The UM-HET3 sibship
UM-HET3 mice are progeny of a cross between two types of F1 hybrids—female F1 mice from matings of BALB/cByJ dams to C57BL/6J sires and male F1 mice from matings of C3H/HeJ dams to DBA/2J sires. These four inbred progenitors are abbreviated CBy, B6, C3H and D2 when referring to mice and strains, and abbreviated C, B, H and D when referring to genotypes and haplotypes. These four fully inbred strains were selected to maximize phenotypic diversity. Young virgin F1 males and females bred at the Jackson Laboratory (JL) JAX facility were transferred to ITP ageing colonies at JL in Bar Harbor Maine, the University of Michigan in Ann Arbor Michigan (UM) and the University of Texas Health Science Center in San Antonio (UT). Breeding cages were set up in spring. First litters were not used. All subsequent litters were used at UT, litters of 6 or more pups were used at JL, and those with 7 or more pups were used at UM. Weanlings were entered into the study over the next 7–8 months.
This study is based on tail samples acquired initially at the three ITP sites in accordance with standards of the Association for the Assessment and Accreditation of Laboratory Animal Care and recommendations of the National Institutes of Health Guide for the Care and Use of Laboratory Animals, including annual reviews and approvals of all protocols. Links to data for ITP papers and data are available at the Mouse Phenome Database75.
All mice in the first four cohort years were born between April 2004 and January 2008. In these cohort years we have numerically well-balanced DNA samples from all sites. Almost no mice were generated in 2008 owing to a funding gap. All mice in the final cohort years used in this study were born between July 2009 and March 2013. The 2009 cohort includes mice from all sites, but 2010 and 2011 include mice only from UM and UT, and cohort years 2012 and 2013 include mice only from UM. The last UM-HET3 mouse in this study died in 21 December 2015. In all years, only a small percentage of mice (<10%) were born between January and April. Mice were weighed at 42 ± 2 days, and at 183 days (6 months), 365 days (12 months), 548 days (18 months) and at 730 days (24 months) with a timing error of ~7 days.
The UM-HET3 sibship segregates for ~10.6 million sequence variants (Supplementary Table 1). All UM-HET3 mice inherit C and B haplotypes from their F1 mothers and H and D haplotypes from their fathers. As a result, the entire sibship segregates for four genotypes, the four two-way combinations of maternal and paternal haplotypes, on all autosomes—CH, CD, BH and BD. Females inherit one entire non-recombinant H-type chromosome X from their paternal grandmothers and a potentially recombined chromosome X from their mothers (recombinations between C and B haplotypes only; Fig. 3g). As a result, females have either CH or BH chromosome X genotypes. Hemizygous males inherit a potentially recombined C or B chromosome X from their mothers. All mitochondria and their genomes in all animals are derived from maternal grandmothers (C) and male chromosome Y is derived from paternal grandfathers (D).
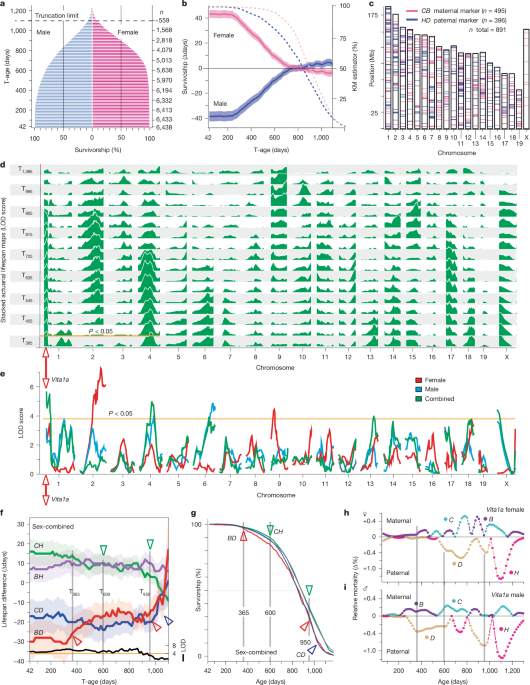
We genotyped 6,872 UM-HET3 mice for which we had full lifespan estimates (n = 3,252 females, n = 3,620 males). Of these genetic siblings 6,438 passed all genotype quality control steps at 891 markers (n = 3401 females, n = 3037 males; see Fig. 1c and ‘Genotype quality control’). To ensure balanced numbers by sex later in life, every ITP cohort initially consists of 51 male and 44 female weanlings per year, per site and treatment category, but with twofold more common untreated controls at each site. This was done to enable overly aggressive males and any wounded mice to be removed while balancing male and female numbers later in life—almost always before 550 days. The earliest minimum inclusion age in this study is T42, the pubescent age at which tails were docked for tissue acquisition, an age equivalent to about 12 years in humans. For numerical convenience we set the first T-age at T35. To compensate for twofold higher male mortality in the first 2 years of life (33% versus 16%), we included 12% more males than females in the T42 survivorship. The earliest death was at 46 days. This left us with a small surplus of 227 males at T365, but by T560, numerical balance was restored and there were 2,930 females and 2,929 males. This age corresponds to roughly 56 years of age in humans. All individual-level data, metadata, survivorships and mortalities per 15-day interval are provided in Supplementary Table 2.
