Cell lines
HT29 (HTB-38), T84 (CCL-248) and 293T (CRL-3216) cells were originally obtained from the ATCC. HT29-Cas9 cells were generated as previously described21. All cell lines were grown at 37 °C and 5% CO2. HT29 cell lines were propagated in McCoy’s 5A medium, T84 lines in DMEM/F-12 and 293T cells in DMEM; all supplemented with 10% FBS. Cell lines were routinely tested for mycoplasma contamination and found to be negative.
Antibodies for immunoblotting, immunoprecipitation and flow cytometry
The following mouse monoclonal antibodies were purchased from the indicated vendors and used at the indicated dilutions: β-actin (A5441, Sigma, 1:2,000), claudin-4 (sc-376643, Santa Cruz Biotechnology, 1:200), Alexa Fluor 647-conjugated claudin-4 (FAB4219R, R&D, 1:100), His tag (MAB050, R&D Systems, 5 μg ml−1), GFP (MA1-952, Invitrogen, 1:2,000). The following rabbit monoclonal antibodies were purchased from the indicated vendors: claudin-3 (83609S, Cell Signaling Technology, 1:1,000), Na/K ATPase α1 (D4Y7E, Cell Signaling Technology, 1:1,000). The following rat monoclonal antibodies were purchased from the indicated vendors: E-cadherin ectodomain (14-3249-82, Invitrogen, 1:250), PE-conjugated E-cadherin ectodomain (147304, BioLegend, 1:100). The following rabbit polyclonal antibodies were purchased from the indicated vendors: claudin-4 (36-4800, Invitrogen; see ‘Co-immunoprecipitation’, Methods), claudin-2 (51-6100, Invitrogen; see ‘Co-immunoprecipitation’, Methods). Rabbit polyclonal antiserum against BFT was produced as previously described61 and was used at 1:1,000. The following secondary antibodies were purchased from the indicated vendors: Alexa Fluor 488-conjugated anti-mouse IgG (A-11001, Invitrogen, 5 μg ml−1), HRP-conjugated anti-rabbit IgG (7074, Cell Signaling Technology, 1:1,000), HRP-conjugated anti-rat IgG (7077, Cell Signaling Technology, 1:1,000), HRP-conjugated anti-mouse IgG (R1005, Kindle Biosciences, 1:1,000).
Plasmids and molecular cloning
The plasmids GFP-C4 (pCMV::EGFP-hCLDN4) and GFP-claudin-2 (pCMV::EGFP-hCLDN2) expressing N-terminally EGFP-tagged human claudins from the CMV promoter were obtained as a gift from J. Turner (Harvard Medical School).
The plasmids GFP-C3 (pCMV::EGFP-hCLDN3), GFP-claudin-4(ΔCTD) (pCMV::EGFP-hCLDN4ΔCTD), GFP-claudin-4(ECS13) (pCMV::EGFP-hCLDN4ECS1-3), GFP-claudin-4(ECS23) (pCMV::EGFP-hCLDN4ECS2-3) and Ecad-mCherry (pCMV::hCDH1-mCherry) were constructed using In-Fusion cloning (Takara Bio). GFP-C3 was generated by replacing the claudin-4 sequence of GFP-C4 with human claudin-3 complementary DNA. GFP-claudin-4(ΔCTD) was generated by excising the claudin-4 amino acids Asn184–Val209 from GFP-C4. GFP-claudin-4(ECS13) was generated by replacing the claudin-4 amino acids Met29–Arg81 from GFP-C4 with the claudin-3 amino acids Met28–Arg80. GFP-claudin-4(ECS23) was generated by replacing the claudin-4 amino acids Thr139–Met160 from GFP-C4 with the claudin-3 amino acids Ser138–Met159. Ecad-mCherry was generated by replacing the mouse E-cadherin sequence from the plasmid Murine E-cadherin mCherry (Addgene, 71366) with the human E-cadherin sequence from the plasmid E-cadherin-GFP (Addgene, 28009). The plasmids GFP-claudin-4(T33S) (pCMV::EGFP-hCLDN4T33S), GFP-claudin-4(V41I) (pCMV::EGFP-hCLDN4V41I) and GFP-claudin-4(T45N) (pCMV::EGFP-hCLDN4T45N) were constructed using the QuikChange II Site-Directed Mutagenesis Kit (Agilent Technologies).
