Growth of C. elegans strains
C. elegans strains were grown on nematode growth medium (NGM) plates seeded with Escherichia coli OP50 at 20 °C. unc-122p::dsRed or gfp were used as co-injection markers (at 40 ng µl−1) to generate transgenic strains unless noted otherwise. vha-6p::NLS::gfp or vha-6p::mCherry was used as a co-injection marker at 10 ng µl−1 with 65 ng µl−1 of 1 kb DNA ladder (New England BioLabs) to a final concentration of 100 ng µl−1 to generate PY12814, PY12815, PY12816, PY12821, PY12822 and PY12823 (Supplementary Table 1). Strains were generated using standard genetic crosses. The PY12829 strain was obtained by deleting glr-9 via gene editing to generate glr-9(oy224) in the PY12818 strain. The genotypes of all strains were verified by sequencing or phenotypic examination. All experiments were performed using one-day-old adult hermaphrodites grown under well-fed conditions for at least three generations. A list of all strains used in this work is provided in Supplementary Table 1.
Molecular biology
1.1 kb of sequences upstream of and including the glr-9 START codon and 1.9 kb of sequences upstream of and including the srg-42 START codon were amplified from wild-type worm lysate and used to drive I3-specific or I3-selective expression. Expression in I1 was driven using 1.1 kb of sequences upstream of the lgc-8 START codon. To generate lgc-8p::glr-7::mStayGold::SL2::mScarlet-I3 and lgc-8p::glr-9::mStayGold::SL2::mScarlet-I3, mStayGold::SL2::mScarlet-I3 was amplified by PCR from gBlock fragments and used to generate PSAB1400 and PSAB1401 by Gibson assembly (Supplementary Table 2). glr-9 cDNA or genomic DNA was amplified from wild-type worm lysate and tagged with gfp for localization and functional rescue experiments. Since glr-7 has a and b isoforms, the genomic region of glr-7 from the START codon of the glr-7b isoform (X: 2,419,477) to the STOP codon of both glr-7a and glr-7b isoforms (X:2,414,064) was amplified from wild-type worm lysate. These sequences were tagged with fluorescent reporter sequences to examine localization of glr-7. Full-length cDNAs encoding GLR-9, GLR-7a and GLR-7b amplified from worm lysates were cloned into the pME18s vector for expression in HEK293T cells. All plasmids were verified by Sanger sequencing. A list of plasmids used in this work is provided in Supplementary Table 2.
CRISPR–Cas9-mediated genome editing
Deletion alleles of glr-9, osm-9 and fipr-26 were generated using an injection mix containing 0.5 μl of Cas9 (IDT), 1.5 μl of 100 μM trans-activating CRISPR RNA (tracrRNA), 1 μl of 50 μM 5′ CRISPR RNA (crRNA), 1 μl of 50 μM 3′-crRNA, 1 μl of 17 μM dpy-10 crRNA as a co-CRISPR injection marker, and nuclease-free water, for a total volume of 10 μl. No DNA donor was used. Worms were maintained at 25 °C after injection. Three days following injection, dumpy or roller F1 progeny were segregated onto individual plates. After the singled F1 worms laid eggs, parents were lysed and then genotyped using PCR.
To generate the flp-6::loxP insertion strain (PY12288, Supplementary Table 1), the injection mix included 0.5 μl of Cas9 (IDT), 2.8 μl of 34 μM crRNA, 0.2 μl of 100 μM tracrRNA, 2 μl of 26 μM donor DNA, 50 ng µl−1 of column-purified unc-122p::gfp, and nuclease-free water for a total volume of 20 μl. Following injection, worms were kept at 25 °C. F1 progeny carrying the unc-122p::gfp co-injection marker were singled onto individual plates and genotyped. The C-terminal loxP site was inserted first, followed by insertion of the N-terminal loxP site. All tracrRNA, crRNAs, and donor DNAs were obtained from IDT.