We genotyped two major categories of mice—those not treated with any dietary intervention and mice treated with a dietary supplement that did not modify lifespan significantly on a per-drug basis using standard statistical criteria76,77,78. These latter mice have been referred to as ‘no drug effect’ (NDE) cases. However, when data are combined across the entire class of these individually ‘ineffective’ agents from 2004 to 2013, there is a highly significant combined effect that is positive on lifespan (Fig. 2i,j).
Husbandry
Mice were weaned into same-sex cages—three males or four females per cage—at 20 ± 1 days (ref. 22). They lived together from weaning to death without any replacements within cages. As a result, most mice lived alone after ~1,000 days, a factor to consider with respect to mortality risks, but not yet integrated into our analysis. From 2004 until 2013, all sites used NIH-31 standard diets. For breeding cages, UM used Purina 5008, UT used Teklad 7912, and JL used Purina 5K52. For mice up to ~122 days, UM used Purina 5001, UT used Teklad 7912, and JL used Purina 5LG6. After 2004, a single control diet—LabDiet 5LG6—was used by all sites. Mice were monitored daily for signs of ill health and aggression and euthanized if moribund. Tails were obtained at 42 ± 2 days for DNA extraction. Body mass at this age was acquired for 2,459 mice, and at half-year intervals for 4,688 mice at 183 days (6 months), and down to 2,208 mice at 730 days (24 months). Our analysis includes four experimental variables—sex, site, dietary drug treatment and cohort year (Supplementary Table 3). The class of nominally ineffective treatments included in this study are listed on Mouse Phenome Database ITP Portal under the acronyms 4OHPBN, CAPEhi, CAPElo, Cur, Enal, FOhi, FOlo, GTE, HBX, I767d, MB, MCTO, MET, NFP, OAA, Res07, Reshi3, Reslo3, Simhi, Simlo and UA.
DNA extraction, sequencing and sequence alignment
Tail tissue was processed using the MagMAX magnetic-bead extraction system and genomic DNA was quantified using a Qubit 4 fluorometer and a high-sensitivity DNA assay kit (Thermo Fisher Scientific). Locus-specific PCR primers were designed using BatchPrimer379 to generate 180-bp amplicons. They were mixed using an optimized Hi-Plex approach at Floodlight Genomics. Sample-specific 12-bp barcodes were added and amplicons were pooled80. Primer sequences for markers with reference SNP rs identifiers are provided in Supplementary Table 4. The amplicons were sequenced to an average of ×1,000 per targeted DNA variant on an Illumina NovaSEQ (2× 150 bp reads). Data were demultiplexed and FASTQ files used for genotyping. FASTQ files were aligned to M. musculus GRCm38.p6/mm10 reference using Bowtie 2 (v2.3.4.1)81. BAM files were sorted and indexed using samtools (v1.6)82, and read group information was added using picard tools (v2.14.1).
Variant calling
We called short sequence variants using bcftools82 in three steps: (1) we created text pileup output for all BAM files using mpileup with mapping quality of 30 or greater; (2) we called using default settings; (3) we removed variants with a read depth of less than 100 across all samples or a QUAL score of less than 100. Sequencing depth per case was used as an additional criterion for filtering reads. Sequence for C57BL/6J, C3H/HeJ and DBA/2J was downloaded from the Wellcome Sanger Mouse Genome Project83 (https://www.sanger.ac.uk/data/mouse-genomes-project) and that for BALB/cByJ was generated by Beierle and colleagues84. From these files we extracted variants that differ among the four founders (Supplementary Table 1). Variants were called in three steps as above, the differences being that the minimum depth was reduced to ×10 and QUAL score to ≥30. Variants that were called reliably in all founders were advanced for genotype phasing and mapping. VCF files with variant data on UM-HET3 and their founders are provided in Supplementary Table 5.
Genotyping by Sequenom MassARRAY
Tails were placed in deep-well plates and sent in three batches to Neogen Corporation. Individuals were genotyped by SNP genotyping by Sequenom MassARRAY MALDI-TOF mass spectrometry85 at a maximum of 270 markers. Fifty markers were removed after quality control, based on deviations from expected allele frequency (0.35 < frequency < 0.65). We chose pairs of linked markers to define maternal and paternal chromosome genotypes.