Recombinant proteins
His6-tagged BFT
The BFT-2 isoform (herein referred to as BFT) was used for all experiments. Recombinant proBFT (residues Asp24–Asp397) with a C-terminal His6-tag was expressed from a modified pET30a vector in Escherichia coli Rosetta(DE3). Overnight cultures of E. coli were subcultured at a ratio of 1:50 into 100 ml of lysogeny broth with kanamycin and chloramphenicol, grown at 37 °C and 240 rpm until they reached an optical density (OD)600 of 0.4–0.6 and then induced with 0.5 mM isopropyl β-d-1-thiogalactopyranoside (IPTG) at room temperature overnight. The following morning, cells were collected by centrifugation at 3,000g for 15 min at 4 °C. Pellets were resuspended in ice-cold buffer A (50 mM Tris-HCl, 500 mM NaCl, 20 mM imidazole, pH 7.5) and sonicated to lyse cells. Cell debris was pelleted by centrifugation at 17,200g at 4 °C for 45 min. The supernatant containing recombinant proBFT was passed through a 0.22-μm filter and loaded onto a HisTrap column (Cytiva Life Sciences) pre-equilibrated with 10 column-volumes (CV) of buffer A. The column was washed with 5 CV of buffer A, then protein was eluted with 5 CV of buffer A containing 250 mM imidazole. Fractions containing the A280 peak were pooled and dialysed into PBS overnight. The resulting proBFT was trypsinized (T6567, Sigma) at a 1:30 w/w ratio of trypsin:proBFT to liberate mature BFT, which was subsequently re-purified on a HisTrap column and dialysed into PBS overnight as above. Concentration was determined by Qubit protein assay (Pierce), purity was assessed by SDS–PAGE followed by Coomassie blue staining (Extended Data Fig. 1a) and purified BFT was aliquoted and stored at −80 °C until use.
Untagged BFT
Recombinant proBFT (residues Asp24–Asp397) with an N-terminal His6-tag was expressed from a modified pET28a vector in E. coli Rosetta(DE3). Overnight cultures of E. coli were subcultured at a ratio of 1:100 into 500 ml of lysogeny broth with kanamycin and chloramphenicol, grown at 37 °C and 240 rpm until they reached an OD600 of 0.4–0.6 and then induced with 0.5 mM IPTG at room temperature overnight. The following morning, cells were collected by centrifugation at 4,000g for 20 min at 4 °C. Pellets were resuspended in ice-cold buffer A (50 mM Tris-HCl, 500 mM NaCl, 20 mM imidazole, pH 7.5) and lysed using a Cell Disruptor (Constant Systems). Cell debris was pelleted by centrifugation at 30,000g at 4 °C for 40 min. The supernatant containing recombinant proBFT was incubated with HisPur Ni-NTA resin (Thermo Fisher) at 4 °C for 1 h, with stirring. The mixture was then poured onto a gravity chromatography column. The flowthrough was allowed to drain and the resin was washed with 10 CV of buffer A, and then protein was eluted with 5 CV of buffer A containing 300 mM imidazole. The eluate containing proBFT was buffer exchanged into 50 mM Tris-HCl, 150 mM NaCl, pH 7.5 using a PD-10 desalting column (Cytiva), then trypsinized at a 1:30 w/w ratio of trypsin:proBFT for 3 h at room temperature to liberate mature BFT. The trypsinization reaction was stopped using 1 mM PMSF, and the trypsinized proBFT was concentrated and buffer exchanged into 20 mM Tris-HCl, 20 mM NaCl, pH 7.5 using an ultrafiltration device. The resulting mixture was injected onto a Mono Q 5/50 GL column (Cytiva) and mature BFT was eluted using a gradient from 20 to 500 mM NaCl. Purity was assessed by SDS–PAGE followed by Coomassie blue staining (Extended Data Fig. 1b). Fractions containing pure mature BFT were pooled, aliquoted, snap frozen in liquid nitrogen and stored at −80 °C until use. The molecular weight and N-terminal sequence of untagged BFT were verified by SEC-MALS and N-terminal sequencing (Supplementary Fig. 10).
CLN4sol and C4sol(C2-ECS)
Recombinant soluble claudins with C-terminal His6-tags were expressed from a modified pET28a vector in E. coli Rosetta(DE3). Overnight cultures of E. coli were subcultured at a ratio of 1:100 into 500 ml of lysogeny broth with kanamycin and chloramphenicol, grown at 37 °C and 240 rpm until they reached an OD600 of 0.4–0.6 and then induced with 0.5 mM IPTG at 28 °C overnight. The following morning, cells were collected by centrifugation at 4,000g for 20 min at 4 °C. Pellets were resuspended in ice-cold buffer A (50 mM Tris-HCl, 500 mM NaCl, 20 mM imidazole, pH 7.5) and lysed using a Cell Disruptor (Constant Systems). Cell debris was pelleted by centrifugation at 30,000g at 4 °C for 40 min. The supernatant containing recombinant soluble claudin was incubated with HisPur Ni-NTA resin (Thermo Fisher) at 4 °C for 1 h, with stirring. The mixture was then poured onto a gravity chromatography column. The flowthrough was allowed to drain and the resin was washed with 10 CV of buffer A, and then protein was eluted with 5 CV of buffer A containing 300 mM imidazole. The eluate containing soluble claudin was buffer exchanged into 50 mM Tris-HCl, 150 mM NaCl, pH 7.5 using a PD-10 desalting column (Cytiva). Soluble claudins were stored at 4 °C until use. Sequences of all recombinant proteins are listed in Supplementary Table 5.
