Sex as a biological variable
In consideration of the clinical context, in which TNBC71, ANRE72,73 and their co-occurrence73,74 are predominantly observed in female patients, we have proactively chosen to model the underlying biology in female mice to maximize translational relevance. Furthermore, there are considerable differences in symptom frequency and severity between male and female patients with ANRE75,76. This sexual dimorphism may be partly explained by sex hormone activity, as ER signalling directly modulates NMDAR expression28,77 in the brain.
Ethical oversight
Human-participant oversight was provided by the Northwell Health Institutional Review Board (IRB 20-0150) for the prospective TNBC cohort and the South Central-Oxford A Research Ethics Committee (REC16/YH/0013) for the ANRE study. Vendor-procured specimens received IRB approvals from their respective institutions, as was verified by each vendor. The authors affirm that all human research was performed in accordance with the relevant guidelines and regulations, and that informed consent was obtained. Human samples included in this study are summarized in Supplementary Table 6.
Animals
All animal experiments were performed in strict accordance with the Cold Spring Harbor Laboratory (CSHL) Institutional Animal Care and Use Committee (IACUC) and the National Institutes of Health Guide for the Care and Use of Laboratory Animals. Mice were kept in specific-pathogen-free conditions under a 24 h 12 h–12 h light–dark cycle. Food and water were provided ad libitum, and the temperature and humidity were maintained at 21–23 °C and 40–60% relative humidity. Mice were allowed to acclimatize for at least 1 week after their arrival to the Cold Spring Harbor Laboratory animal facility. Adult female mice between 9 and 14 weeks of age were used. All mice were randomly assigned to experimental groups, and the sample size was predetermined according to the type of experiment, on the basis of common practice. For the majority of experiments, the investigators were not blinded to group allocation as they performed both the experiment and analysis, making blinding not possible. For tumour experiments, 8-week-old female wild-type BALB/c mice were obtained from Jackson Laboratory. As per our IACUC-approved protocol, a tumour size of greater than 20 mm was defined as a humane end point and this limit was never exceeded in any of the experiments. For antibody infusion experiments, 8-week-old female wild-type C57BL/6 mice were obtained from Jackson Laboratory. Outbred wild-type adult female X. laevis frogs (aged 2–4 years) were ordered from Xenopus 1.
Patient samples
Tumour and matched plasma specimens were obtained from three cohorts of individuals with stage I–III TNBC who received surgical resection as part of standard clinical management. All studies were approved by Institutional Review Boards, and written informed consent was obtained from all of the participants. All specimens were fully de-identified before analysis. The Northwell Health cohort (n = 48) was established through an academic partnership under IRB protocol 20-0150. Peripheral blood samples were collected either before or after surgery on the day of surgical resection. Tumour tissue was available for a subset of patients, depending on the availability of excess clinical material. Tumour specimens were viably preserved by mincing fresh tissue and storing in cryopreservation medium at −80 °C. The vendor-supplied cohort (n = 5) comprised paired plasma and tumour specimens obtained from Biochain Institute under IRB-approved protocols with the participants’ informed consent. Tumour sections were preserved as cold acetone-fixed frozen tissue and stored at –80 °C. A third cohort consisted of a single frozen tumour section obtained from Origene (patient ID: FR001D525), collected under IRB-approved protocols with the participant’s informed consent. This sample was used for Visium HD spatial transcriptomic profiling. TNBC status was confirmed in all cases by immunohistochemistry according to standard clinical criteria: ER and progesterone receptor (PR) expression each <1% of tumour nuclei, and HER2 scored as 0–1+ by immunohistochemistry or 2+ with negative fluorescence in situ hybridization. Plasma samples from all cohorts were processed using standardized protocols and stored at –80 °C until analysis. For the Northwell cohort, DFS data were abstracted by manual review of electronic health records by the Northwell biorepository team. DFS was defined as the interval from surgery to first recurrence or death, with censoring at the most recent clinical follow-up. To assess the normal range of anti-NMDAR antibody titres, a panel of plasma samples from healthy women 18–30 years of age (n = 23) was obtained from BioIVT under IRB-approved protocols with the participants’ informed consent. All of the patient samples in this study served as a discovery set, and no formal power calculation was performed.
B cell isolation and antibody cloning
The OX1 antibody was isolated from one patient with a confirmed diagnosis of anti-NMDAR encephalitis (ANRE), on the basis of cerebrospinal fluid antibody testing, with written informed consent as part of a research study approved by the University of Oxford with ethical approval REC16/YH/0013. Peripheral blood mononuclear cells (PBMCs) were isolated from patient blood samples and, after thawing, PBMCs were labelled using fluorophore-conjugated biotinylated GluN1-ATD baits along with B cell surface marker antibodies (CD19, CD20, IgG) for 30 min at 4 °C. GluN1-ATD-binding B cells were enriched by fluorescence-activated cell sorting and sorted into 96-well plates containing single-cell culture medium. After 10–14 days of culture, the cell culture supernatant was prescreened for NMDAR reactivity using GluN1-transfected HEK293 cells by IF microscopy. Positive wells were identified, and antibody sequences were determined using the RNA extraction using the RNeasy kit (Qiagen), followed by reverse transcription and amplification of heavy- and light-chain immunoglobulin variable regions using primers specific for human IgG, followed by Sanger sequencing.
