Animal experiments
All animal work in this study was approved by the Animal Care Committee of The Centre for Phenogenomics (TCP) (AUP 25-0100H) and by the Institutional Animal Care and Use Committee of the Center for Comparative Medicine (CCM) at Baylor College of Medicine (AN-9134 and AN-9136). All mice housed at TCP were maintained under standard housing conditions with a 12-h light–12-h dark cycle (lights on at 07:00; lights off at 19:00), an ambient temperature of 21–23 °C and relative humidity of 40–60%. All mice housed at CCM were maintained under standard housing conditions with a 12-h light–12-h dark cycle (lights on at 06:00; lights off at 18:00), an ambient temperature of 20–22 °C and relative humidity of 30–70%. All mice had ad libitum access to food and water. Timed breeding of C57BL/6J mice was set up, and whole litters were collected from eight independent embryonic time points: E9, E10, E11, E12, E13, E14, E16 and E18. Sexes for all embryos were determined from tail clippings using PCR protocols. Tissues from two female and two male littermates were selected at each time point to undergo dissociation.
FCG experiments
The FCG mice were originally provided by J. Danska, and additional mice were purchased from The Jackson Laboratory (strain 039108). The strain was maintained on a C57BL/6J background. XY mice with testes (XY−-Sry) were bred with wild-type C57BL/6J females to generate the four types of offspring. Caudal cerebellum was collected from E16 and E18 embryos. Genotypes for all embryos were determined from tail clippings using PCR protocols. Cerebella from one XX embryo with ovaries (XX), one XX embryo with testes (XX-Sry), one XY embryo with ovaries (XY−) and one XY embryo with testes (XY−-Sry) were collected at each time point for tissue dissociation.
DNA extraction
DNA was extracted from mouse tissue using the QIAGEN DNeasy Blood and Tissue Kit (QIAGEN, 69506) according to the manufacturer’s instructions.
PCR reactions
Genomic DNA was amplified with the following primer pairs: SX_F (5′-GATGATTTGAGTGGAAATGTGAGGTA-3′), SX_R (5′-CTTATGTTTATAGGCATGCACCATGTA-3′), SRY_F (5′-AGCCCTACAGCCACATGATA-3′) and SRY_R (5′-GTCTTGCCTGTATGTGATGG-3′). X and Y chromosomes were distinguished by PCR as previously described53. In brief, SX_F and SX_R amplified the pseudoautosomal Sly and Xlr genes on the Y and X chromosome, respectively. The presence or absence of the Sry gene in FCG offspring was determined using the SRY_F and SRY_R primers. PCR reactions were performed in a final volume of 20 µl with 1.5 mM MgCl2, 200 µM dNTPs, 0.5 µM primers, 0.4 µl of Phire Hot Start II DNA Polymerase (Thermo Fisher Scientific, F-122L) and the following PCR parameters: initial denaturation at 98 °C for 30 s, 30 cycles with denaturation at 98 °C for 5 s, annealing at 60 °C for 5 s and extension at 72 °C for 5 s, followed by final extension at 72 °C for 1 min. PCR products were analysed on a 1.5% agarose gel with a 100-bp DNA ladder (FroggaBio, DM001-R500) and visualized with ethidium bromide.
Patient-derived PFA-EPN tissues
All PFA-EPN tissues used in this study were obtained with properly informed consent of patients. All experimental procedures were performed in accordance with the Research Ethics Boards (REB 1000055059) at The Hospital for Sick Children.
Tissue handling and dissociation
Fresh patient tumours were collected at the time of surgical resection. Tumour tissue was mechanically dissociated followed by collagenase-based enzymatic dissociation as previously described9. Whole hindbrain was dissected from E9, E10, E11 and E12 embryos; one incision was made between the boundary of the midbrain and hindbrain, and a second between the hindbrain and spinal cord. Whole cerebellum was dissected from E13 and E14 embryos. Caudal cerebellum was collected from E16 and E18 embryos; rostral structures were removed from the cerebellum. All embryonic mouse dissections were performed using Dumont fine forceps (Fine Science Tools, no. 5) under a Leica stereoscope. All mouse tissue was dissociated using the Papain Dissociation System (Worthington Biochemical Corporation, LK003153) according to the manufacturer’s instructions. Cells were filtered using a 40-µm strainer and resuspended in phosphate-buffered saline with 0.1% bovine serum albumin (BSA).
