousal– neurons alone; (4) bulk firing rate and pupil size.
For the first analysis (two random populations), we matched the size of the populations with that of the Arousal+ and Arousal– neurons. This resulted in a prediction with a similar number of parameters as the combined model, thus controlling for the increased number of degrees of freedom of the combined vs. bulk model. We estimated the degrees of freedom for the different HRF models by counting the number of non-zero weights out of the 30 possible (from −2.1 to 6.6 s in 0.3 s steps). We did so for each cross-validation fold, and then averaged across folds in each session, and then across recording sessions. We find an average of 22.6 degrees of freedom for the bulk model, 40.7 for the combined model, and 41.3 for the random combined model. The latter yielded predictions even worse than the bulk model, so the increased degrees of freedom alone cannot account for the superior performance of the combined model.
Finally, we looked at predictions obtained by applying biphasic filters to the bulk firing rate (Supplementary Fig. 4). We tested four such filters, obtained by taking the weighted difference between the filter we obtained for Arousal+ neurons (weight k) and the filter we obtained for Arousal– neurons (weight 1 – k), with k taking the values 0, 0.33, 0.66 and 1. This was done for each recording session using the filters obtained for this specific session.
Fit evaluation
To compare whisking-evoked activity for different types of signals (firing rate, actual and predicted blood volume), we identified whisking events as previously, aligned the signal to these events and averaged across them. All traces were z-scored throughout the whole recording session prior to this alignment. The combined predictions showed higher correlations with the actual data than the bulk model, both in the visual cortex (average Pearson correlation of predicted versus actual whisking-evoked activity for short bouts: 0.88 versus 0.52, P = 10−3 with a t-test across n = 10 independent recording sessions), and the hippocampus (0.81 versus 0.62, P = 2 × 10−2).
To evaluate fit quality, we computed the coherence throughout the session between actual and (cross-validated) predicted blood volume, which is like correlation in the frequency domain. Specifically, we estimated the magnitude squared coherence estimate using Welch’s method with 15 s windows and 7.5 s overlap between windows. We also computed the coherence separately for different parts of the session (Extended Data Fig. 6): spontaneous activity (grey screen) versus visual stimulation (flickering checkerboards).
To compare different predictions, we ran paired t-tests between the average coherence between actual and predicted blood volume (for all frequency bands below 0.7 Hz) for pairs of predictions (n = 10 independent recording sessions per region).
Classification of arousal states
We used pupil size and eye movements to define four putative states: rapid-eye-movement (REM) sleep, non-rapid-eye-movement (NREM) sleep, quiet wake and active wake. We base this classification on previous work showing that pupil size alone well predicts the different sleep and wake states19. For each recording session, we plotted eye movements against pupil size and manually identified two critical points. Starting from intermediate pupil size, in which eye movements are scarce, we identified the points at which the levels of eye movements start increasing: (1) when going to smaller pupil size—probably reflecting REM sleep; (2) when going to higher pupil size—probably reflecting active wake. We then used the mid-point between these thresholds as an arbitrary transition from NREM sleep to quiet wake.
For each session, we defined 3 s intervals, measured the median of different metrics in each interval and binned them on the basis of the median pupil size within the corresponding interval. We defined an arousal index which corresponds to the interpolated pupil size for each session, so that the previously determined thresholds correspond to 0 (REM transition) and 1 (active wake transition). We then averaged metrics across sessions for similar values of the arousal index. This allowed us to combine different sessions, in which slightly different camera angles prevented us from directly comparing the absolute values of pupil size and eye movements. We validated our definition by looking at other behavioural metrics as a function of pupil size (eye position, whisking).
To evaluate prediction accuracy across states, we cut each recording session into 10 s intervals and assigned a state to each using the same method. We then computed the Pearson correlation between actual and predicted blood volume within each of these intervals and took the median correlation across all intervals corresponding to each state to obtain a prediction accuracy by state for each recording session. We then compared prediction accuracy across models in each state with a t-test across sessions (n = 10 independent recording sessions per region). We corrected for multiple comparisons using the Benjamini–Hochberg procedure.
As a control, we also fit different models for each state separately. In each recording session, for each state we fit different filters to predict blood volume for firing rate using only time points belonging to intervals assigned to this state. We reconstructed a single prediction for the whole session by combining the cross-validated predictions obtained for each state with these different filters.