Cell culture
Sf9 insect cells (Thermo Fisher Scientific, 11496015) were maintained in a non-humidified shaker (125 rpm) at 27 °C in CCM3 medium (Cytiva). HEK293S GnTI- cells (American Type Culture Collection, CRL-3022) were maintained in a humidified shaker (115 rpm) at 37 °C and 8% CO2 in FreeStyle 293 expression medium (Thermo Fisher Scientific) supplemented with 2% FBS. 4T1-NMDAR cells were maintained in a humidified incubator at 37 °C and 5% CO2 in RPMI 1640 media with L-glutamine, supplemented with 10% tetracycline-free FBS (Takara Bio) and 100 U ml−1 penicillin–streptomycin. All of the cell lines were routinely tested for Mycoplasma contamination with consistently negative results.
Purification of NMDARs
Human GluN1a–GluN2B NMDA receptor (NMDAR) proteins were expressed in Sf9 cells as described previously78. In brief, Sf9 cells were grown to a density of 6 × 106 cells per ml before infection with 30 ml per 1 l cells of freshly collected P1 virus. Cells were collected 48 h after infection by centrifugation at 3,000g for 20 min at 4 °C and frozen as cell pellets. Cells were thawed and resuspended in HEPES-buffered saline (HBS) with glycine/glutamate (HBSG) buffer (20 mM HEPES, 150 mM NaCl, pH 7.5, 1 mM glycine and 1 mM glutamate) in the presence of 1 mM phenylmethylsulfonyl fluoride (PMSF) and lysed by two passes through an Avestin cell disruptor operating at 10,000 psi. Membrane fraction was collected by ultracentrifugation at 185,511g for 30 min at 4 °C in the Ti-45 rotor (Beckman Coulter). The pelleted membrane was solubilized in HBSG with 0.5% lauryl maltose neopentyl glycol (LMNG) at a ratio of 10 ml buffer per 1g of membrane, with gentle stirring for 30 min. The insoluble fraction was removed by ultracentrifugation and the supernatant was passed through a 5 ml Strep-Tactin Sepharose column (IBA Lifesciences) by gravity flow. The column was washed sequentially with wash buffer 1 (HBSG with 0.002% LMNG), wash buffer 2 (HBSG with 3 mM Mg-ATP), and then wash buffer 1. NMDAR protein was eluted using wash buffer with 3 mM desthiobiotin. For the SK5A-matured-apo structural analysis, NMDAR was initially purified using HBS buffer in the absence of glycine and glutamate. Eluted proteins were concentrated to 500 µl and separated by size-exclusion chromatography using the Superose 6 10/300 column (Cytiva) in a HBSG with 0.002% LMNG running buffer. To generate biotinylated NMDAR, Sf9 cells were infected with a baculovirus construct engineered with an AviTag at the C terminus of GluN1. Biotinylation was performed by incubating concentrated NMDAR protein overnight with 10 mM MgCl2, 4 mM ATP, 0.3 mM d-biotin and 10 μM His-tagged BirA. BirA was removed by passing the reaction mixture over a Co-NTA column, and biotinylated NMDAR proteins were dialysed against HBSG with 0.002% LMNG to remove excess biotin.
Purification of IgG and Fab
Recombinant IgG and Fab proteins were expressed using the BacMam expression system, using a plasmid comprising codon-optimized heavy- and light-chain sequences assembled with a P2A sequence under the control of the cytomegalovirus promoter (Fig. 3a). The amino acid sequence for 003-102 (ref. 15) IgG was obtained from the ABCD database79 (identifier ABCD_AU745), while the sequence for mGO53 (ref. 40) was provided by H. Wardemann. GnTI- cells were grown to a density of 2–3 × 106 cells per ml before infection with 40 ml per 1 l cells of freshly collected P1 virus. Sodium butyrate was added to a final concentration of 5 mM at the time of infection. After 4–5 days, the cell supernatant was collected by centrifugation at 3,000g for 10 min and dialysed against 200 mM NaCl and 20 mM boric acid, pH 9.0 (for IgG) or 200 mM NaCl and 20 mM Tris, pH 8 (TBS) with 20 mM imidazole (for Fab). IgG2 protein32 was purified from hybridoma supernatant. For IgG purification, the dialysed supernatant was passed through a rProtein A column (Cytiva, 17549801; or Genscript, L00464), washed with 1 M NaCl, 50 mM boric acid, pH 9.0, 1 M NaCl and 50 mM Tris, pH 8.0, and eluted with 200 mM NaCl and 50 mM citric acid, pH 3.0. The elution fractions were immediately neutralized with 1 M Tris. For Fab purification, the dialysed supernatant was passed through a metal chelating Sepharose column (Cytiva) charged with cobalt, washed with TBS containing 20 mM imidazole, TBS containing 40 mM imidazole and eluted with TBS containing 250 mM imidazole For structural studies, eluted proteins were dialysed into HBSG with 0.002% LMNG. For mouse phenotyping studies, eluted proteins were dialysed into an excess of PBS (137 mM NaCl, 2.7 mM KCl, 8 mM Na2HPO4 and 2 mM KH2PO4, pH 7.4) and were sterile filtered using a 0.22 μm filter under aseptic conditions. For structural characterization, IgG and Fab proteins were expressed with human constant domains. For mouse phenotyping studies, SK3D and mGO53 (non-binding control) were expressed as mouse IgG2a with a C-terminal Flag tag to verify delivery. IgG2a was selected, as it is the murine equivalent of IgG1 (ref. 80), which predominates in human ANRE15.