scRNA-seq library preparation and sequencing
Single-cell suspensions were assessed with a trypan blue (Wisent, 609-130-EL) count. We aimed to load 10,000–16,000 cells per sample using the Chromium Controller in combination with the Chromium Single Cell 3’ V3 and V3.1 Gel Bead and GEM Kits (10X Genomics). Individual cells were partitioned into gel beads-in-emulsion (GEMs), followed by reverse transcription of barcoded RNA and cDNA amplification. Individual single-cell libraries with indices and Illumina P5/P7 adapters were generated with the Chromium Single Cell 3’ Library kit and Chromium Multiplex kit. The libraries were sequenced on an Illumina Novaseq6000 or Illumina NovaSeq X Plus sequencer.
Primary cell culture
All samples used in this study were obtained with properly informed consent of patients. All experimental procedures were performed in accordance with the Research Ethics Boards at The Hospital for Sick Children. Patient-derived PFA-EPN (MDT-PFA4, MDT-PFA5, MDT-PFA6, MDT-PFA7, MDT-PFA9 and MDT-PFA15) and ST-EPN (ST1 and ST4) cell lines were established as previously described7. PFA-EPN and ST-EPN cell lines were cultured in EPN-C medium, which is defined as NeuroCult NS-A basal medium (human)(STEMCELL Technologies, 05750), supplemented with N-2 (Thermo Fisher Scientific, 17502048), B27 (Thermo Fisher Scientific, 12587010), 2 mM GlutaMAX (Thermo Fisher Scientific, 35050061), 100 U ml−1 penicillin (Thermo Fisher Scientific, 15240062), 100 µg ml−1 streptomycin (Thermo Fisher Scientific, 15240062), 0.25 µg ml−1 Gibco amphotericin B (Thermo Fisher Scientific, 15240062), 150 µg ml−1 BSA (Sigma-Aldrich A8412), 2 µg ml −1 heparin (Sigma-Aldrich, H3393), 10 ng ml−1 EGF (Sigma-Aldrich, E9644) and 10 ng ml−1 FGF (PeproTech, 100-18B). Laminin-coated Primaria plates (Corning, 353803) were prepared by treating plates with poly-l-ornithine (Sigma-Aldrich, P4957) for 30 min, followed by treatment with laminin (Sigma-Aldrich, P4957). Cells were seeded on laminin-coated plates, cultured in 1% hypoxia (1% oxygen, 94% nitrogen and 5% carbon dioxide), and the medium was replaced every three days.
DIPG cell lines (DIPG004 and DIPG007) were maintained in Matrigel Corning-coated plates with DIPG medium (equal parts of Neurobasal A and Dulbecco’s modified Eagle’s medium with F12 (DMEM/F12) supplemented with B27 (without vitamin A), HEPES buffer (10 mM), MEM sodium pyruvate solution (1 mM), MEM non-essential amino acids (1×), GlutaMAX-I supplement (1×), antibiotic–antimycotic (1×), heparin (2 μg ml−1), human-EGF (20 ng ml−1), human-bFGF (20 ng ml−1), PDGF-AA (10 ng ml−1) and PDGF-BB (10 ng ml−1)).
Hormone treatment experiments
Testosterone (Sigma-Aldrich, T1500), DHT (Sigma-Aldrich, A8380), progesterone (Sigma-Aldrich, P8783) and oestradiol (Sigma-Aldrich, E1024) were reconstituted in high-purity ethanol (VWR, TS61509-5000). All of the cell culture models, PFA-EPN, ST-EPN and DIPG cells were seeded (1 × 106 cells) in laminin-coated Primaria plates. Twenty-four hours after seeding, cells were treated with the indicated hormones and incubated for seven days. For those seven days, each cell line was treated with fresh medium replacement with exogenous hormone(s) every 48 h. Ethanol alone was used as vehicle control. After seven days of treatment, treated and control cells were lifted using accutase (Thermo Fisher Scientific, A1110501) and cell counts (proliferation) and viability were measured using a Countess 3 Automated Cell Counter (Thermo Fisher Scientific). Cell counts were normalized to the vehicle control for each sample. Each treatment group was assessed in two biological replicates, each with two or three technical replicates. A one-way ANOVA followed by a Tukey’s post-hoc test using GraphPad Prism (v.10.2.0) software was performed to assess differences in cell growth.