Phasing SNP genotypes
SNP genotypes were phased using R v4.0.2 (script at https://github.com/DannyArends/UM-HET3). We generated fully phased haplotypes for all sets of maternal and paternal autosomes and chromosome X. Markers fall into those that unambiguously define maternal haplotypes (n = 486, C versus B alleles and pink lines in Fig. 1c) and paternal haplotypes (n = 396, H versus D alleles and blue lines in Fig. 1c). We generated diploid genotype probabilities at all positions from phased chromosomes using the calc.genoprob() function in R/qtl. Markers were reviewed and those that had a call rate <30% were excluded. When we plot genotype or haplotype effects (Extended Data Figs. 2–5), we call genotypes and haplotypes when imputation certainty is greater than 80%. We used R/qtl to estimate number of recombinations per mouse. The mean ± s.d. value was 25.1 ± 8.6. Just over 370 animals had recombination numbers more than 2× s.d. above this value. We excluded 59 mice that had more than 80 recombinations, a value likely to be caused by sample cross-contamination. The final genotype files are available at GeneNetwork.org86,87,88 by setting Species = Mouse, Group = Aging Mouse Lifespan Studies (NIA UM-HET3), and Type = DNA Markers and SNPs and pressing the Info button. Supplementary Table 6 lists all genotypes.
Genotype quality control
Markers and individuals with greater than 90% missing data were removed from further analysis (n = 333). We converted data from genotypes into parental haplotypes (see ‘Phasing SNP genotypes’). Markers were checked for Mendelian inheritance errors and removed when missing data after conversion exceeded 90%. We integrated MonsterPlex and Sequenom marker sets based on genomic positions. We inspected the marker map for linkage between neighbouring markers and removed a single marker (chr. 4_58784823) which did not show linkage to its neighbours. We also removed 20 individuals that died of unnatural causes and 22 that we inadvertently genotyped that had received an effective drug supplement. Finally, we constructed an R/qtl cross object for imputation using the remaining mice, imputed their haplotypes based on the full population (n = 6,438). Supplementary Table 7 shows the R/qtl cross object used for most analyses.
Inversions in the UM-HET3
There is a large inversion on chromosome 6 of the C3H/HeJ progenitor strain89 between 51 Mb and 94 Mb which will affect regional map precision. The recombination fraction in this interval is 0.00025 versus an expectation of 0.21. At least 42 Mb of the paternal chromosome is locked in linkage. We detected a total of 13 potential inversions ranging in size of 3.5 to 54 Mb. Six suppress recombination on the maternal chromosome and seven suppress recombination on the paternal chromosome (Supplementary Table 8). There is overlap of maternal and paternal inversions on chromosomes 4, 7 and 11.
Lifespan estimates depend on age of entry
Based on work by König and colleagues38,90, we know that in wild commensal populations of mice, 35–40% of pups do not survive beyond weaning. At birth, mean lifespan can be as low as 196 days. By 13 days, the mean expectancy has improved to 250 days for males and 500 days for females. We make these points to emphasize that lifespan estimates are made with reference to an operational starting point—a minimum ascertainment base age for a population. Researchers often incorrectly imply that mean lifespan is estimated from the date of birth, but this neglects earlier and often unknown numbers of deaths—referred to as left truncation. In our case we can only measure lifespan for UM-HET3 progeny that lived to be pubescent juveniles. When we say that the mean lifespan of the 6,438 sibs in this study is 844 ± 210 days (mean ± s.d., median of 870 days), we mean that this is the mean age at death conditioned on a mouse having lived to at least 42 ± 2 days—the age at which tail tips were taken for DNA extraction.
In a similar way we can determine the mean lifespan or life expectancy of the survivorship subsets of UM-HET3 mice that lived to be at least 185, 365, 545, 725 or 1,100 days (roughly 0.5, 1, 1.5, 2 and 3 years, respectively). The mean conditional lifespans of those five survivorships or age cohorts are 847 ± 206, 861 ± 188, 890 ± 160, 941 ± 126 and 1,162 ± 54 days (mean ± s.d.), respectively. The oldest mouse was a female that died at 1,456 days. We refer to these different ages at which lifespan can be computed as either survivorships or cohorts.
The actuarial procedure consists of sequentially studying progressive older and therefore smaller survivorships from the radix population of 6,438 mice in Fig. 1a. The minimum T-age is 42 days. Between T42 and T50 days a single male died. We truncated survivorships upward in 15-day steps from T35 to T1,100, a convenient step size for plots that can also be rationalized as roughly the range of survivorship T-ages (1,065 days) divided by the square root of the radix (n of 6,438). The first anchor point age was defined as 35 days in order to align plots at 365 days (1 year), 725 days (~2 years) and 1,100 days (~3 years) using 72 steps. We explored both right and bilateral truncation but do not cover the analyses here.
The actuarial mapping strategy is affected both by lower sample size and by right skew at higher truncations. To ensure that our results are not affected by this, we used three approaches. First, we use a non-parametric quantile regression in combination with standard parametric mapping to make sure associations are detected using both methods91. Second, we limited ourselves to a maximum T-age of T1,100. Third, we used a time-dependent hazard function to validate Vita loci.