ELISA analysis
EvenCoat Streptavidin Coated Plates (R&D Systems, CP004) were incubated overnight at 4 °C with 1.5 μg ml−1 biotinylated GluN1–GluN2B NMDAR protein diluted in wash buffer (HBSG with 0.002% LMNG). NMDAR-immobilized wells were blocked with incubation buffer (wash buffer supplemented with 1% BSA). Monoclonal antibodies (human or mouse Fc) and mouse plasma samples were added to wells in duplicate to a final volume of 50 μl. A standard curve was constructed using 003-102 (ref. 15) engineered with either a mouse (IgG2a) or human (IgG1) Fc at a maximum concentration of 450 ng ml−1. After multiple washes with wash buffer, NMDAR-specific IgG binding was detected using horseradish peroxidase (HRP)-conjugated anti-human (Thermo Fisher Scientific, 31413, 1:8,000 dilution) and anti-mouse (Thermo Fisher Scientific, 62-6520, 1:4,000 dilution) IgG. Monoclonal antibodies, plasma samples (1:20), and secondary antibodies were all diluted in incubation buffer to reduce non-specific signals. To visualize the signal, 100 μl of TMB buffer (R&D Systems, DY999B) was added for 20 min, after which time the reaction was stopped with 50 μl of stop solution (R&D Systems, DY994). Absorbance at 450 nm (relative to 540 nm) was detected using the BioTek Synergy Neo2 Reader (Agilent Technologies). Interpolated concentrations relative to the 003-102 standard curve (Fig. 2f) were determined using the GainData ELISA Calculator (Arigo Biolaboratories).
To assess antibody polyreactivity, Maxisorp plates (Thermo Fisher Scientific, 439454) were coated with dsDNA (10 µg ml−1), LPS (10 µg ml−1) or insulin (5 µg ml−1) in PBS pH 7.4, blocked with 1% BSA in PBS-Tween (0.05%), and developed using protein G-HRP with antibodies diluted in blocking buffer using established protocols81. To test for BSA cross-reactivity, Maxisorp plates were coated with BSA (10 µg ml−1) in PBS pH 7.4, washed with HBSG containing 0.01% LMNG and blocked with 3% non-fat dry milk in the same buffer. Antibodies were diluted in incubation buffer containing 0.5% non-fat dry milk, and detection was performed using cross-adsorbed secondary antibodies.
For the analysis of human patient samples, we modified our indirect anti-NMDAR ELISA to mitigate against the potential for assay interference from non-specific antibody binding. In brief, for each sample, we subtracted the background optical density (OD) value for wells without immobilized NMDAR82, the wash buffer was modified to contain 0.01% LMNG, the blocking buffer was modified to contain 2% BSA and the sample dilution buffer was modified to contain 10 µg ml−1 polyclonal goat IgG (Southern Biotech, 0109-01). Finally, detection was performed using (Fab)2 goat anti-human HRP (Jackson Immunoresearch, 109-036-098, 1:12,000 dilution) to mitigate against heterophile antibody binding.
SPR analysis
Biotinylated NMDAR proteins diluted in HBSG with 0.002% LMNG at a concentration of 10 μg ml−1 were immobilized onto streptavidin-coated sensor chips (Cytiva, BR100398/X100 or 29104992/T200) for 180 s at a flow rate of 5 μl min−1. The running buffer was HBSG with 0.002% LMNG, with a flow rate of 30 μl min−1. IgG proteins were dialysed against an excess of running buffer and then serially twofold diluted to generate five diluted samples per assay. For each antibody, the appropriate concentration range was estimated from ELISA data. IgG samples were injected sequentially from lowest to highest concentration for 400 seconds at a flow rate of 10 μl min−1 according to the Biacore single-cycle kinetics protocol. The final dissociation period was 2,400 s. For all antibodies except for SK3D-matured and OX1-matured, data were collected using the Biacore T200 system; SK3D-matured and OX1-matured data were collected using the Biacore X100 system. A new sensor chip was used for each IgG to avoid interference from residual bound IgG. The experimental data were analysed by fitting a kinetic binding model in the anabel83 R package using the default parameters (no decay or drift).