Western blots
PFA cells were cultured for 24 h with vehicle, 50 nM or 100 nM testosterone or 50 nM or 100 nM DHT to assess the nuclear response of ARs. Total cell lysates from three million cells were prepared using RIPA buffer with added protease inhibitors (Sigma-Aldrich, 11836170001). To obtain nuclear and cytoplasmic fractions, cell pellets from three million cells were resuspended in NE-PER Nuclear and Cytoplasmic Extraction Reagents (78833) as per the manufacturer’s protocol.
Protein concentration was measured using a Pierce BCA Protein Assay (Thermo Fisher Scientific, A55865). Fifty micrograms of whole-cell lysate or fractionated lysate were subjected to SDS–PAGE on a 7% polyacrylamide gel. The resulting protein was semi-dry transferred to a polyvinylidene difluoride (PVDF) membrane (Bio-Rad, 1704272) using the Trans-Blot Turbo Transfer System (Bio-Rad, 1704150), and blocked using 5% skimmed milk in TBST (20 mM Tris, 150 mM NaCl and 0.1% Tween 20, pH 7.5). Membranes were incubated with the following primary antibodies overnight: GAPDH (Abcam, ab9484, mouse, 1:600; Abcam, ab9485, rabbit, 1:2,500); lamin B1 (Abcam, ab16048, rabbit, 1:1,000); and AR (Cell Signaling Technology, D6F11, rabbit, 1:1,000). Membranes were then washed with TBST three times, incubated with anti-rabbit, HRP-linked antibody (Cell Signaling Technology, 7074S, goat, 1:3,000) or anti-mouse, HRP-linked antibody (Cell Signaling Technology, 96714S, goat, 1:40,000) and washed with TBST three times. Membranes were incubated with HRP substrate (Thermo Fisher Scientific, WP20005) for 1 min, before imaging on a Li-COR Odyssey Fc Imaging System.
Western blot images were quantified using ImageJ (v.1.54g; National Institutes of Health; NIH)54. Integrated density values were measured for all bands using a consistent region of interest (ROI) across samples. AR band density was normalized to the housekeeper band. Statistical significance was evaluated using the Wilcoxon test performed on four biological replicates. P < 0.05 was considered statistically significant.
AR inhibition
The PROTAC-based AR inhibitor MTX-23 (MedChemExpress, HY-148771) and enzalutamide (MedChemExpress, HY-70002) were dissolved in dimethyl sulfoxide (DMSO; Sigma-Aldrich, D8418), and DMSO was used as a vehicle control. PFA-EPN cells were seeded at a density of 3,000 cells per well in 96-well Primaria plates (Corning), because PFA-EPN is a slow-growing tumour and requires a higher initial density to establish growth, whereas ST-EPN and DIPG cells were seeded at 1,000 cells per well in the same plates. Twenty-four hours after seeding, cells were treated with enzalutamide or MTX-23 for seven days with varying concentrations. Medium with the indicated dose of MTX was replaced every three days. Cell viability was measured using either a Countess 3 Automated Cell Counter (Thermo Fisher Scientific) or the CyQUANT Direct Cell Proliferation Assay (Thermo Fisher Scientific, C7026), as per the manufacturer’s protocol at end-point. Cell counts were normalized to the vehicle control for each sample. Each treatment group was assessed in two biological replicates, each with three technical replicates. A one-way ANOVA followed by a Tukey’s post-hoc test using GraphPad Prism (v.10.2.0) software was performed to assess differences in cell growth.