To generate nuclear SL2::mStayGold insertions (PY12818, PY12819, PY12820, Supplementary Table 1), the injection mix included 0.5 μl of Cas9 (IDT), 2.8 μl of 34 μM crRNA, 0.5 μl of 100 μM tracrRNA, and pRF4 rol-6(gf) plasmid as an injection marker. Repair templates were obtained by PCR from each plasmid, gel purified and cleaned up using AMPureX beads, and 500 ng was added to the injection mix. Injected worms were singled and 96 F1 progeny were singled from plates that contained rollers, allowed to starve out the plate, and then screened by PCR for the expected change. The sequences of the crRNAs and donor DNA are listed in Supplementary Table 3.
Fluorescent reporter imaging
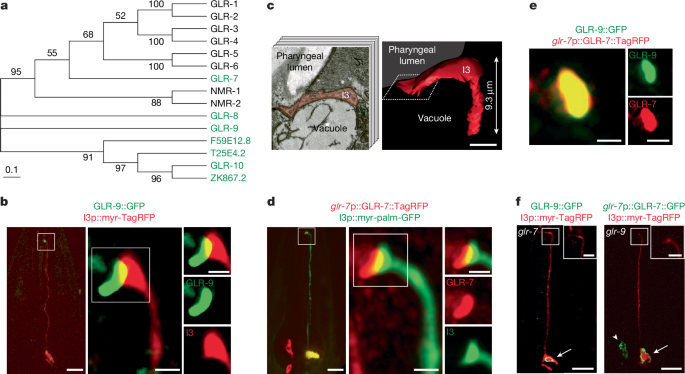
One-day-old adult worms were immobilized with 10 mM (−)-tetramisole hydrochloride (Sigma-Aldrich L9756) and mounted on 10% agarose pads on microscope slides. Images in Fig. 1b, left, and Fig. 1d, left, were acquired on an inverted Zeiss Axiovert with a Yokogawa CSU-X1spinning disk confocal unit and a Photometrics Quantum 512SC camera. Slidebook 6.0 (Intelligent Imaging Innovations, 3i) software was used to acquire images with z-step sizes of 0.2 or 0.27 μm using a 63× oil immersion objective. High resolution images in Fig. 1b, right, Fig. 1d, right, and Fig. 1e were acquired at 0.21–0.23 μm z-intervals using a 100× oil immersion objective on an inverted LSM880 Airyscan confocal microscope using ZEN software (Zeiss). Histograms were adjusted for visualization after Airyscan processing to optimize resolution and contrast. Images in Figs. 1f, 5d and Extended Data Figs. 4c and 5a,b were acquired at 0.2 μm z-intervals using a 63× oil immersion objective on an inverted two-camera spinning disk confocal microscope (Leica DMI6000B with a CSU-W1 spinning disk head and two Andor Neo sCMOS cameras) and Andor IQ3.5 Software. All images were acquired in the Brandeis Light Microscopy Facility (RRID: SCR_025892) and processed using FIJI/Image J.
Reconstruction of I3 sensory ending
Serial section electron microscope (EM) images of I3 were obtained from Wormimage (https://www.wormimage.org). I3 was identified as ‘blue 14’ on the electron microscopy images (with the assistance of D. Hall), specifically from the sections N2T_114522 to N2T_114583 in the legacy series2,60. The TrakEM2 plugin in Fiji was used to manually align I3 images. Following the alignment, manual segmentation was performed using 3dmod v4.11 from the IMOD suite (https://bio3d.colorado.edu/imod/) to generate the 3D reconstruction of the I3 sensory ending.
Acute salt stress assay
Acute responses to high salt were examined using a protocol adapted from ref. 37. Six-centimetre unseeded NGM plates supplemented with the indicated salt concentrations were divided into two sections to assess the behaviours of wild-type and experimental worms in parallel on the same plate. Growth-synchronized 1-day-old adult hermaphrodites were transferred from NGM plates containing 50 mM NaCl to high salt-containing plates to initiate the assay. Each assay included ten worms of each genotype, and the time at which all ten worms were unresponsive to touch with a platinum wire was recorded. Unresponsive was defined as failure to move after prodding with a platinum wire at the nose and tail, and cessation of pharyngeal pumping.