Single-particle cryo-EM
For all antibodies except for SK5B-germline, purified and concentrated NMDAR and antibody proteins were combined at a 4:1 molar ratio, with target final concentrations of 4 mg ml−1 and 5.5 mg ml−1, respectively, and incubated on ice for 30 min. For structural characterization of SK5B-germline, SK5B-germline IgG was combined with NMDAR (purified in the absence of glycine) at a 1:1 molar ratio, with a final NMDAR concentration of 0.2 mg ml−1 and incubated on ice for 30 min. Amine-to-amine cross-linking was achieved during a 2 h incubation with 2 mM BS3 (Thermo Fisher Scientific, 21580), followed by reaction quenching with 20 mM Tris. The reaction mixture was concentrated 20-fold to achieve a target final NMDAR concentration of 4 mg ml−1. Before blotting, glyco-diosgenin was added to IgG-NMDAR mixtures at a final concentration of 0.1%. Glow discharging was performed using the PELCO easiGlow glow discharge cleaning system (Ted Pella) for 35 s under 15 mA. The grids were blotted with 3.5 μl of sample and plunge-frozen into liquid ethane using the FEI Vitrobot Mark IV system (Thermo Fisher Scientific) at 20 °C with 80% humidity, with a blot time of 4.5 s and under level 7 blot force. Holey carbon grids (Quantifoil R 2/1, R 1.2/1.3 and R 0.6/1) and holey gold grids (UltrAuFoil R 1.2/1.3) were used for blotting. Data were collected as described previously31,43. In brief, micrographs were acquired on the Titan Krios system operating at 300 keV using the Gatan K3 Summit direct electron detector coupled with the Gatan Imaging Filter quantum energy filter with a magnification of ×105,000 (0.827–861 Å px−1). EPU software (Thermo Fisher Scientific) was used for image acquisition, with a defocus range of −0.8 to −2.2 μm, an exposure time of 2–4 s and a total dose ranging from 49 to 71.7 e− Å−1 over 30 frames. Single-particle cryo-EM data processing was performed using WARP84, Relion85 and cryoSPARC86 (v.3.2.0 to v.4.6.0). The precise data-processing steps for each dataset (n = 9) are summarized in Supplementary Figs. 6–8 and Supplementary Note 1. Models for GluN1 and GluN2B were built from Protein Data Bank (PDB) entry 7saa (ref. 87). Models for antibody Fab domains were built using homologous PDB entries88 or ModelAngelo89 where map resolution was sufficient. Antibody constant domains were not modelled as they could not be consistently visualized. Masked local refinements of the ATD heteromer–antibody interface were used to model the interaction interface. Models were built using UCSF ChimeraX90 and WinCoot91, with real-space refinement performed in PHENIX92 (non-bonded weight set to 500). Models built from locally refined maps were used to characterize residue-level paratope-epitope interactions in antibody-NMDAR complexes. A summary of data collection and refinement statistics is shown in Supplementary Tables 4 and 5.
TEVC
Antibody-dependent receptor modulation was tested by TEVC using X. laevis oocytes. Xenopus oocytes were collected from the frogs raised in the animal facilities of Cold Spring Harbor Laboratory and defolliculated before use. Complementary RNAs (cRNAs) for the GluN1 and GluN2B subunits were injected into defolliculated oocytes. Subsequently, the oocytes were incubated in recovery medium (50% L-15 medium (Hyclone) buffered by 15 mM Na-HEPES at a final pH of 7.4) supplemented with 100 μg ml−1 streptomycin and 100 U ml−1 penicillin at 18 °C for 24–48 h. The TEVC recording was performed at room temperature in TEVC buffer (5 mM HEPES, 100 mM NaCl, 0.3 mM BaCl2, 10 mM Tricine, at a final pH of 7.4). The agonist solutions were freshly prepared on the day of measurement from 1 M stock solutions of glycine and glutamate, and final concentrations of 100 μM for each of the agonists were obtained. The antibody solutions were dialysed against excess agonist-containing TEVC buffer before serial dilutions. For each antibody, a total of five serial dilutions was obtained by threefold dilutions of the 333 nM solution, with each antibody concentration ranging from 4–333 nM. Current resulting from NMDA receptor activation was measured using agarose-tipped microelectrodes (0.4–0.9 MΩ) at a holding potential of −60 mV. Data were acquired using the PatchMaster program (HEKA) and analysed by Origin (OriginLab). The relative effect of antibody treatment was calculated as described in Extended Data Fig. 11a.
In vitro transfection experiments
The Grin1-2b expression construct (Fig. 2a; sequence deposited at GenBank PX651955) was generated from the PB-TRE-dCas9-VPR plasmid93 (a gift from G. Church, Addgene, 63800). The insert between NheI and PmeI was replaced with a codon-optimized insert comprising the canonical mouse GluN1 (UniProt, P35438, GluN1a variant) and GluN2B (UniProt, Q01097) amino acid sequences with a P2A linker. No fluorophore was included to minimize immunogenicity94. Stably transfected 4T1-NMDAR cells were generated by co-transfecting parental 4T1 cells (a gift from M. Egeblad, with identity confirmed by genome sequencing) with the Grin1-2b expression construct and the piggyBac transposase vector (System Biosciences, PB210PA-1) using Lipofectamine 3000 according to the manufacturer’s instructions. Stably transfected cells were selected by hygromycin (250 μg ml−1) antibiotic treatment for 14 days, before selecting single-cell clones by limiting dilution. Grin1-2b transcript induction was determined by performing full-length mRNA sequencing on two vehicle-treated and three 10 ng ml−1 DOX-treated (Sigma-Aldrich, D9891) cell aliquots after 48 h of treatment. Raw RNA-seq reads were aligned and quantified using the nf-core/rnaseq95 pipeline with a custom GRCm38 reference containing the Grin1-2b construct, and differential expression analysis was performed using the edgeR package for R96. NMDAR protein induction was determined using immunoblots for GluN1 and GluN2B on cell lysates (permeabilized with 1% Triton X-100) after incubating cells with 0–100 ng ml−1 DOX for 48 h. To minimize potential cytotoxicity from NMDAR expression, MK801 was added to the medium to a final concentration of 20 μM during DOX induction.