We tested a complementary approach for mapping under the assumption that loci contribute to variation in the time-dependent hazard function as in our previous work17 but with refinements. We carried out a test at each marker of the null hypothesis that the (time-dependent) hazard function of death does not depend on genotypes. To do this, we used a model where the hazard function is allowed to depend smoothly on age with haplotype and baseline covariates, using natural splines with three degrees of freedom. For computational tractability we created risk sets with 50-day windows92.
Implementation of actuarial mapping
We mapped in 15-day survivorship steps from T42 to T1,100. Mapping was performed using both conventional four-way mapping of genotypes at 891 markers (Fig. 1c) or by restricting analysis to each of the parental haplotypes—C versus B on the maternal chromosomes (n = 495 markers), and H versus D on the paternal chromosomes (n = 396 markers). Note that chromosome X only segregates for maternal C and B haplotypes in both sexes (Fig. 3g). In general, mapping all four genotypes provides better power. This method defined all loci in Tables 1 and 2 and full LOD scores across all survivorships are illustrated in Extended Data Figs. 1 and 2. The null model (model(H0)) and alternative model (model(Halt)) were fitted at each marker using the following linear models:
$${\rm{model}}({{\rm{H}}}_{0}):{\rm{lifespan}}={\rm{sex}}+{\rm{site}}+{\rm{cohort}}+{\rm{treatment}}+{\rm{error}}$$
$${\rm{model}}({{\rm{H}}}_{{\rm{alt}}}):{\rm{lifespan}}={\rm{sex}}+{\rm{site}}+{\rm{cohort}}+{\rm{treatment}}+{\rm{gtp}}+{\rm{error}}$$
Sex has two levels (M, F), site has three levels (JL, UT and UM), cohort year has nine levels (no animals were generated in 2008), treatment has two levels—untreated controls and cases that received a drug that had no significant effect on lifespan. The gtp (genotype probability) term is an n row by four-column matrix, where n is the number of individuals used in the mapping, and columns are genotype probabilities for markers computed by R/qtl (v1.48-1)93,94. Sex was dropped as a co-factor from the regression model when mapping was stratified by sex. LOD scores were computed by comparing the fit of the null model with the fit of the alternative:
$$\mathrm{LOD}=(n/2)\times {\log }_{10}(\mathrm{sum}(\mathrm{residuals}{({{\rm{H}}}_{0})}^{2})/\mathrm{sum}(\mathrm{residuals}{({{\rm{H}}}_{\mathrm{alt}})}^{2}))$$
Mapping on chromosome X requires special care due to the four-way cross structure and greater difficulty defining highly reliable markers for haplotype, particularly close to both telomeres.
Actuarial effect size plots, their errors and interpretation
We use the four-way mapping algorithm implemented in R/qtl94,95 while controlling for sex, site, cohort year and drug treatment. We compute maps for survivorships starting from T42 up to T1,100 (Fig. 1a,d–f). We introduce a new type of actuarial plots in Fig. 1b,d,e. The x axis defines the minimum inclusion age (T-age) of each survivorship. The expected increase in error terms in older survivorships with lower sample sizes are mitigated by the reduced range over which animals die within progressively older survivorships—from 1,410 days at T42 to 356 days at T1,100. Errors of lifespans of survivorships are relatively stable—from 844 ± 210 (mean ± s.d.) in the T42 base population (s.d./range of 15%) to 1,162 ± 54.4 (mean ± s.d.) in the last T1,100 survivorship (s.d./range of 15%). When interpreting effect size plots note that increases and decreases in mortality rates among subgroups are relative to the mean lifespan of that T-age.
The T-age of a survivorship always refers to an entire age range. Actuarial truncation in a forward direction means that the oldest-old mice are embedded in every survivorship. Reverse truncation flips the polarity by truncating from oldest to youngest and defines exitships. The 1,099T exitship consists of mice that died before 1,100 days and complements the T1,100 survivorship. The combination of forward, reverse and even two-sided truncations is useful to dissect potential causes of mortality in age-restricted subpopulations. In this paper we restrict attention to forward truncations.
Significance testing for actuarial analyses
We evaluated the significance of mapping results across two axes: (1) the spatial location of markers and their independence across the genome; and (2) the temporal axis and the number of effectively independent actuarial tests using left truncation that progressively excludes mice that died before reaching the T-ages. We mapped using 891 markers but given the significant linkage between markers we effectively tested about 442 independent genome locations. This number of independent tests was estimated using the simpleM method with a block length ranging from 10 to 500 markers and a 10-marker step size96. The threshold at a genome-wide type 1 Bonferroni error rate of 10% is 3.65 LOD (–log10(0.1/442) at 5% it is 3.95, and at 1% it is 4.65. These are conservative thresholds. For maternal and paternal maps, the density of recombination is reduced by half, and corresponding thresholds are 3.44, 3.65 and 4.34.