Screening CRISPR-KO libraries
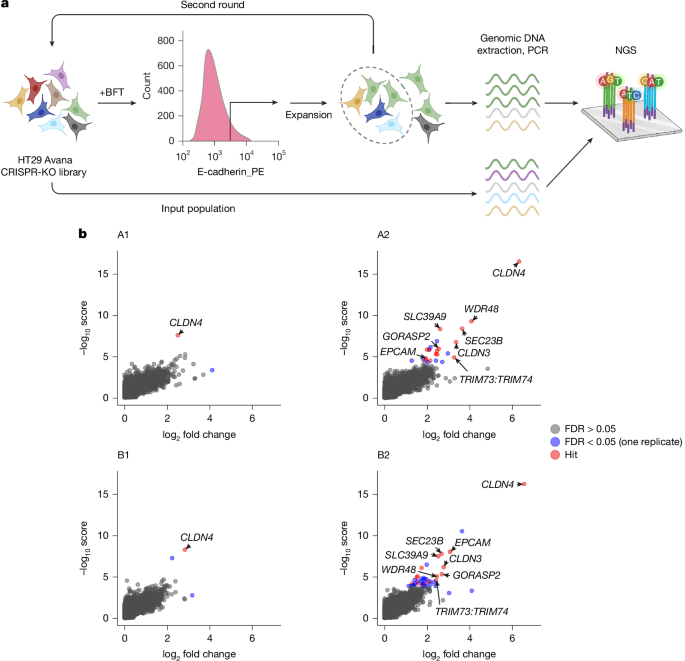
The Avana CRISPR gRNA library in HT29-Cas9 cells, containing four gRNAs targeting 18,675 annotated human protein-coding genes, was generated in two replicates (A and B) as previously described21. Libraries A and B were maintained at more than 500× representation in T225 flasks. Two independent screens, consisting of two rounds each of BFT selection, were performed on libraries A and B. One confluent T225 flask (around 120 million cells) of each library was treated with His6-tagged BFT (1.5 ng ml−1, 70 pM) in serum-free McCoy’s 5a medium for 1 h at 37 °C and 5% CO2. BFT-treated flasks were rinsed twice with Dulbecco’s PBS (DPBS) and treated with 12 ml of Cellstripper (Corning) for 40 min at 37 °C. Then, 15 ml of pre-warmed 2% FBS in DPBS was added and cells were gently dissociated by pipetting up and down. Cells were pelleted at 200g for 5 min, washed with 2% FBS/DPBS, pelleted again and resuspended in pre-warmed digestion solution (DPBS with 2% FBS, 100 μg ml−1 DNase I, 500 μg ml−1 collagenase D, 5 mM MgCl2 and 2 mM CaCl2) to further dissociate cell clumps. Cells were pelleted and washed with ice-cold FACS buffer (DPBS with 2% FBS and 2 mM CaCl2), and then resuspended in ice-cold FACS buffer with 1:100 PE-conjugated E-cadherin antibody and stained for 40 min at 4 °C with end-over-end rotation.
Cells were sorted on an SH800S Cell Sorter (Sony) using a 70-μm nozzle and the top 10% of live, single cells by surface E-cadherin staining were collected. Collected cells were plated, expanded and subjected to the same procedure for the second round of selection. After each round of selection and expansion, genomic DNA was extracted from 10 million cells using the Blood and Cell Culture DNA Kit (Qiagen) and DNA fragments containing the gRNA sequences were amplified by PCR to attach Illumina adaptor and barcode sequences as previously described34 (Extended Data Fig. 2a). The same DNA extraction and PCR were performed on the unselected libraries A and B for comparison of gRNA abundances. PCR products were gel purified and sequenced on an Illumina MiSeq. Reads were trimmed to eliminate adaptor sequences using CLC Genomics Workbench (Qiagen) and the rank and statistical significance of genes after each round of selection was determined using MAGeCK22. The null distribution of the 1,000 Avana control gRNAs was used to compute P values for all genes using the MAGeCK ‘test’ function with the option ‘–norm-method control’. Hits were defined as genes significant at an FDR of 0.05 in round 2 of both screens, A and B. Full gene-level datasets for each screen replicate and round are available as Supplementary Tables 1–4.