4T1-NMDAR insertion sequencing
To sequence the insert region in stably transfected 4T1-NMDAR clones, we designed a custom 396-probe sequencing panel (Twist Biosciences) targeting the 12 kb insert with significant overlap. Genomic DNA was extracted from 4T1-NMDAR clones (n = 5) using the Qiagen DNeasy kit and sequenced on the NextSeq 500 platform. Reads were aligned to the mouse reference genome (GRCm39), and insertion sites (breakend variants) were identified using Manta97 and manually annotated.
Cytotoxicity assay
4T1-NMDAR cells were seeded into each well of a 48-well plate at a density of 2,500 cells per well in complete growth medium as described above. After 24 h, the medium was replaced with medium containing DOX at 10 ng ml−1 and 1 µM MK-801. After an additional 48 h, the medium was replaced with medium containing 47 nM MK-801, as well as IgG (mGO53, SK3D or SK5G) at 33 nM or vehicle (PBS). For dose–response evaluation, cells were incubated with 0–200 nM SK3D IgG. After a final 24 h of antibody incubation, cell viability was measured by sulforhodamine B (SRB) assay (Thermo Fisher Scientific, A14769-06) according to the manufacturer’s protocol.
In vivo tumour experiments
For wild-type BALB/c experiments, a total of 50,000 4T1-NMDAR cells suspended in 20 μl of 1:1 PBS and Cultrex (R&D Systems, 3433-005-01) was injected into the left inguinal mammary fat pad of female BALB/cJ mice (aged 8–10 weeks). For Ighj-knockout BALB/c mouse experiments, female homozygote mice were obtained from Cyagen (C001345). In total, 100,000 4T1-NMDAR cells suspended in 40 μl of 1:1 PBS and Cultrex (R&D Systems, 3433-005-01) were injected into the left inguinal mammary fat pad of female BALB/c Ighj-knockout mice (aged 8 weeks). Once tumours reached a volume of 100mm3 (approximately 12 days after injection), an osmotic pump (Alzet, 1004) containing 75 mg ml−1 filter-sterilized ifenprodil (dissolved in 40% DMSO, 40% water and 20% Cremaphor EL) was implanted subcutaneously. After 48 h, filter-sterilized DOX was added to the drinking water at a final concentration of 0.25 mg ml−1. Mice were monitored regularly until palpable tumours formed, after which the diameter of each tumour was measured every 3–4 days using callipers. Tumour samples were obtained by dissection of mice after euthanasia by cervical dislocation after 14–21 days of DOX treatment for wild-type experiments and 7 days for Ighi-knockout experiments. Tumour samples were separated into fractions for cryo-embedding in optimal cutting temperature compound (OCT) and single-cell suspensions, which were immediately stored in prechilled MACS Tissue Solution (Miltenyi Biotec, 130-100-008) before dissociation. In one experimental sequence, paired tumour and TDLNs were dissected separately are used to prepare tumour suspensions. Blood samples were obtained from tail bleeds and terminal cardiac bleeds and collected in heparin-coated capillary tubes, which were kept on ice before centrifugation at 14,000g for 5 min at 4 °C. The supernatant (plasma) was stored at −80 °C.
Tumour dissociation
Dissected mouse tumours and TDLNs were processed into single-cell suspensions using the mouse Tumour Dissociation Kit (Miltenyi Biotec, 130-096-730) and the gentleMACS Dissociator (Miltenyi Biotec) according to the manufacturer’s protocol. Viably frozen human tumours (minced tissue frozen in cryopreservation medium) were processed into single-cell suspensions using the human Tumour Dissociation Kit (Miltenyi Biotec, 130-095-929) and the gentleMACS Dissociator (Miltenyi Biotec) according to the manufacturer’s protocol. After red blood cell lysis, cell suspensions were cryopreserved in medium containing 10% DMSO, 50% FBS (tetracycline free) and 40% RPMI 1640.
Bulk RNA-seq analysis
For all bulk RNA-seq, total RNA was extracted using the RNeasy Lipid Tissue Mini Kit (Qiagen, 74804) according to the manufacturer’s protocol. RNA was then submitted for full-length paired-end (2 × 150 bp) mRNA-seq with poly(A) pull-down. For mouse tumour profiling, raw RNA-seq reads were aligned and quantified using the nf-core/rnaseq95 pipeline with a custom GRCm38 reference containing the Grin1-2b construct, and differential expression analysis was performed using the edgeR package for R96. Relative proportions of immune cell types for imputed using the mMCP-counter package for R98.