Colony formation assay and LDAs
For colony formation assays, PFA cells were dissociated using accutase (Thermo Fisher Scientific, A1110501) and stained with DAPI (Sigma-Aldrich, D9542) to assess viability. Single viable cells were sorted into a 96-well plate format and cultured under hypoxic conditions (1% O2). Serial dilutions of PFA cells were performed, ranging from a seeding density of 1,024 cells per well to 32 cells per well. Six cell doses, each with at least six technical replicates, were tested. To evaluate the effects of hormone treatments, cells were plated in the presence of vehicle control or 50 nM of testosterone, oestradiol or progesterone. To assess the effects of AR antagonists, cells were plated with vehicle control or 5 µM enzalutamide or MTX-23. To examine the effects of AR antagonists in a hormonally active environment, cells were pre-treated with 50 nM testosterone for at least seven days, followed by plating in the presence of vehicle control or 5 µM enzalutamide or MDT-23. Plates were incubated for one to two weeks to allow for colony formation and were subsequently scored as ‘positive’ (colony present) or ‘negative’ (no colony). Media and hormone treatments were replenished every three days. Statistical analysis of limiting dilution data was performed using the ELDA55 R package statmod (v.1.5.0).
Survival analysis
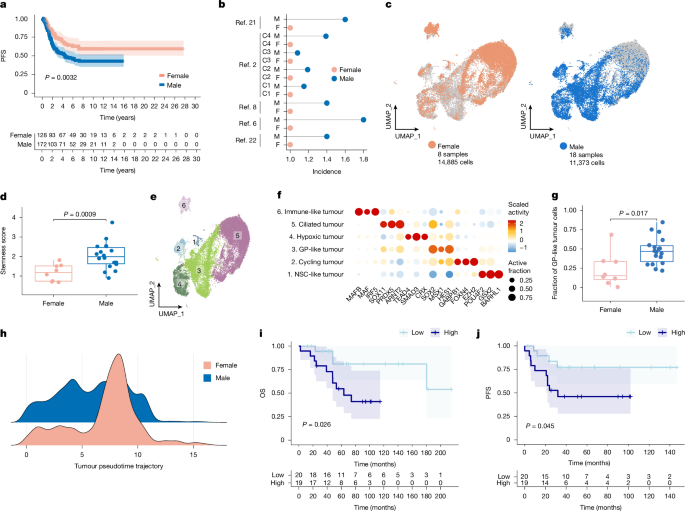
A previously reported22 multicentre cohort of 599 PFA-EPNs was re-analysed to assess the effect of biological sex specifically. PFS was determined using the Kaplan–Meier method and P values were determined using the two-sided log-rank test. Univariable and multivariable Cox proportional hazard regression was used to estimate hazard ratios including 95% confidence intervals. All statistical analyses were performed using the R statistical environment (v.4.3.2), with R packages survival (v.3.5-7) and ggplot2 (v.3.4.4).
scRNA-seq analysis of human PFA-EPN data
Published PFA-EPN cohorts
In addition to our newly generated scRNA-seq PFA database, we added published scRNA-seq PFA 10X Genomics samples from EGAS00001003170 (ref. 9) and GSE125969 (ref. 23). The samples’ copy-number statuses and ependymoma subgroups were classified on the basis of methylation arrays.
Copy-number analysis
The copy-number status of samples in the scRNA-seq PFA cohort was determined using methylation arrays. EPIC and Infinium Human Methylation 450K BeadChip arrays were analysed and combined using the minfi (v.1.34.0) package, with conversion with combineArrays to 450K. The converted samples were combined with normal brain controls (GSE74193, samples from infants and children, 450K BeadChip array)56,57. Using conumee (v.1.18.0) the combined intensities were extracted and normalized on the basis of the controls with a linear regression model58. Segments were calculated with a conumee wrapper based on the DNAcopy package, using the following parameters: significance threshold of 0.001; 50,000 permutations; and a minimum of 5 markers for a changed segment (https://bioconductor.org/packages/release/bioc/html/DNAcopy.html). Methylation-derived copy-number calls were then compared to inferCNV (v.1.4.0)59 calls based on the single-cell data at the single-cell levels compared with reference clusters from the lymphoid cell types60.