Salt acclimatization assay
Acclimatization experiments were performed using an adaptation of a previously published protocol37. Thirty-five growth-synchronized 1-day adult hermaphrodites were transferred from NGM growth plates (50 mM NaCl) to NGM plates containing 250 mM NaCl, 250 mM KH2PO4 or 250 mM sucrose and seeded with 200 μl E. coli OP50. Worms were maintained at 20 °C for 3 h. Following 3 h of acclimatization, worms were transferred to unseeded plates containing 500 mM NaCl, 500 mM KH2PO4 or 500 mM or 1 M sucrose, and tested for responses after prodding with a platinum wire after 5, 10, 15 and 30 min. Control and experimental worms were examined in parallel. A subset of experiments was performed by researchers blinded to the genotype.
Salt recovery assay
Recovery from paralysis on 400 mM NaCl was examined using an adaptation of previously published protocols14,51. Bacteria-seeded growth plates were supplemented with 400 mM NaCl. Growth-synchronized 1-day-old adult hermaphrodites were transferred from growth plates containing 50 mM NaCl to the assay plates and were examined every 15 min up to 4 h. Each assay included ten control and experimental worms examined in parallel.
Glycerol avoidance assay
The avoidance assay was performed as previously described61. In brief a ring of 8 M glycerol with xylene cyanol (Sigma-Aldrich X4126) was pipetted onto an unseeded NGM plate dried overnight. For each assay, 10 one-day-old adult hermaphrodites were transferred to the centre of the glycerol ring. After 10 min, the number of worms inside and outside the ring was counted. Wild-type and glr-9(oy180) mutants were examined in parallel.
Auxin-induced degradation
To make 4 mM auxin-containing plates, 400 mM auxin (1-naphthaleneacetic acid, Sigma 317918) was dissolved in ethanol and added to NGM containing the indicated salt concentrations. Control plates contained equal concentrations of ethanol alone. Plates were seeded with 100 μl E. coli OP50, dried overnight, and stored at 4 °C wrapped in aluminium foil to prevent light exposure.
At least 70 young adult hermaphrodites were transferred from NGM growth plates to plates containing 250 mM NaCl supplemented with either auxin or ethanol. After 3 h acclimatization, worms were tested for movement on plates containing 500 mM NaCl supplemented with either auxin or ethanol. Worms were examined at 5, 10, 15 and 30 min for responses to touch with a platinum wire. Each assay included 30 worms per condition and per genotype and was conducted over multiple days. Experimental and control strains were examined together in the same assays in each experiment.
Quantification of pumping frequency
Pumping frequency was analysed on NGM agar plates containing 50 mM or 250 mM NaCl and seeded with bacteria. The bacterial solution was prepared by diluting the supernatant of centrifuged E. coli OP50 with LB broth at a 1:50 ratio and 1 ml of this solution was spread to form a thin bacterial layer on a 10 mm plate. Individual worms were moved to these tracking plates and pumping was recorded for 3 min at 40 frames s−1. Worm behaviour was captured using a custom worm tracker consisting of a 760 nm IR LED (LED760L, Thorlabs), a 4× objective lens (XLFLUOR 4× 340, Olympus), and a CMOS camera (BFS-U3-23S6M-C, FLIR). To calculate the pumping rates, 2 min of pumping events from the recording was analysed using custom software written in Python (https://github.com/venkatachalamlab/Yeon-2025-pumping-analysis). In brief, the contour of the worm’s body was extracted from every frame of the video, and the centre line passing through the head and tail vertices and the terminal pharyngeal bulb was identified. A two-dimensional kymograph was then generated using this centre line as the region of interest for each frame. Pumping defined as morphological changes in the pharyngeal grinder was analysed by observing the movement of the corresponding region on the kymograph.
Microfluidics behavioural assay
Behavioural assays using microfluidics devices were performed essentially as previously described62. After degassing the assembled microfluidic devices, the outlet port was loaded with 5% v/v poloxamer surfactant (Sigma P5556) containing 2 mg ml−1 xylene cyanol to eliminate bubbles. Twenty-to-thirty one-day-old adult hermaphrodites were transferred to unseeded NGM plates, washed with water, and loaded into two separate arenas within a single device. glr-9 mutants were assessed in the same device, with wild-type controls placed in the adjacent arena. After allowing the worms to disperse (<5 min), the stimulus flow containing either 75 mM NaCl or 250 mM NaCl and 2 mg ml−1 xylene cyanol for visualization was initiated. 20 min videos were recorded at 2 Hz using a PixelLink camera. Microfluidic devices were cleaned after each experiment by washing with water and 100% ethanol before reuse. Custom MATLAB software was used to process and analyse all recordings, and data were visualized using custom R scripts (https://doi.org/10.5281/zenodo.13748735)63. Mean residency histograms and chemotaxis indices were analysed as previously described62.