Flow cytometry
To prepare adherent cells for flow cytometry, they were trypsinized and then the trypsin was neutralized with serum-containing medium before cells were resuspended in flow cytometry buffer (PBS with 2% FBS). Cell surface NMDAR protein induction was determined by staining 4T1-NMDAR cells with Alexa-Fluor-488-conjugated 003-102 IgG and measuring the fluorescence signal using the Guava easyCyte system; unstained cells were used as controls. For analysis of cell surface NMDAR expression, cryopreserved tumour suspensions were thawed and stained with LIVE/DEAD Violet (Thermo Fisher Scientific, L23105) before staining with an antibody cocktail containing anti-CD45 (S18009F, PE/Fire 810), anti-GluN1 (003-102, conjugated to Alexa Fluor 488) and anti-GluN2B (IgG2, conjugated to PerCP-Cy5.5). For analysis of mouse intratumoural B cell populations, cryopreserved tumour suspensions were thawed and stained with LIVE/DEAD Violet (Thermo Fisher Scientific, L23105) before staining with an antibody cocktail containing anti-CD45 (S18009F, PE/Fire 810), anti-CD19 (1D3, APC-Cy7), anti-CD138 (Brilliant Violet 711), anti-IgD (11-26 c.2a, Alexa Fluor 488), anti-CD3 (17A2, PE-Cy5) and anti-CD86 (GL1, Brilliant Ultraviolet 737). Staining buffer (PBS with 2% tetracycline-free FBS) included mouse FcR blocking reagent (Miletnyi Biotec, 130-092-575) or human FcR blocking reagent (BioLegend, 422301) as appropriate, as well as Brilliant Stain Buffer (Thermo Fisher Scientific, 00-4409-42). For tumour suspension flow cytometry, cells were analysed using the Dual Fortessa Cell Analyzer (Becton Dickinson). Flow cytometry data were analysed in FlowJo (BD Biosciences). Antibody conjugation was performed using the Lightning-Link kit (Abcam, ab236553) for 003-102 and IgG2.
Western blot
Cells were lysed in TBS containing 1% Triton X-100 and 1 mM PMSF (60 min, 4 °C), and lysates were clarified by centrifugation (21,000g, 60 min, 4 °C). Proteins were separated by SDS–PAGE (8% polyacrylamide), transferred to nitrocellulose and blocked in 5% milk. Membranes were probed with anti-GluN1 (Cell Signaling, 5704, 1:1,000) or anti-GluN2B (Cell Signaling, 4207, 1:1,000) antibodies, followed by HRP-conjugated secondary antibodies. Signals were detected using ECL Prime reagent (Amersham).
scRNA-seq analysis
On the day of sequencing, mouse tumour cell suspensions were thawed and stained with LIVE/DEAD Violet (Thermo Fisher Scientific, L23105) before an antibody cocktail containing anti-CD3 (17A2, PE-Cy5), anti-CD19 (1D3, PE/AF700) and anti-CD45 (S18009F, PE/Fire 810). Staining buffer (PBS with 2% tetracycline-free PBS) included mouse FcR blocking reagent (Miletnyi Biotec, 130-092-575). Cells were then combined with biotinylated NMDAR protein conjugated with Streptavidin-PE-Oligo complexes (10x Genomics, BEAM-Ab conjugates) according to the manufacturer’s protocol (10 Genomics, CG000595). When pooling samples from multiple mice, each suspension was incubated with anti-mouse hashing antibodies (BioLegend, TotalSeq-C). For pilot experiments before the public release of the 10x Genomics BEAM-Ab conjugates, we conjugated NMDARs to TotalSeq-C Streptavidin-PE-Oligo (5:1 molar ratio) and quenched the reaction with 200 nM biotin. Cells were sorted using a SH800 Cell Sorter (Sony); representative compensated gating plots to enrich for NMDAR-binding B cells are shown in Supplementary Fig. 3d. To profile cancer cells, the B cell fraction was mixed with CD45− cells before library preparation. To profile TNBC cryopreserved tumour suspensions, cells were thawed, washed and immediately loaded onto the 10x Chromium chip without sorting or additional processing.