Alignment and preprocessing
scRNA-seq samples from the PFA cohort were aligned to the human genome using CellRanger (v.3.1.0)61 to human reference hg19 (v.3.0.0) with default settings. CellRanger-filtered count matrices per sample were preprocessed individually with the Seurat pipeline and filtered further (nFeature_RNA > 400, percent.mt < 3/4 quartile+ 3 times interquartile, genes expressed > 3 cells) and doublets were identified by DoubletFinder (v.2.0.3)62. Filtered samples were then combined with the published cohort (GSE125969) with Seurat. The biological batch effect arises from inherent biological differences between individual PFA tumour samples. Our dataset consists of three distinct cohorts of PFA tumour samples. To account for batch effects within and across these cohorts, we grouped the PFA samples into three separate groups and applied Harmony to correct for batch-related variations within each group. Specifically, we provided two variables for Harmony’s batch correction: sample identity (to address differences between individual samples) and group identity (to correct for batch effects across the three cohorts). Given that we anticipate batch effects both at the individual sample level and across sample groups, we set θ = 1 for both variables to achieve a balanced correction. Single-cell data were normalized with the Seurat SCTransform (v.0.3.2)63 workflow and regularized negative binomial regression with mitochondrial percentage and difference between the G2M and S phase scores regressed out64,65. Principal components to use for nearest-neighbour analysis and UMAP dimension reduction were selected using a quantitative elbow plot approach (https://hbctraining.github.io/scRNA-seq/lessons/elbow_plot_metric.html).
Cell-type annotations
Clusters were identified in the integrated object with the default Seurat Louvain algorithm and cluster markers were identified with the MAST (v.1.25.2)66 test with a 0.25 log fold change threshold, 0.1% expression threshold and minimum 10% expression in cluster. Top markers were selected by log fold change. Clusters were annotated on the basis of published markers and matching to built-in SingleR (v.1.0.1)67 databases and the mouse glial lineage. Tumour and microenvironment separation was validated on the basis of inferCNV analysis of PFA samples with chromosome 1q gain, with lymphoid cells as a reference.
Tumour cell-type trajectory inference
To assist with annotations of tumour subgroups and their hierarchy from stem-like to differentiated, trajectory inference for tumour cells was performed with Monocle3 (v.1.3.1)68,69. The normalized data from Seurat were imported to Monocle3 for UMAP dimensional reduction and cell clustering with default parameters. Root cells (with three standard deviations over the median) were selected on the basis of the CancerSEA (v.0.1.0) database stemness signature score. Next, a principal graph was generated through the UMAP using the learn_graph function, depicting the path through development70. The graph was subsequently used to order cells along the developmental program as pseudotime.
Cell composition analysis
Sex differences in cluster proportions were analysed by dividing the number of cells in the cell type of interest by the total number of cells in the full sample or the sample tumour fraction. Next, a Student’s t-test was used to analyse differences in cell-type fraction between male and female samples.
Differential abundance analysis
Differential abundance between male and female PFA tumour cells was assessed using miloR71 (v1.2.0). A k-nearest neighbour (k-NN) graph was constructed with the buildGraph function from the first 30 Harmony dimensions (d = 30), connecting each cell to its 30 nearest neighbours (k = 30). Differential abundance testing was performed using the testNhoods function, with all other parameters set to default.
Differential expression and pathway analysis
Differential expression between male and female samples within clusters was performed with the MAST test with a log2 fold change threshold of 0.25 and a minimum expression threshold of 10%. To avoid samples with large cell counts, our sex-biased gene lists were further filtered for genes that also had a significant sex bias between samples in average expression across the tumour fraction (AverageExpression function from Seurat). Next, we performed pathway analysis with the gprofiler2 (v.0.2.1)72 package with a multiple-testing g:SCS threshold of 0.1. Average expression of the male-biased gene signature by cell was visualized with the Seurat ModuleScore function.
TF analysis
Raw counts from the combined PFA object were used as input for the single-cell regulatory network inference and clustering (SCENIC) Python pipeline (pySCENIC, v.0.10.3)73,74. GENIE3 was used to infer gene co-expression networks within each cell and combined with the human RcisTarget database included with SCENIC to identify TF targets among the genes. Next, TF networks were binarized as ‘on/1’ or ‘off/0’ with automatic AUCell thresholds. Top regulatory networks by cell type were selected on the basis of fold change after filtering for networks with at least 10% binary difference in expression between clusters. Sex-biased transcriptional networks were selected from the top 20 networks with the highest relative difference in activity by sex within clusters, then filtered for networks with significant sex bias between samples in TF average gene expression across the tumour fraction (AverageExpression function from Seurat). Average expression of the male-biased TF signature by cell was measured with the Seurat ModuleScore function and compared between samples across the tumour fraction with the AverageExpression function.