Calcium imaging
Imaging in I3 and I1
Custom microfluidic devices for calcium imaging were fabricated as described previously62. Images were acquired using an Olympus BX52WI microscope with a 40× oil objective and a Hamamatsu Orca CCD camera at 4 Hz with 4 × 4 binning. All stimuli were diluted in filtered Milli-Q water, and worms were paralysed with 10 mM (−)-tetramisole hydrochloride (Sigma L9756) dissolved in water prior to loading on the unseeded NGM. Stimuli dilutions were prepared fresh in water on the day of each imaging session. Responses to 30 s of water, 30 s of stimuli and 30 s of water delivered by a perfusion system were recorded in I3. For GCaMP fluorescence analysis, recorded videos were processed using Fiji as previously described62. In brief, images were aligned using the Template Matching plugin, and cell body and background fluorescence were calculated using manually drawn ROIs. Background-subtracted fluorescence intensity values were used for analysis in R Studio. The baseline fluorescence intensity (F0) was calculated as the average fluorescence value over 5 s prior to stimulus onset. R Studio was used to compute the response mean and s.e.m. (https://doi.org/10.5281/zenodo.13748735)63. Peak ΔF/F0 values were calculated as the maximum change in fluorescence relative to F0 within 10 s following stimulus addition. GCaMP6s was driven under the glr-9 promoter for imaging in I3, and under the pha-4-promoter264 for imaging in I1. I1 was identified by its unique anatomical position. Control and experimental worms were interleaved in each imaging session.
Pan-pharyngeal neuron imaging
Volumetric calcium imaging was performed using a microfluidic device as previously described65,66,67 using the OH18562 strain64. Responses were recorded to 60 s water, 30 s 50 mM or 250 mM NaCl, 30 s water delivered via a perfusion system. Pan-pharyngeal neuronal soma imaging was performed using a piezo-driven 60× objective lens (UPLSAPO60XS2, Olympus) and a sCMOS camera (Kinetix, Teledyne) mounted on a spinning disk confocal microscope (Dragonfly 200, Andor). Calcium responses were recorded at a rate of 2.67 volumes per second, with each volume consisting of 25 frames per 15 ms. For volumetric imaging analysis, each of the 20 imaged pharyngeal neurons was identified on the basis of the neuroanatomical map68,69. All 20 neurons were tracked across all frames using the ZephIR multi-object tracking algorithm70. F0 was calculated as the average fluorescence value over 30 s prior to stimulus onset, ΔF was calculated as F − F0.
Heterologous expression and calcium imaging in HEK293T cells
HEK293T cells were provided by L. B. Vosshall at the Rockefeller University. Frozen stocks were stored in liquid nitrogen or at −80 °C. Cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM; High Glucose, Wako) supplemented with 10% (v/v) fetal bovine serum. Cells were routinely passaged at least twice a week. For plasmid transfection, cells were seeded on Type I-C collagen-coated 35 mm glass-bottom dishes (Nitta Gelatin). Just before transfection, the culture medium was replaced with Advanced DMEM/F12 (Thermo Fisher) supplemented with 1% (v/v) GlutaMAX and 2.5 µM 4-phenylbutyric acid. The vectors with or without the glr-9 and glr-7 cDNAs (1.0 µg per dish) were transiently transfected into HEK293T cells using X-tremeGENE HP DNA Transfection Reagent (Roche) according to the manufacturer’s instructions. Cells were analysed 24–36 h after transfection.