Locus confidence intervals
We used the standard 1.5 LOD drop to estimate the confidence interval of linkage97. We also estimated the temporal duration of action of loci, defined as the range of T-ages over which a locus has strong effects. The start and stop T-ages of these temporal confidence intervals are those that immediately precede or succeed survivorships that are genome-wide significant. For example, if linkage is above LOD of 3.95 from T365 to T725 survivorships, then the T-age confidence interval of the locus is from T350 to T740. This criterion is not conservative when only a few survivorships have significance. In this latter case, we defined the T-age range that brackets the peak T-age by at least a 1.5 LOD drop.
We evaluated the impact on significance testing of mapping multiple nested survivorships using a Cauchy combination test98 at each marker across all survivorships (Table 1). Cauchy P values were adjusted for multiple testing using a Benjamini and Hochberg correction to account for multiple testing across markers, and the Cauchy –logP in Table 1 of ≥1.50 is significant at a P of less than 5%.
Locus names and ambiguities
Locus names are prefixed with Vita, Soma and Mass identifiers followed by chromosome and a letter suffix (for example, Vita1a, Soma2b and Mass11c). When the 95% confidence intervals of two adjacent QTLs overlap, and the genotype and haplotype effects appear to be the same, loci are generally assigned the same Vita identifier. For such ‘merged loci’, the 95% confidence interval is reduced to the overlapping interval. Determining whether a locus is detected in different subsets of the sibship is somewhat subjective, examples being Vita3a and Vita14b that could have been split into two loci by sex and T-age. We provide data for linkage scores in all three categories (sexes combined, females and males) for Vita loci (Supplementary Table 9), Soma loci (Supplementary Table 10) and Mass loci in (Supplementary Table 11).
Testing for sex interactions
There are significant differences between male and female stratified survivorship maps. To investigate this formally and to detect significant sex-marker interactions modulating age-specific lifespan we tested sex interactions of Vita loci at the T-age with the peak LOD (Fig. 3a). We computed the LOD score difference of two models against each other:
$${\rm{model}}({{\rm{H}}}_{0}):{\rm{longevity}}={\rm{sex}}+{\rm{site}}+{\rm{cohort}}\,{\rm{year}}+{\rm{treatment}}+{\rm{gtp}}+{\rm{error}}$$
$${\rm{model}}({{\rm{H}}}_{{\rm{alt}}}):{\rm{longevity}}={\rm{sex}}+{\rm{site}}+{\rm{cohort}}\,{\rm{year}}+{\rm{treatment}}+{\rm{gtp}}+{\rm{gtp}}:{\rm{sex}}+{\rm{error}}$$
Since we are only interested in significant sex interactions at previously detected Vita loci (excluding only VitaXa) we used a Bonferroni correction based on 28 markers. A LOD score above 2.75 is significant (–log10(0.05/28)).
Testing pairwise epistatic interactions
A two-dimensional analysis of marker-marker interactions affecting life expectancies was performed by testing the following models against each other in four survivorships—T42, T365, T740 and T905.
$${\rm{model}}({{\rm{H}}}_{0}):{\rm{lifespan}}={\rm{sex}}+{\rm{site}}+{\rm{cohort}}\,{\rm{year}}+{\rm{treatment}}+{{\rm{gtp}}}_{{\rm{m}}1}+{{\rm{gtp}}}_{{\rm{m}}2}+{\rm{error}}$$
$${\rm{model}}({{\rm{H}}}_{{\rm{alt}}}):{\rm{lifespan}}={\rm{sex}}+{\rm{site}}+{\rm{cohort}}\,{\rm{year}}+{\rm{treatment}}+{{\rm{gtp}}}_{{\rm{m}}1}+{{\rm{gtp}}}_{{\rm{m}}2}+{{\rm{gtp}}}_{{\rm{m}}1}:{{\rm{gtp}}}_{{\rm{m}}2}+{\rm{error}}$$
We computed the difference in fit between H0 and Halt as explained in ‘Implementation of actuarial mapping’. To limit numbers of tests, we initially tested interactions between the 29 Vita loci (Table 2), excluding all pairs of loci on the same chromosome (Fig. 5a). This resulted in the matrix of 387 non-syntenic tests of pairs of loci per sex. We used Bonferroni corrections for most work. With 387 Vita–Vita tests, a LOD of approximately 4.0 is appropriate, but for more comprehensive tests of all loci in all 4 survivorships (11,768 tests) the corresponding P < 0.05 threshold is 5.37, a value that limits the yield of interactions to only 50 (Supplementary Table 12). By contrast, the Benjamini and Hochberg test suggests that 966 interactions have FDR of less than 0.005 (Supplementary Table 12.2). We opted for a compromise and analysed a total 289 unique epistatic interactions that are visualized in part in Fig. 5g,h for the T42 survivorship, and much more comprehensively Extended Data Fig. 9 for all 4 survivorships. All interactions have LODs greater than 3.8. Of a total of 11,768 tests fewer than 4% have LODs ≥ 3.8 (Supplementary Table 12).