Microarray analysis and survival
Published datasets generated using Affymetrix HG-U133 Plus 2 chip microarrays (GSE125861 and GSE64415) were used for survival analysis6,23. Only primary PFA samples that received both radiation and gross total resection with full survival data were kept to account for variation in prognosis due to treatment. Analysis was performed as previously described23, with robust multichip average normalization and a pipeline with the affy (v.1.66.0), affyPLM (v.1.64.0) and simpleaffy (v.2.64.0) R packages and the hgu133plus2.db database75,76. For scoring the male-biased gene signatures, the R package jetset (v3.4.0) was used for selecting the optimal probes for each gene. Then, the average of the rma counts was taken for each signature, and samples were divided into low and high groups by the median value77. OS and PFS were correlated with signature expression by the Kaplan–Meier estimator and plotted with the survival (v.3.3-1) and survminer (v.0.4.9) R packages with default parameters (https://cran.r-project.org/web/packages/survminer/index.html).
Comparison of AR expression in bulk RNA-seq data
The Stand Up to Cancer bulk RNA-seq dataset (EGAD00001006046) was published previously7. Primary PFA-EPN and ST-EPN tissue samples were integrated and normalized with the DESeq2 pipeline, and log-transformed counts of AR were compared with a two-sided Student’s t-test.
scRNA-seq analysis of mouse data
Alignment of raw reads
CellRanger (10X Genomics) pipelines (v.3.0.2 and v.5.0.1) were used to process the raw sequencing data for the mouse developing hindbrain and the FCG data, respectively. The raw base call (BCL) files were demultiplexed into FASTQ files with default parameters. The FASTQ files for mouse developing hindbrain data and the FCG data were then aligned to the reference mouse genome mm10 (v.3.0.0) and mm10 (v.2020-A) (10X Genomics), respectively. The count function was used to generate raw gene–barcode count matrices. It is worth noting that, to eliminate batch effects introduced by the technical process, the male sample and the female sample from normal development at E16 and E18 were reprocessed using the same alignment pipeline as was used for the FCG dataset. Alignment quality control metrics are provided in Supplementary Table 1.
Quality control and normalization
Quality control was performed on individual samples before their integration and normalization. Low-quality and outlier cells were identified and excluded from the datasets. We considered low-quality cells as those with fewer than 500 genes expressed, and cells with mitochondrial gene-expression fractions greater than 3/4 quartile + 3 times interquartile. Low-abundance genes were also excluded from the datasets as genes expressed in fewer than ten cells. Outlier cells were identified and excluded as cells with number of genes and unique molecular identifier (UMI) counts greater than 3/4 quartile + 3 times interquartile and less than 1/4 quartile − 1.5 interquartile. Doublets were detected and filtered out using the R package DoubletFinder (v.2.0.3); the doublet rate used the multiplex rate from 10X Genomics. Clusters with high mitochondrial gene-expression fraction but low number of genes expressed and UMI counts were considered as low-quality populations and thus manually removed from further analysis. Individual samples were merged and normalized together using SCTransform (v.0.3.5) using the parameter variable.features.n = 3000 and regressing unwanted variance associated with mitochondrial and cell-cycle content.
Cell-cycle analysis
We used cell-cycle phase-specific annotations to characterize the cell-cycle status of each individual cell64. We assigned a score to each cell on the basis of its expression of S and G2/M phase markers using the CellCycleScoring function from the R package Seurat (v.4.3.0)65. These marker sets should be anticorrelated in their expression levels, and cells that do not express either are probably not actively cycling and are likely to be in G1 phase.
Clustering analysis, visualization and annotation
Clustering analysis was performed with the R package Seurat (v.4.3.0). Linear dimensionality reduction was performed using principal component analysis (PCA) to determine statistically significant principal components. We considered the possibility of a biological batch effect arising from inherent biological differences between individual mouse samples rather than technical variability. A data harmonization method known as Harmony (v.0.1.1)78 with parameter theta = 0 was applied to remove sample-specific biological differences, addressing variations specific to cell types and states. The clusters of cells were identified by a shared nearest neighbour (SNN) modularity optimization-based clustering algorithm from Seurat. We then visualized these clusters using UMAP embedding. Cell types were assigned by manual annotation using known cell-type marker genes and computation of DEGs using the FindAllMarkers function in the Seurat package. The non-default parameters of the FindAllMarkers method are MAST test, minimum log fold change threshold of 0.1 and genes that are detected in a minimum of 10% of cells. DEG markers for each cluster are provided in Supplementary Table 2.