Stable cell line generation
The following gRNA sequences were designed using CRISPick (https://portals.broadinstitute.org/gppx/crispick/public) and were cloned into either the plasmid lentiGuide-Puro (Addgene, 52963, to transduce HT29-Cas9 cells) or lentiCRISPR v2 (Addgene, 52961, to transduce T84 cells): CAAGGCCAAGACCATGATCG (CLDN4) and GGGCAACGCGCAGCACACGA (CLDN3). Lentiviruses were produced through co-transfection of the corresponding gRNA plasmid with psPAX2 (Addgene, 12260) and pCMV-VSV-G (Addgene, 8454) into 293T cells seeded in six-well plates as previously described21. HT29-Cas9 or T84 cells were transduced with gRNA-expressing lentiviruses and mixed populations of knockout cells were selected with puromycin (1 µg ml−1 for HT29-Cas9 cell lines or 2 µg ml−1 for T84 cell lines). C4/C3-KO cell lines were produced by transduction of polyclonal C4 single-knockout lines with lentiGuide-Hygro (Addgene, 139462) harbouring the CLDN3 gRNA, followed by selection with hygromycin B (800 µg ml−1 for C4-KO T84 cells and 400 µg ml−1 for C4-KO HT29 cells). Monoclonal knockout HT29-Cas9 cell lines were isolated by limiting dilution and used for all experiments. Polyclonal T84 knockout cell populations were further selected by FACS with surface claudin-4 and claudin-3 staining to remove remaining claudin-positive cells, and the resulting polyclonal knockout cell lines were used for all experiments. C4-KO T84 cells required two rounds of sorting to remove all claudin-4-positive cells. Cells derived from the first round of sorting, used for monolayer experiments with basolateral BFT, are referred to as C4-pKO (Extended Data Fig. 4b–d). Disruption of the targeted open reading frames was confirmed by Sanger sequencing and immunoblot.
The C4-KO::GFP-C4 HT29 cell line was constructed through delivery of a gRNA-resistant GFP-C4 gene into C4-KO HT29 cells by lentiviral transduction. The lentiviral transfer plasmid pLenti-GFP-C4-rescue encoding EGFP-claudin-4, harbouring silent mutations in the protospacer adjacent motif (PAM) and adjacent protospacer sequence that prevent targeting by the indicated CLDN4 gRNA sequence, was constructed using the QuikChange II Site-Directed Mutagenesis Kit and In-Fusion cloning. First, the gRNA-resistant plasmid GFP-claudin-4(mutPAM) (pCMV::EGFP-hCLDN4mutPAM) was constructed by site-directed mutagenesis of GFP-C4. Then, pLenti-GFP-C4-rescue was generated by replacing the EGFP stop codon in the lentiviral transfer plasmid pLenti-CMV-GFP-Hygro (Addgene, 17446) with the gRNA-resistant claudin-4 sequence from GFP-claudin-4(mutPAM). Lentivirus was produced through co-transfection with psPAX2 and pCMV-VSV-G into 293T cells as above and used to transduce C4-KO HT29 cells, and mixed populations of transduced cells were selected with 800 µg ml−1 hygromycin B.
BFT cell rounding assay
Cells seeded in 96-well plates were treated with His6-tagged BFT (0.8 ng ml−1, 37 pM) in serum-free culture medium at 37 °C and 5% CO2. Phase contrast images (and green fluorescence images, for C4-KO::GFP-C4 cells) of live cells were acquired at the indicated timepoint (Nikon ECLIPSE Ts2, ×20 objective).
Cellular BFT binding assays
Cells seeded in 12-well plates were rinsed once with serum-free culture medium, treated with His6-tagged BFT (100 ng ml−1, 4.6 nM) in serum-free culture medium at 37 °C and 5% CO2 for 40 min and rinsed twice with ice-cold PBS to remove unbound BFT. Cells were then either dissociated into single cells for bound BFT detection by flow cytometry or lysed directly in the plate for detection by immunoblot. For flow cytometry analysis of BFT binding, cells were dissociated using TrypLE express for 10 min at 37 °C, stained with a mouse anti-His6 primary antibody followed by an anti-mouse Alexa Fluor 488 secondary antibody and analysed on an Attune NxT flow cytometer (Thermo Fisher). Flow cytometry data analysis was performed in FCS Express 7 (De Novo Software). In all cases, events were gated on forward and side scatter to select a uniform cell population and exclude debris and doublets. For 293T cell experiments in which the cleaved E-cadherin ectodomain was detected in the corresponding culture supernatants, 400 μl of each culture supernatant was concentrated by ultrafiltration in a 10-kDa molecular weight cutoff spin column (Amicon) and equally concentrated supernatant volumes were analysed by immunoblot. Immunoblot methods are described below. For flow cytometry gating strategy, see Supplementary Fig. 7.