Visualizing genetic modulation of mortality rates
We plotted age-dependent mortality and the HRs of haplotypes to give more insight into the specific ages over which genetic differences operate with more or less force. We computed mortality rates of maternal and paternal haplotype pairs (C versus B, H versus D) for males and females separately. We used LOESS smoothers with span parameters of 0.2 (Extended Data Fig. 4), and adaptive LOESS with a span of 0.3 (refs. 99,100,101) with shorter spans over ranges with high numbers of deaths (800–950 days) and longer spans over ranges with few deaths (Fig. 2f). log2 HRs greater than 0.6 represent relatively strong genetic effects (HR > 1.5 or HR < 0.67 in Fig. 2f).
Body mass versus lifespan analysis—the Soma loci
Body masses were measured at 42 days (right before tails were biopsied for genotyping); and at 183, 365, 548 and 731 days (6, 12, 18 and 24 months). We used these data to map loci for body masses for both sexes so that we could align and compare body mass loci with Vita and Soma loci (for example, Fig. 5a–c). Sample sizes for mapping Mass loci range from a low of 2,459 at 42 days to a high of 4,688 at 183 days (Supplementary Table 11.1). Models used are like those used for mapping Vita loci, but the variable is body mass at each of five ages.
Actuarial analysis of Soma loci
To investigate the effects of body mass on lifespan, we adjusted body mass and lifespan using the model:
$$\begin{array}{l}{{\rm{Lifespan}}}_{({\rm{adjusted}})}={\rm{mean}}({\rm{Lifespan}})+{\rm{residuals}}({\rm{Lifespan}} \sim {\rm{Sex}}\\ \,+{\rm{Site}}+{\rm{Cohort}}+{\rm{Treatment}})\end{array}$$
$$\begin{array}{l}{{\rm{BW}}}_{({\rm{adjusted}})}={\rm{mean}}({\rm{BW}})+{\rm{residuals}}({\rm{BW}} \sim {\rm{Sex}}\\ \,+{\rm{Site}}+{\rm{Cohort}}+{\rm{Treatment}})\end{array}$$
Where BW is body mass. This model includes mice for which we have lifespan and body mass data at one or more of the five weight ages. We computed the sex-stratified Spearman rank order correlations between body mass at 42, 183, 365, 548 and 730 days with lifespan in 15-day increments (Fig. 4d–p). We compute the –logP of the difference in correlation between males and females using the cor.test function in R. The average correlation in the combined population between adjusted body mass at 183 days (~6 months) and adjusted lifespan is rho = –0.206. When computed for males, correlation is stronger (rho = –0.284), while it is weaker for females (rho = –0.110).
CTL mapping
We used correlated trait locus (CTL) mapping37 to determine if a distinct set of loci modulate correlations between body mass and subsequent lifespan (Tables 3–4). Before using this procedure, we adjusted body mass and lifespan using sex, site, cohort year, and drug treatment as covariates. At each marker we stratified survivorships based on genotypes. Ideally, the sample size for each subgroup would be above 400, but in some cases, N was as low as 200 due to genotype uncertainty and to the lower sample sizes at T42 (N = 2549) and T730 (N = 2208). We computed rho correlations for each genotype with n > 100 and the z-scores associated with differences:
$$z=0.5\times \log ((1.0+\rho )/(1.0-\rho ))$$
Here cor is a four-value vector containing the observed correlations for CH, BH, CD and BD genotypes. We compute the sum of squares (sumOfSq) by multiplying the observed sample sizes (ss) of the CH, BH, CD and BD genotypes to the squared z-scores:
$${\rm{sumOfSq}}={\rm{sum}}({{\rm{ss}}}^{* }{{\rm{z}}}^{2})$$
We compute the squares of sums (sqOfSum):
$${\rm{sqOf}}{\rm{Sum}}={\rm{sum}}({{\rm{ss}}}^{* }{{\rm{z}}}^{2})$$
Using these values, and the sum of the sample sizes on which each correlation is based, we compute the critical value (Cv) which follows a chi-square distribution under the null hypothesis that all correlations (z-scores) are from the same distribution:
$${\rm{Cv}}={\rm{sumOfSq}}{\rm{\mbox{–}}}({\rm{sqOfSum}}/{\rm{sum}}({\rm{ss}}))$$
The Cv is converted to a P value using the chi-squared distribution using the pchisq() function in R, with P[X > x] and three degrees of freedom (number of genotypes – 1).
We adjust the significance threshold for multiple tests using the p.adjust function in R at a 5% FDR (refs. 39,102). The 5% FDR threshold is approximately 2.75 –logP. While still stringent, this threshold is less harsh than that applied to Vita loci that use a highly conservative Bonferroni correction. Our rationale for this difference is our use of conservative non-parametric rho for comparing correlations of body mass to lifespan. Differences in correlation correspond to days gained or lost relative to an average individual that can be converted to effect sizes measured in days gained or lost per gram of body mass. We computed the linear regression coefficient for body mass on lifespan for the four genotypes as:
$${\mathrm{Lifespan}}_{(\mathrm{adjusted})}=\mathrm{mean}(\mathrm{Lifespan})+\beta \times {\mathrm{bodyweight}}_{(\mathrm{adjusted})}$$
where mean(Lifespan) is the intercept of the total population (all genotypes combined), while the β coefficient is the estimated effect size of body mass on lifespan based on the subpopulation defined by each genotype (Table 4). This leads to effect sizes relative to an averaged mouse. While there are caveats with respect to interpreting these effects because they are computed relative to this mean, these values give readers a sense of the impact of Soma loci on life expectancies.