Validation of cell-type annotation in scRNA-seq data
Cell-type predictions in the mouse developing hindbrain data were performed using CoRAL (v.4.0.1; https://github.com/fungenomics/CoRAL), an annotation pipeline in which an ensemble of reference-based annotation tools are used to compute a weighted consensus mapping score to label individual cells.
We used a prenatal mouse cerebellar developmental atlas9 as a reference (five developmental samples from E10 to E18; n = 34,382 cells), which we reprocessed and refined to obtain more granular labels as follows. For each sample, RNA libraries were scaled using Seurat v.4.3.065 to 10,000 UMIs per cell and log-normalized. UMI counts and mitochondrial content were regressed from normalized gene counts, and the residuals were z-scored gene-wise. Dimensionality reduction was performed using PCA on the top 2,000 most variable features, keeping the top 30 principal components as input for clustering with an SNN algorithm (resolution 1). This resulted in 119 clusters, which were labelled on the basis of age, cluster number and cell class information from the original publication9. Cell class was assigned on the basis of the proportion of the original cell type in the new cluster: if the proportion of one original cell type was greater than 75% in the new cluster, that label was used; otherwise, a label consisting of two prominent cell types was used. Twenty clusters were manually reannotated on the basis of marker-gene expression.
Eight different annotation tools were benchmarked and trained on this reference: ACTINN79, scClassify80, SciBet81, singleCellNet82, SingleR67, Symphony83, Seurat and Support Vector Machines. Benchmarking was performed by fivefold cross validation using the average of F1 scores across labels as readout. The predictions of each of these methods were combined into a weighted consensus score using the CAWPE algorithm84, in which the weights for each method are based on their individual benchmarking performance. The final label in the query dataset is assigned to the label with the maximum consensus score.
Pseudotime cell-state trajectory reconstruction
Pseudotime analysis was performed using the R package Monocle3 (v.1.3.1). The preprocessed and normalized data from Seurat were imported to Monocle3 for UMAP dimensional reduction and cell clustering with default parameters. A principal graph was generated through the UMAP using the learn_graph function, depicting the path through development. The graph was subsequently used to order cells along the developmental program as pseudotime using Hes3 expression greater than 1 in E9 cells at the start of the program. To recapitulate the developmental trajectory in the glial lineage, cell-state transition directions were inferred using the Python package CellRank (v.1.4.0)85. CellRank’s PseudotimeKernel function was used to compute directed transition probabilities on the basis of a k-NN graph and Monocle3’s pseudotime ordering. The compute_transition_matrix function was used to compute a cell–cell transition matrix with parameter threshold_scheme = ‘soft’, nu = 0.5.
Reconstruction of gene regulatory and TF networks
The activity of specific TFs in each cell type was inferred in the mouse glial lineage cells using the package pySCENIC (v.0.10.3), implemented in Python (v.3.8.2). Co-expression modules between TFs and putative target genes were estimated by GENIE3. We then performed cis-regulatory motif analysis using RcisTarget, and indirect targets lacking motif-binding sites were removed. The resulting regulatory module (regulon) activities in each cell were then scored and binarized with AUCell. Regulon activity was computed independently in male and female cells of the glial lineage and subsequently integrated on the basis of shared regulons for visualization. The top sex-biased regulons were ranked by z-scaled AUC score. Inferred regulons and their activity across clusters by sex in the dataset are provided in Supplementary Table 4.