Sorted cells were counted using the Countess II FL automated cell counter using 1:1 acridine orange/propidium iodide (AO/PI) stain (Revvity CS2-0106) before loading the 10× Chromium chips according to the manufacturer’s recommendations. Single-cell gene expression, B cell receptor (BCR), antigen capture (BEAM) and feature barcoding (used for cell hashing) libraries were prepared using the Single Cell 5′ Gene Expression v2 kit with the feature barcoding library prep kit (10× Genomics, 1000263 and 1000541). For pilot experiments before the release of the 10x Genomics BEAM-Ab conjugates, single-cell gene expression and BCR libraries were prepared using the Single Cell 5′ Gene Expression kit v1.1 (10× Genomics, 1000165). TotalSeq-C libraries were generated by spiking in 1 μl of 0.5 μM Totalseq-C cDNA primer (CGGAGATGTGTATAAGAGACAG) to the cDNA amplification step and performing a 2× solid-phase reversible immobilization (SPRI) cleanup on the supernatant of the cDNA SPRI cleanup. Final TotalSeq-C libraries were generated by amplifying cDNA with the AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCG and CAAGCAGAAGACGGCATACGAGATCTCAAGTCGTCTCGTGGGCTCGGAGATGTGTATAAG primers. Libraries were sequenced using the Illumina NextSeq 2000 system. Sequencing data were processed using the Cell Ranger v.7.2.0 multi pipeline (10x Genomics) for BEAM datasets. A custom reference genome was generated to include the Grin1-2b construct. Transcriptional data were analysed using Seurat99 (v.5.1.0) for R. B cell receptor sequencing data were analysed using Loupe VDJ Browser (10x Genomics) and enclone100 (10x Genomics). Germline inference was performed using the partis software library101 using a BALB/c-specific VDJ reference. Mitochondrial mutations were detected using the VarTrix software library (10x Genomics, https://github.com/10XGenomics/vartrix). For downstream analyses, 4T1 cancer cells were defined by inclusion within the transcriptional cluster with high Hmga2 expression and an alt/alt genotype call for the chrM:1576-T-C variant. TNBC cancer cells were defined by copy-number variation profiling implemented in the inferCNV package for R102.
Bioinformatic prioritization of antibody sequences
To prioritize B cell clones most likely to produce NMDAR-specific antibodies, we manually integrated multiple bioinformatic parameters, including B cell differentiation status, immunoglobulin class switching events, clonal expansion patterns and antigen specificity scores. B cells were classified by their differentiation state and whether they had undergone class switching to IgG, indicating a more mature, antigen-experienced phenotype. Clonal relationships were determined by identifying B cells that shared common ancestral sequences within expanded phylogenetic groups (clonotypes), as implemented in enclone100 (10x Genomics). For each B cell, we calculated an antigen specificity score calculated from NMDAR–oligo and albumin–oligo counts103, which quantifies the probability of NMDAR binding versus non-specific albumin binding. These integrated parameters enabled us to rank and prioritize B cell clones on the basis of their likelihood of producing high-affinity, NMDAR-specific antibodies, which were then further validated functionally through recombinant expression.
Validation of mouse-derived antibody sequences
To screen either tumour- or TDLN-derived sequences for binding, expression plasmids were cloned containing variable regions for each antibody sequence (Twist Biosciences) on an IgG1 framework (Fig. 3a). For tumour-derived sequences, antibodies were expressed recombinantly and purified by protein A affinity chromatography as described above, before screening by NMDAR-specific ELISA. For TDLN-derived sequences, given the higher number of sequences, screening was performed by transient transfection. In brief, HEK293 GnTi- cells grown in serum-free Freestyle 293 medium were seeded into 24-well plates at a density of 500,000 cells per well, before transfection using TransIT-2020 (Mirus Bio, MIR5404) according to the manufacturer’s protocol, with the inclusion of untransfected controls and a strong binder (003-102) positive control. Transfection efficiency was verified by GFP fluorescence. After 72 h of transfection, the cell culture supernatant for each well was collected. Specific NMDAR binding was screened by ELISA using serially diluted supernatant starting at 1:2. For TDLN-derived sequences, a positive result was defined as OD > 0.1 with twofold dilution.
In vivo intracerebroventricular antibody infusion experiments
Mice were allowed to acclimatize for at least 1 week after their arrival at the Cold Spring Harbor Laboratory animal facility. We performed stereotaxic brain surgery on isoflurane-anaesthetized mice to implant a sterile stainless-steel cannula (Alzet, 0004760) into the right lateral ventricle. This intracerebroventricular cannula was connected to a subcutaneous programmable and refillable pump (Alzet, iPRECIO SMP-310R) loaded with sterile PBS and set at an infusion rate of 0.5 μl h−1. Post-operatively, mice were allowed to recover for at least 5 days before enrolment in the study, at which point, the contents of the pump reservoir were removed and replaced with either mGO53,SK3D or SK5G (mice selected at random). Seizures were induced with a single intraperitoneal dose of 20 mg per kg filter-sterilized PTZ (Selleck Chemicals, S4587) diluted in PBS. Mice were euthanized if a seizure extended for more than 5 min. Seizure activity in the 10 min after intraperitoneal injection was monitored by filming mice and was scored according to established criteria104. Mice were infused with either mGO53, SK3D or SK5G for 14 days, with the pumps refilled once after 7 days. At the end of the study, mice were perfused with 0.9% saline before dissecting brains, which were cryo-embedded in OCT.
Metabolic cage profiling
Mice were singly housed in cages as part of the Comprehensive Lab Animal Monitoring System (CLAMS) from Columbus Instruments. This platform allows precise monitoring of food and water intake, activity and energy expenditure (by open circuit calorimetry). Data were extracted into tabular format using Oxymax for Windows (v.5.71) and analysed using custom R scripts. Energy expenditure was computed using measured concentrations of oxygen and carbon dioxide, as described previously105. Data entries that generated negative values for VO2/VCO2 or outlier respiratory exchange ratio values (<0.3 or >1.6) were removed. Total activity was computed using CalR (v.2)106. Energy expenditure data were adjusted for relevant covariates using ANCOVA analyses implemented using the glm function in R, according to established guidance in the field107,108. For each ANCOVA analysis, we confirmed that the treatment–covariate interaction term was not significant. In a separate experiment, healthy female C57BL/6 mice received a single intraperitoneal dose of 0.5 mg ketamine after acclimatizing to metabolic cages for 1 week.