To perform calcium imaging, HEK293T cells in glass-based dishes were loaded with Fura-2/AM (2.5 µM; Thermo Fisher) according to the manufacturer’s instructions. After washing the cells with Ringer’s solution (140 mM NaCl, 5.6 mM KCl, 2.0 mM MgCl2, 2.0 mM CaCl2, 1.25 mM KH2PO4, 9.4 mM D-glucose, 2.0 mM sodium pyruvate, and 5 mM HEPES; pH 7.4), the low-cation solution contained 266 mM sorbitol, 5 mM HEPES, 2.3 mM NMDG and 0.1 mM CaCl2 (pH 7.4, adjusted with NMDG) was applied with a custom-built gravity-flow perfusion system. Salt stimulation was performed by 1 mM NaCl solution (1 mM NaCl, 259 mM sorbitol, 5 mM HEPES, 2.3 mM NMDG and 0.1 mM CaCl2 (pH 7.4, adjusted with NMDG)), 50 mM NaCl solution (50 mM NaCl, 167 mM sorbitol, 5 mM HEPES, 2.3 mM NMDG and 0.1 mM CaCl2 (pH 7.4, adjusted with NMDG)), and 140 mM NaCl solution (140 mM NaCl, 5 mM HEPES, 2.3 mM NMDG and 0.1 mM CaCl2 (pH 7.4, adjusted with NaOH)) and 0.5 mM EGTA solution (0.5 mM EGTA-2Na, 282 mM sorbitol, 5 mM HEPES and 2.3 mM NMDG (pH 7.4, adjusted with NMDG)). Fluorescence at 510 nm excited at 340 or 380 nm provided by a LED light source (pE-340fura; CoolLED) was monitored with a CCD camera (ORCA-R2; Hamamatsu Photonics). Fluorescence ratios (F340/F380) were calculated using HCImage software (Hamamatsu Photonics). Positive and negative response magnitudes (ΔF ratio) were defined as the maximum change from baseline. The baseline was defined as the average fluorescence ratio during the 5 s immediately preceding stimulation.
RNA sequencing
Worms were synchronized by bleaching adults, collecting eggs, and then arresting hatched L1s in the absence of food for ∼16 h. Around 1,000 worms were cultured on E. coli OP50 seeded at 1 ml per 10 cm plate at 20 °C until the 1-day-old adult stage. Worms from a total of four 10 cm plates were used per sample. Worms were washed with M9 containing 0.01% Triton X-100 (Sigma T8787) and placed onto either 10 cm OP50 seeded NGM plates or NGM plates supplemented with 250 mM NaCl for 3 h at 20 °C. Worms were then washed off the plates with M9 containing 0.01% Triton X-100. Total RNA extraction was performed using TRIzol (Invitrogen 15596026) and chloroform (Sigma 288306), followed by RNA Clean & Concentrator-5 (Zymo Research R1014) to extract total RNA. The quality and concentration of total RNA was assessed using Agilent High Sensitive RNA ScreenTape analysis (Agilent 5067-5579).
For each sample, 100 ng µl−1 of total RNA was used as input to generate a sequencing library using the xGen Broad Range RNA library preparation Kit (IDT 10009865) with AMPure XP Beads (Beckman Coulter A63880). Agilent DNA ScreenTape (D1000 5067-5582) assays were used to assess the quality of the library prior to sequencing. Libraries were paired-end sequenced at 75 × 75 bases on a NextSeq 1000 system (Illumina). Sequencing reads were adapter trimmed using Trim Galore (https://doi.org/10.5281/zenodo.7598955), mapped to the WBcel235 C. elegans genome, and counted using STAR71 with alignMatesGapMax 2500. Differential expression analysis was performed in R studio with DESEQ2 (ref. 72). The principal component analysis, heat maps and volcano plots were generated using the ggplot package in R. Venn diagrams were created using VennDiagram in R. GO terms, Wormbase, WomCat, Uniprot and PANTHER were used to categorize individual genes listed in Supplementary Data 1.
Statistical analyses
Statistical analyses were performed using Prism v10.4.1 and R Studio 2025.05.01 + 513. Comparisons to wild-type were performed with two-tailed unpaired or paired t-test, Kruskal–Wallis test, two-tailed Welch’s t-test, or one-way ANOVA across the genotype and condition unless indicated otherwise. Corrections for multiple comparisons were performed using Dunnett’s, Dunn’s or Tukey’s tests. Scatter plots and bar graphs were generated and analysed using Prism v10.4.1, and estimation plots were generated using R Studio 2025.05.01 + 513. The statistical tests used, significance values, the number of animals, and number of independent experiments are indicated in the legend to each figure. Exact P values for acclimatization assays are shown in Supplementary Data 2.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.