Differential expression and pathway enrichment analysis
To perform functional enrichment analysis of the sex-biased gene list in the glial lineage, DEGs between male and female within clusters were identified using the FindMarkers function from Seurat (v.4.3.0), with the parameters MAST test, minimum log fold change threshold 0.1 and genes detected in a minimum of 10% of cells. To ensure that the sex-biased DEGs were not driven by unequal numbers of cells from males and females, an equal number of cells from both sexes was used. Specifically, we kept all of the cells for the sex that had fewer cells. For the sex with more cells, we randomly selected the same number of cells as that used in the sex with fewer cells. For the differential expression comparisons of sex hormones (mouse with ovary versus mouse with testis), sex chromosomes (XX versus XY) and four genotypes (XXF, XYF, XXM and XYM) in the FCG dataset, we used the same pipeline as that described for the developing glial lineage. However, genes that were detected in a minimum of 5% of cells were analysed with the FindMarkers function, with the aim of recovering genes that were highly expressed, but in few cells. The resulting DEGs (Supplementary Tables 5 and 7) were filtered to include only those with an average log2 fold change greater than 0.1, P < 0.05 and an absolute difference in the fraction of expressing cells between comparison groups greater than 0. The filtered DEGs were ranked by average log2 fold change and subsequently analysed using gprofiler2 (v.0.2.1) for pathway enrichment, with multiple-testing correction applied using the g:SCS algorithm (Supplementary Tables 6 and 8).
Integration of PFA tumour data with the single-cell developmental glial lineage
Human genes from the PFA tumour fraction were converted to mouse genes with the convert_human_to_mouse_symbols function from the nichenetr package86. Raw counts from shared genes in PFA tumour and mouse glial lineage were merged and normalized with SCTransform with regression of mitochondrial percentage and difference between the G2M and S phase scores. Cross-species integration was performed with the Seurat Fast integration using the reciprocal PCA (RPCA) workflow on normalized counts (https://satijalab.org/seurat/articles/integration_rpca.html). Cells were integrated in an unsupervised manner using the 3,000 most variable features. Our dataset includes a similar number of male and female tumour cells, ensuring that the selection of the 3,000 variable genes was not biased toward either sex. The primary batch effect in this integration arises owing to differences between the two species: the human PFA tumour dataset and the mouse developmental dataset. To correct for this batch effect, RPCA identifies anchors between the two datasets using the FindIntegrationAnchors() function. In this approach, one dataset is projected into the PCA space of the other, ensuring that the integration process respects mutual nearest-neighbour relationships across species. This mutual neighbourhood constraint helps to identify biologically relevant anchors while minimizing technical artefacts. A key parameter in this process is k.anchor, which defines the number of nearest neighbours considered when selecting anchors between datasets. Increasing k.anchor enhances the alignment strength by incorporating a broader set of neighbours during anchor selection. For our integration, we set k.anchor = 20 to optimize the alignment between the PFA tumour and developing mouse datasets, ensuring a more accurate cross-species comparison. Principal components to use for nearest-neighbour analysis and UMAP dimension reduction were selected using a quantitative elbow plot approach as before, after clustering with the Seurat Louvain algorithm. Clusters were annotated on the basis of their composition of previously annotated mouse and PFA tumour clusters. Markers were identified with the MAST test, 0.25 log fold change threshold, 0.1% expression threshold and minimum 10% expression in cluster Top markers were selected by log fold change. Sex differences in integration were analysed in a similar manner to differences in tumour composition by comparing ratios of integrated cell type in the tumour fraction between male and female samples. Pseudotime and trajectory analysis were performed using the same workflow as for the mouse glial lineage, with ordering initiated from E9 cells with SOX3 expression greater than 1, because HES3 was not detected in this dataset after human–mouse gene conversion.
Deconvolution of PFA tumours with the single-cell mouse developmental glial lineage
For BayesPrism (v.2.2.2) deconvolution, the tumour fraction in each sample was converted to pseudobulk with the Seurat AverageExpression function. Human genes were converted to mouse and raw average counts of shared genes were used for the analysis. A glial lineage reference was constructed by keeping only human and mouse shared genes in the glial lineage raw count matrix, then filtering them for Ribosomal, XY-linked and non-protein-coding genes. The top 300 markers for each glial cluster were identified with the MAST test with a log fold change threshold of 0.6 and at least 5% expression in cluster. Deconvolution was then performed with default BayesPrism parameters. For CHETAH (v.1.18.0) deconvolution, a reference was constructed from the full mouse glial lineage Seurat object and ribosomal genes were removed as suggested in the CHETAH workflow. PFA tumour cells were matched on the basis of 500 genes selected by CHETAH, with a 0.1 threshold and other parameters at default settings. Fractions of deconvoluted cell types and nodes were compared between male and female samples with a Student’s t-test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.