Transfection
293T cells were seeded at either 2.7 × 105 live cells per well in 12-well plates or 2.5 × 104 per well in 96-well plates coated with poly-d-lysine (Sigma) the day before transfection. Plasmid DNA (1 μg per 12-well plate well) diluted in OptiPro-SFM serum-free medium (Thermo Fisher) to a total volume of 97 μl was mixed with 3 μl of TransIT-LTI transfection reagent (Mirus Biosciences) and incubated for 20 min at room temperature to form transfection complexes. For 96-well plate transfections, the same described quantities of plasmid DNA and TransIT-LTI were used to transfect 10 wells. In the case of co-transfection with GFP-claudin and Ecad-mCherry plasmids, the plasmids were added in a 1:1 molar ratio to the same total DNA mass, such that the ratio of total DNA mass to TransIT-LTI volume was maintained. Transfection complexes (100 μl per 12-well plate well or 10 μl per 96-well plate well) were then added dropwise to adherent 293T cells. Unless otherwise specified, experiments were performed at 20 h post transfection.
Co-immunoprecipitation
HT29-Cas9 cells seeded in a 10-cm dish were rinsed once with serum-free culture medium, treated with His6-tagged BFT (100 ng ml−1, 4.6 nM) at 37 °C and 5% CO2 for 40 min and rinsed twice with ice-cold PBS to remove unbound BFT. Cells were then lysed in immunoprecipitation lysis buffer (25 mM Tris-HCl, 150 mM NaCl, 1 mM EDTA, 1% NP-40 and 5% glycerol, pH 7.4), incubated on ice for 5 min and clarified by centrifugation at 13,000g at 4 °C. Protein concentration was determined by BCA assay (Pierce) and 2 mg of clarified lysate per immunoprecipitation reaction was pre-cleared by incubation with 80 μl of Control Agarose Resin (Pierce) for 1 h at 4 °C to reduce non-specific binding. Each pre-cleared lysate was then incubated overnight at 4 °C with 20 μl of Protein A/G Plus Agarose (Pierce) crosslinked with 10 μg of rabbit polyclonal antibody against either claudin-4 or claudin-2 by disuccinimidyl suberate. The following day, resins were washed three times with Tris-buffered saline and proteins were eluted at pH 2.8 for 5 min at room temperature. Lysate and immunoprecipitated fractions were analysed by immunoblot as described below.
E-cadherin flow cytometry
HT29-Cas9 cell lines seeded in 96-well plates, or 293T cells seeded in 96-well plates and transfected with the indicated plasmids, were treated with the indicated His6-tagged BFT concentrations in serum-free culture medium for the indicated time. Cells were rinsed twice with 200 μl of PBS per well, then dissociated by incubation with 100 μl of Cellstripper (Corning) at 37 °C. An equal volume of 2% FBS in DPBS was added and cells were gently dissociated by pipetting up and down. Cells were transferred to a round-bottom 96-well plate, pelleted at 300g for 3 min, washed once in ice-cold FACS buffer and resuspended in antibodies diluted in FACS buffer (PE-E-cadherin and Alexa Fluor 647-claudin-4, 1:100) for 30 min at 4 °C in the dark with gentle rocking. Cells were washed twice with ice-cold FACS buffer, resuspended in 100 μl of ice-cold FACS buffer per well and analysed on either an Attune NxT (ThermoFisher) or Aurora (Cytek) flow cytometer via the attached autosampler.
All flow cytometry data analysis was performed using FCS Express 7 (De Novo Software). Events were gated on forward and side scatter to select a uniform cell population and exclude debris and doublets. As C4-KO::GFP-C4 cells were found to have a lower average claudin-4 expression level than WT HT29-Cas9 cells, these cells were additionally gated to exclude cells with surface claudin-4 levels less than the 5th percentile of WT HT29-Cas9 cells. The log10-transformed PE median fluorescence intensity (MFI) values for each well were determined as a readout of surface E-cadherin level. Normalized surface E-cadherin levels for each well were calculated on the basis of non-toxin-treated, identically stained positive control wells and unstained negative control wells using the formula Ecadnorm, i = (MFIi − MFIneg)/(MFIpos − MFIneg), where MFIi is the log10-transformed MFI of well i, and MFIneg and MFIpos are the log10-transformed MFIs of the corresponding negative and positive control wells, respectively. Curve fitting of the normalized surface E-cadherin levels was performed in GraphPad Prism 10 using the model ‘[Inhibitor] vs. response − Variable slope (four parameters)’. Fold resistance values were calculated by dividing the obtained half-maximal inhibitory concentration (IC50) values by the IC50 value of WT HT29-Cas9 cells. For flow cytometry gating strategies, see Supplementary Figs. 6, 8 and 9.