Broad-sense haplotype-based heritability
We compute broad-sense genotype-based heritability by fitting a full model including the haplotype probabilities of the 29 top markers of all Vita loci:
$$\begin{array}{l}{{\rm{lifespan}}}_{({\rm{T}}-{\rm{age}})}={\rm{sex}}+{\rm{site}}+{\rm{cohort}}+{\rm{treatment}}+{{\rm{gtp}}}_{{Vita}1a}\\ \,+{{\rm{gtp}}}_{{Vita}1b}+\ldots +{{\rm{gtp}}}_{{VitaXa}}+{\rm{error}}\end{array}$$
Broad-sense haplotype-based heritability is estimated by fitting this model to all survivorships. When stratifying by sex, sex is dropped from the model as well as from the H2e component described in the computation of environmental variance component below. Computing heritability is done by taking the following approach, adapted from Falconer103 and Lynch and Walsh104 using five steps:
Step 1: The vector of partial total variances (PTV):
PTV = (σ2 – σ2residuals)/r
Step 2: An adjustment factor adj—the sum of the mean sum of squares for the fixed effect including the residual variance:
adj = sum(σ2)/r
Step 3: The contribution of each parameter in PTV to the model:
C2p = PTV/adj
Step 4: The total broad-sense genotype-based heritability:
H2h = C2Vita1a + C2Vita1b + … + C2VitaXa
Step 5: The environmental variance:
H2e = C2sex + C2site + C2cohort + C2treatment
where σ2 is the vector containing all the means of sums of squares (including the residual) computed by the ANOVA model. σ2residuals is the residual mean sum of squares. r is the average number of replicates for each genotype. H2h can be interpreted as the broad-sense genotype-based heritability as a sum of all Vita loci. H2e is the environmental variance estimate of the known environmental covariates. Unexplained variance H2u can be computed as:
$${{H}^{2}}_{{\rm{u}}}=1-({{H}^{2}}_{{\rm{h}}}+{{H}^{2}}_{{\rm{e}}})$$
To obtain upper and lower bounds on H2h and H2e we generated 50 bootstrap resamples and fit the ANOVA model to each using a random subset of 90% of the survivorship. We computed median and standard deviations across all bootstraps to estimate errors of H2h and H2e as a function of survivorship T-age.
Candidate genes analysis
All annotated features located within the 95% confidence interval were downloaded using BioMart105. We applied several criteria to prioritize positional candidate genes: (1) first, we considered only the protein-coding genes that reside in the 95% confidence intervals; (2) we then selected genes with annotated non-synonymous SNPs segregating in the population, and these were further ranked by the potential deleterious effect based on the ENSEMBL variant effect predictor106; and (3) another priority score for the candidate genes was based on whether they were listed as ageing and longevity genes in GenAge107. We also compared our positional candidate genes against known human, C. elegans, D. melanogaster and Saccharomyces cerevisiae genes reportedly associated with age in GenAge. To accomplish this, we use biomaRt to convert mouse gene symbols to the corresponding species-specific orthologous gene symbols. Supplementary Tables 13 and 14 provide access to all gene models and variant types segregating in Vita and Soma loci.
Gene set over-representation analysis
We analysed gene set enrichment in Vita and Soma loci using R. Gene annotations were retrieved from Ensembl via biomaRt, retaining only mouse protein-coding genes, but excluding olfactory receptor genes (Olf*, n = 1,427), vomeronasal receptor genes (Vmn*, n = 595), predicted genes (Gm*, n = 1,246) and RIKEN cDNA sequences (n = 490). For each locus we tested set over-representation against our filtered whole-genome background (n = 18,830) using clusterProfiler for Gene Ontology (Biological Process, Molecular Function, Cellular Component), KEGG pathways and Reactome pathways. An adjusted P cut-off of 0.2 was applied to capture marginally significant enrichments for subsequent analysis. Results are summarized in Supplementary Table 16.