Longitudinal Neuropixels recordings
To verify the stability of the correlation with whisking across days, we implanted chronic Neuropixels73 in two mice (one in visual cortex, one in somatosensory cortex) and tracked neurons across days using UnitMatch74. We then compared the correlation with whisking across recording sessions for pairs of matched neurons, depending on how many days separated them. During the recordings, mice were allowed to run on the wheel and were either facing a dark screen or presented with natural images. We excluded sessions in which the mouse ran more than 90% of the time.
Brainwide Neuropixels recordings
We used Neuropixels recordings in the mouse brain performed in our consortium of laboratories as detailed elsewhere40. In the present analysis we included all sessions in which both video recordings and passive periods were available (n = 324 sessions). We focused our analysis on good units (n = 18,791 neurons) recorded in the passive periods, which occurred after the mice had performed the task.
In these experiments, mice were head-fixed onto a fixed platform (not a running wheel), so there was no locomotion. Whisking was measured as the motion energy in a region of interest focused on the whisker pad, averaged across 1 to 2 cameras, binned in 300 ms windows, and normalized as previously (between first percentile and maximum).
We mapped neurons recorded across different mice and sessions to similar brain regions as measured in fUSI. To do so, we started with each neuron’s label in the fine-grained Allen CCF labels. We then sorted the target brain regions by hierarchical level, starting with the lowest. For a given region, we included all neurons belonging to descendants of a region except if they had been already included in another child region. For example, VENT could contain neurons belonging to the subregions of VL, VAL but not VP, since the latter was also a target region of interest. This procedure aimed to mimic what had been done for fUSI voxels, as well as ensuring that each neuron only featured in one target region. We excluded from further analysis regions containing less than 50 neurons for which spikes were detected during the passive periods.
Spikes of each neuron were binned in the same 300 ms bins as whisking. For each neuron, we z-scored the firing rate of each neuron across time within its recording session, detected whisking events as previously and aligned this signal to the whisking events. For each type of event (brief or longer bouts), we only included neurons for which at least five bouts were detected.
We computed the Pearson correlation with whisking in two splits of the session. We defined neurons as Arousal+ if this correlation was above 0.05, and as Arousal– if it was below −0.05. To count the number of reliable Arousal+ and Arousal– neurons in each region, we considered only neurons meeting the criterion in two splits of the session. We also computed an index measuring the relative bias to Arousal+ neurons (as used in Fig. 6d) in each region by taking the difference between the number of Arousal+ neurons and Arousal– neurons over their sum.
To establish the stability of our population’s definition across behavioural contexts, we compared for each neuron the correlation with whisking computed on the passive part of the session versus the active part—that is, when the mice performed the task. We then computed the correlation of these values across neurons for each region (Extended Data Fig. 9).
To look at whisking-related firing rate, we classified neurons on the basis of one split and computed whisking-related changes in the other split (Figs. 5 and 6 and Extended Data Fig. 8). All whisking-related changes are baseline-subtracted (for each region), using a window of 0.5–2 s before whisking onset.
Predicting whisking-related brainwide blood volume
To predict changes in blood volume (measured in the brainwide fUSI experiments) from the firing of neurons (measured in the brainwide Neuropixels recordings), we used the HRFs found from the combined fUSI–Neuropixels recordings, averaged across visual cortex and hippocampus. We assigned a neuron to Arousal+ or Arousal– groups on the basis of half of the recording session and used its average firing rate on whisking bouts on the other half of the session. For each region, we then averaged this cross-validated firing rate across all Arousal+ or Arousal– neurons and convolved it with their respective filters. We first rescaled the Arousal+ filter by the ratio of the proportion of Arousal+ neurons in this region to the average proportion of Arousal+ neurons in the simultaneous fUSI–Neuropixels recordings (across hippocampus and visual cortex). We did the same for Arousal– filters and summed the contributions from both filters. For the predictions from bulk firing rate, we used the corresponding filter with the same scaling for all regions and used firing for the same half as for the other prediction. Finally, we applied a global scaling factor to each type of prediction (the same across all regions). This scaling factor was obtained by minimizing the mean squared error between the predicted and actual blood volume −1.2 to 5 s from whisking onset, for brief and long bouts, across all regions. We similarly computed the mean squared error for each region separately and compared those between the combined and bulk prediction (Fig. 6c).
Visualization
The 3D brain visualizations were generated using Urchin.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.