Immunoblotting
Unless otherwise indicated, cells were lysed directly in tissue culture plates in NP-40 buffer (1% v/v Igepal CA-630, 50 mM Tris-HCl, 150 mM NaCl, pH 7.5) with an added cOmplete Mini protease inhibitor tablet (Roche). Cells were solubilized with orbital shaking for 30 min at 4 °C, debris were pelleted by centrifugation at 17,200g for 30 min at 4 °C and clarified lysates were stored as single-use aliquots at −80 °C until used for immunoblotting. Clarified lysates were thawed on ice and protein concentrations were determined using a BCA protein assay kit (Pierce). Lysates (20 μg of protein) were heated in 1x NuPAGE LDS sample buffer (Invitrogen) and 100 mM DTT at 70 °C for 20 min and loaded into NuPAGE 4–12% Bis-Tris Mini protein gels (Invitrogen), and electrophoresis was performed at 200 V for 35 min in NuPAGE MES SDS running buffer (Invitrogen). Proteins were transferred to nitrocellulose membranes at 100 V for 1 h in a wet tank filled with NuPAGE transfer buffer (Invitrogen) supplemented with methanol (10% if transferring one gel, 20% if transferring two gels). Evenness of transfer was checked by Ponceau S staining and stain was removed by rinsing in PBS-T (PBS with 0.05% Tween 20, pH 7.4). Blots were blocked in blocking buffer (5–10% dry milk in PBS-T) for 1 h at room temperature, then cut horizontally into pieces if multiple proteins were to be probed. Blots were incubated with primary antibodies diluted in blocking buffer for 1 h at room temperature, washed three times for 5 min each in PBS-T, then incubated with HRP-conjugated secondary antibodies diluted in blocking buffer for 1 h at room temperature, and then finally washed again three times for 5 min each in PBS-T. Blots were developed in SuperSignal West Pico PLUS chemiluminescent substrate (Thermo Fisher) for 5 min, placed in plastic folders and imaged on an iBright CL1500 (Invitrogen).
Blots were stripped and re-probed only if two proteins with very similar electrophoretic mobilities needed to be detected (in this case, either untagged claudin-3 and untagged claudin-4 or untagged claudin-4 and BFT). Blots were stripped in Restore stripping buffer (Thermo Fisher) for 30 min at room temperature, washed three times for 5 min each in PBS-T, re-blocked and re-probed as indicated above.
BFT and CLNsol pulldown assay
Untagged BFT (20 μg) alone or mixed with either His6-tagged CLN4sol or C4sol(C2-ECS) (20 μg) was incubated in binding buffer (20 mM Tris-HCl, 250 mM NaCl, 5 mM imidazole, 5% glycerol, pH 7.5) in 200-μl total volume for 1 h on ice. Incubated mixtures were centrifuged at 16,100g and 4 °C for 10 min. Post centrifugation, 20 μl of the input mixtures was removed for analysis. Then, 50 μl of HisPur Ni-NTA agarose slurry (Thermo Fisher) was added to the remaining mixtures. After 30 min of tube rotation at 4 °C, agarose was collected by centrifugation (4,000g, 2 min) and washed three times with wash buffer (20 mM Tris-HCl, 250 mM NaCl, 20 mM imidazole, 5% glycerol, pH 7.5). Bound proteins were eluted in 30 μl of elution buffer (20 mM Tris-HCl, 250 mM NaCl, 300 mM imidazole, 5% glycerol, pH 7.5). Input and bound fractions were analysed by SDS–PAGE on handmade 20% acrylamide gels followed by Coomassie blue staining. Input mixtures were also analysed by immunoblot using a mouse penta-His primary antibody (Qiagen, 1:5,000) and IRDye 800CW-conjugated goat anti-mouse secondary antibody (LI-COR, 1:10,000). The fluorescent immunoblot was imaged on an Odyssey Fc imager (LI-COR).
T84 monolayer culture and TER measurements
T84 cell lines were seeded at 165,000 live cells per well in 0.33-cm2 polyester Transwell inserts with a 0.4-μm pore size (Corning, 3470). Apical and basolateral media were replaced with fresh complete medium every 2 days and experiments were performed 7–9 days post seeding. The day before the experiment, the apical and basolateral media were changed to serum-free medium. The following day, initial TER measurements were recorded in a tissue culture hood with an EVOM2 (World Precision Instruments) using an STX2 electrode. The apical or basolateral medium was then immediately replaced with His6-tagged BFT (100 ng ml−1 for apical treatments or 2 ng ml−1 for basolateral treatments) in serum-free medium and cells were returned to the CO2 incubator. TER measurements were taken at the indicated timepoints after BFT treatment. For time course analysis, TER measurements were normalized by dividing them by the corresponding baseline TER value.