C. elegans candidate gene screening of Vita9b
Out of 262 protein-coding genes in Vita9b, we highlighted 98 with missense variants and a subset with C. elegans orthologues using Ortholist2108 (Supplementary Table 16). From these, we selected a subset of 15 genes (Fig. 6g) represented in the Ahringer C. elegans RNAi library109 for testing: (1) acds-10 (orthologue of mouse Acad11); (2) chhy-1 (Hyal1, Hyal2 and Hyal3); (3) oct-1 (Slc22a13); (4) col-135 (Col6a6); (5) frm-8 (Frmpd1); (6) let-413 (Lrrc2); (7) F30A10.3 (Ip6k2); (8) lin-41 (Trim71); (9) lis-1 (Poc1a); (10) stt-3 (Stt3b); (11) C41D11.3 (Csrnp1); (12) znf-782 (Zfp445); (13) pes-4 (Pcbp4); (14) pho-6 (Acp3); and (15) dpf-5 (Apeh). We also screened using the L4440 empty vector control and daf-2 (Igf1r) as positive control. The sequence-verified single RNAi clones were inoculated in 10 ml lysogeny broth (LB) with ampicillin (100 μg ml−1) and tetracycline (50 μg ml−1) overnight at 37 °C. The next day, 5 ml of LB with 0.5 mM isopropyl β-D-1-thiogalactopyranoside was added and inoculants were incubated for 30 min at 37 °C with shaking for pre-induction. The bacterial inoculants were centrifuged (3,700 rpm for 15 min) and the pellet was resuspended in 10 ml liquid NGM with 50 µg ml−1 IPTG, 100 µg ml−1 ampicillin and 13.32 mg ml−1 nystatin.
Gravid adult stage TJ1060 [spe-9(hc88);rrf-3(b26)] C. elegans were synchronized by bleaching. Resulting L1 stage worms were suspended in concentrated heat-inactivated OP50 bacteria in liquid NGM and transferred into U-bottom 96-well plates in a final volume of 100 µl. The worms were grown on an orbital shaker (Heidolph Titramax 1000) at 600 rpm until they reached the L4 stage (2.0–2.5 days) at 25 °C to prevent progeny production. When worms reached L4 stage, 20 µl of bacterial inoculant of each RNAi clone were added into single wells of the U-bottom 96-well plates containing worms. At least 30 worms per well and 8 wells per clone were used for two independent screens. The MicroTracker (InVivo Biosystems) system was used to measure motility in liquid in 96-well plates by infrared beam breaks. Plates were measured daily for a period of at least 90 min. Significance of differences in activity were estimated using the AUC (Fig. 6f) for either the full lifespan (1–30 days relative lifelong motility, Fig. 6g) or only after 14 days (Fig. 6h) using a two-tailed t-test of control-normalized AUC values assuming unequal variance and using a Bonferroni correction for 15 tests.
Mendelian randomization analysis
We used Mendelian randomization methods to rank the translational relevance of genes within Vita loci against their human orthologues. We extracted lists of protein-coding genes within Vita1a and Vita9b with significant human cis-expression quantitative trait loci (cis-eQTLs) in blood from eQTLGen (31,684 individuals)110 as exposures (Supplementary Table 15). As outcome variables, we used summary statistics on parental longevity111,112 from the IEU OpenGWAS project113 and on human longevity114 from the GWAS Catalog115. We tested for potential causal effects of variation in expression on: (1) extreme longevity of at least one parent (>95 years of age, IEU OpenGWAS.io trait ebi-a-GCST003395)111; (2) both parents in the top 10% of longevity (IEU OpenGWAS.io trait ebi-a-GCST006698)112; (3) mother’s age at death (IEU OpenGWAS.io trait ebi-a-GCST006699)112; (4) father’s age at death IEU OpenGWAS.io trait (ebi-a-GCST006700)112; (5) mean parental age at death (IEU OpenGWAS.io trait ebi-a-GCST006702)112; (6) individual survival into the top 10% (GWAS Catalog: GCST008598)114; and (7) individual survival into the top 1% (GWAS Catalog: GCST008599)114. In many instances, multiple cis-eQTLs were detected in the GWAS summary statistics, allowing them to be used as instrumental variables. Because linkage disequilibrium pruning would leave only a small number of independent variants, we applied an established principal component analysis-based approach that aggregates correlated instrumental variables into linkage disequilibrium-based principal components (principal component instrumental variables)116 that were evaluated using the 1000 Genomes Project panel117 with PLINK118 v1.90b6.21. We retained top principal components explaining more than 99% of genetic variance and used the inverse-variance weighted method to test for potential causal effects. When only a single instrumental variable was available for both the exposure and the outcome, or when principal component analysis yielded a single component, causal effects were estimated using the Wald ratio method implemented in TwoSampleMR (v0.6.2)63. For associations that were significant, we additionally performed linkage disequilibrium pruning and conducted a leave-one-out sensitivity analysis, sequentially removing each independent variant and estimating effects of the remainder. We evaluated heterogeneity of estimates using Cochran’s Q test, and horizontal pleiotropy was tested. In this analysis we were sensitive to key assumptions of Mendelian randomization analysis62,119. The assumption of relevance is reasonably well satisfied but the assumptions of independence and exclusion are only partially satisfied. However, in the context of using Mendelian randomization methods to rank candidate genes in which negative findings are as informative as positive findings, this procedure improves bidirectional translation between mouse and human studies of mortality and longevity.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.