HT29 cell BFT neutralization assay
Untagged BFT at the indicated concentration was pre-incubated with a 100-fold molar excess of either CLN4sol or C4sol(C2-ECS) for 1 h at room temperature, then applied to HT29 cells for 30 min at 37 °C. Cells were then lysed for E-cadherin detection by immunoblot.
BFT caecal injection assay
All procedures were conducted in accordance with the guidelines approved by the Boston Children’s Hospital Institutional Animal Care and Use Committee (no. 00002233). CD-1 outbred mice (5 weeks old, body weight around 17–20 g) were purchased from Envigo (030-US) and only the female mice were used. Mice were group housed with up to five animals per cage on a 12:12-h light:dark cycle, with free access to food and drink in individually ventilated specific-pathogen-free cages (temperature 22 ± 2 °C; relative humidity 40–60%). All mice used were healthy and were not involved in any previous procedures or drug treatments. For BFT caecal injection experiments, untagged BFT (10 μg), CLN4sol (300 μg) or C4sol(C2-ECS) (300 μg) was incubated either alone or mixed together in 200 μl of PBS for 30 min at room temperature. Mice were anaesthetized with isoflurane and the caecum was exposed via laparotomy. The BFT, CLNsol or BFT:CLNsol mixture was injected directly into the caecum, followed by wound closure with stitches. Buprenorphine (0.1 mg kg−1) was injected subcutaneously immediately after surgery for analgesia. Mice were allowed to recover, and 4 h after surgery, the mice were euthanized. The caecum was collected, rinsed with PBS to remove the contents and fixed with 10% formalin overnight for further histology and immunofluorescence analysis.
The fixed caecum was dehydrated with 70% ethanol, 96% ethanol, 100% ethanol, then xylene (each step twice for 30 min), then infiltrated with melted paraffin at 58 °C for 45 min and embedded into the block. The tissue blocks were sectioned into 6-μm-thick sections. For H&E staining, sections were heated at 42 °C for 30 min and treated with xylene (twice for 5 min), 100% ethanol (thrice for 1 min), tap water (1 min), Gill 3 haematoxylin (2 min), tap water (2 min), eosin (3 min), 100% ethanol (thrice for 1 min) and xylene again (twice for 2 min). Sections were mounted with DPX mountant (Sigma) for histology analysis. Stained sections were coded and scored blindly. For immunofluorescence analysis, sections were heated at 42 °C for 30 min, then deparaffinized with xylene (twice for 10 min). The sections were air dried, then rehydrated with 100% ethanol, 70% ethanol, 50% ethanol, 30% ethanol, water and PBS (each step for 5 min). The rehydrated sections were immersed in slightly boiling 10 mM sodium citrate buffer pH 6.0 for 10 min, washed with PBS for 5 min, then incubated with blocking buffer (5% FBS, 0.1% Triton X-100 in PBS) for 1 h and with 1:100 rabbit anti-E-cadherin antibody overnight (3195T, Cell Signaling Technology). The next day, sections were washed three times with PBS and incubated with 1:100 DyLight488-conjugated donkey anti-rabbit IgG (406404, BioLegend) for 1 h at room temperature. Then the sections were washed three times with PBS and mounted with DAPI Fluoromount-G (SouthernBiotech). Images were recorded with an LSM880 upright confocal system (Zeiss) and processed in ImageJ.
H&E-stained sections were scored in a blinded fashion for epithelial shedding and oedema (Supplementary Fig. 11). Measurement of caecal crypt length was performed as described in ref. 62. Specifically, ten distinct crypts were randomly chosen from each section. Only crypts with an intact architecture spanning from the crypt base to the luminal surface were selected, and the selected crypts were distributed as evenly as possible across the entire caecal mucosa. Crypt length was measured using ImageJ. The average value of the ten measurements was used as the readout for each mouse.
Analytical SEC and SEC-MALS
BFT and CLN4sol were incubated in SEC buffer (50 mM Tris-HCl, 150 mM NaCl, pH 7.5) supplemented with 1 mM PMSF, either alone, in a 1:4 mixture for 1 h (analytical SEC) or in a 1:1 mixture overnight (SEC-MALS) at 4 °C. Samples were centrifuged at 16,100g for 10 min at 4 °C, then injected onto a Superdex 200 Increase 10/300 column (Cytiva) pre-equilibrated in SEC buffer. The column was maintained at 4 °C (analytical SEC) or 18 °C (SEC-MALS) and run at a flow rate of 0.5 ml min−1. Analytical SEC was run using an AKTA purifier (Cytiva) and SEC-MALS was run using a Prominence HPLC system (Shimadzu) connected to a DAWN HELEOS II multi-angle light scattering detector and an Optilab T-REX refractive index detector (Wyatt Technology). Data processing and analysis were performed in ASTRA 7 (Wyatt Technology). A dn/dc value of 0.185 ml g−1 (typical for proteins in aqueous buffer) was assumed for calculations and BSA (Wyatt Technology) was used as a protein standard.
Surface biotinylation
293T cells seeded in 12-well plates and transfected with the indicated GFP-claudin plasmid were rinsed gently with PBS++ (PBS with 2.5 mM CaCl2 and 1 mM MgCl2). Sulfo-NHS-biotin (0.5 mg ml−1 in PBS++, Thermo Fisher) was added directly to adherent cells and surface biotinylation was performed for 30 min at 4 °C with gentle rocking. Remaining unreacted sulfo-NHS-biotin was quenched by gently rinsing cells twice with ice-cold Tris-buffered saline and cells were lysed in NP-40 buffer. Cells were solubilized with orbital shaking for 30 min at 4 °C, debris were pelleted by centrifugation at 17,200g for 30 min at 4 °C and protein concentrations of clarified lysates were determined using a BCA protein assay kit (Pierce). Lysates were diluted to 1 mg ml−1 and 100 μl of each lysate was incubated with 50 μl of NeutrAvidin agarose beads (Pierce) for 1 h at 4 °C with gentle rotation. Beads were washed four times with NP-40 buffer and biotinylated proteins were eluted by heating in 1x NuPAGE LDS sample buffer (Invitrogen) with 100 mM DTT at 70 °C for 20 min. Input and biotinylated fractions were analysed by immunoblotting as described above. Band intensities were quantified using ImageJ.
Fluorogenic peptide cleavage assay
The quenched fluorescent peptide Mca-KAQPVEA-Lys(Dnp) was custom synthesized by GenScript and initially dissolved as a stock solution in DMSO. Reactions (100-μl final volume) consisting of the peptide diluted to the indicated concentrations in 1 × PBS were prepared in triplicate in a black/clear-bottom 96-well plate (Thermo Scientific, 165305) on ice. BFT was added to each well last at a final concentration of 457 nM and the assay plate was immediately transferred to a CLARIOstar Plus microplate reader (BMG Labtech) pre-heated to 37 °C. Fluorescence measurements (excitation = 330 nm, emission = 390 nm) were acquired every 4 min for 6 h at 37 °C. Fluorescence units were converted to nM peptide hydrolysed using a standard curve of fluorescence units versus initial peptide concentration at 340 min post BFT addition, when all reactions had gone to completion. Km and Vmax values were determined by nonlinear curve fitting in GraphPad Prism using the model ‘Michaelis Menten’. The kcat value was calculated by dividing the obtained Vmax by the BFT concentration in the assay, 457 nM.
AlphaFold modelling
To model the hypothetical ternary complex between claudin-4, BFT and E-cadherin, we used AF3 implemented in the AlphaFold server (https://alphafoldserver.com/)43. The following input sequences were used: human claudin-4 (UniProt ID O14493; amino acids M1–P186, excluding the unstructured cytoplasmic CTD), BFT-2 (UniProt ID O05091; amino acids A212–D397, excluding the N-terminal prodomain), human E-cadherin (UniProt ID P12830; amino acids V480–A735, excluding the unstructured cytoplasmic CTD and the outer three extracellular domains EC1–3) and one zinc ion. Predictions were run without using templates. Ten predictions with different random seed values were generated by the AlphaFold server and the prediction with the best interface-predicted template modelling (ipTM) score was chosen (random seed = 1035423868, ipTM = 0.62). Figures were generated in UCSF ChimeraX. The membrane-embedded claudin-4 crystal structure (PDB: 7KP4) was downloaded from the MemProtMD database63. Alignment of the AlphaFold model to the membrane-embedded claudin-4 structure was performed using the ‘matchmaker’ command in ChimeraX.
Plotting and statistical analysis
GraphPad Prism 10 was used to plot BFT dose–response curves, T84 monolayer studies, fluorogenic peptide cleavage data and all displayed bar graphs. Scatterplots in Extended Data Fig. 2d were generated using ggplot2 version 3.4.4 in R version 4.2.2. Sequence alignments were generated using Clustal Omega or EMBOSS Needle64 and the alignment shown in Extended Data Fig. 3c was created using ggmsa version 1.0.3. Data are shown as mean ± s.d unless otherwise indicated in the figure legend. The number of independent biological replicates for each experiment is reported in the legend.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.