Bioinformatic analysis of human data
Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) RNA profiling (microarray) data were downloaded from cBioPortal, while The Cancer Genome Atlas (TCGA)-BRCA RNA-seq data were downloaded from UCSC Xena Browser. TCGA reverse-phase protein array data from TCGA-BRCA were downloaded from cBioPortal. TNBC annotations for METABRIC and TCGA were extracted from a published study109. For each dataset, tumours were stratified by ER, PR and HER2 status, and GRIN1/GRIN2B expression levels were compared using Wilcoxon rank-sum tests. Human thymus scRNA-seq data33 were downloaded from https://cellxgene.cziscience.com/e/63b0a978-5820-4c3a-96a8-af098c1b0166.cxg/ and processed using Seurat99 (v.5.1.0) for R.
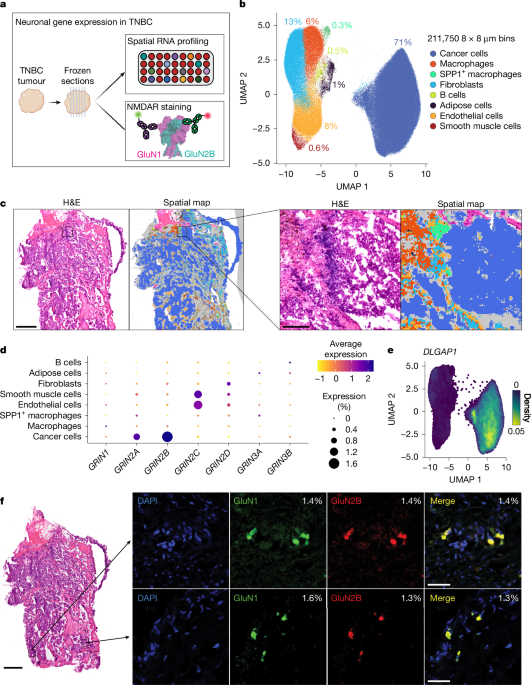
Spatial RNA profiling
Vendor-supplied slides (Origene, patient ID: FR001D525) were fixed in 4% paraformaldehyde (PFA) in PBS for 30 min at room temperature then washed in 1× PBS for 1 min before proceeding to step 3.2 (H&E staining) to 3.7 (decrosslinking) of the Visium FFPE Handbook (10x Genomics, protocol ID CG000684). Slides were imaged using an Olympus VS200. The Human Visium HD Spatial Gene Expression assay (10x Genomics, PN:1000675) was performed according to the manufacturer’s instructions (10x Genomics CG000495), and libraries were sequenced on the Illumina NextSeq 2000 system to target 900 million reads per capture area. Sequenced reads were mapped to the Visium Human Transcriptome Probe Set v.2.0 and aligned to their spots in the paired H&E images using Space Ranger v.3.0.0 (10x Genomics). White-space regions were manually removed using Loupe Browser v.8.0.0 (10x Genomics) before downstream analysis using Seurat99 (v.5.1.0) for R. Visium analysis in Seurat was performed according to the authors’ documentation (https://satijalab.org/seurat/articles/visiumhd_analysis_vignette).
Immunohistochemistry
Immunohistochemical slides were stained on a DISCOVERY ULTRA Research Staining System (Roche) according to the manufacturer’s protocol. Acetone fixation was used for all immunohistochemical stains, without an additional permeabilization step. After fixation and quenching of endogenous peroxidases, the slides were incubated with primary antibodies at 37 °C for 1 h. The Discovery multimer detection system (Discovery OmniMap HRP and/or Discovery DAB, Roche) was used to amplify and/or detect immunosignals. The primary antibodies used were anti-DYKDDDDK (Flag) tag (Cell Signaling Technology, 14793, 1:500) and HRP-conjugated anti-GluN2B (IgG2 (ref. 32), 1:250). Slides were scanned using the Olympus VS200 scanner. Scanned images were analysed using QuPath110 (v.0.5.1)
In situ hybridization (RNAscope)
RNAscope was performed using RNAscope 2.5 HD Detection Reagent Red (322360, Advanced Cell Diagnostics) according to the manufacturer’s protocol. Mouse tissue sections were hybridized with a custom RNAscope probe that was designed to bind specifically to the Grin1-2b insert.
IF analysis
Sections from fresh-frozen TNBC tissue (Origene) were fixed in 4% PFA for 15 min at room temperature and washed in PBS. Sections from fresh frozen mouse 4T1 tumours were fixed in either 4% PFA (for immune cell marker antibodies) at room temperature or acetone (for anti-NMDAR antibodies) at 4 °C for 10 min and washed in PBS. PFA-fixed tissue sections were then permeabilized with 0.2% Triton X-100 in PBS and washed in PBS. All of the tissue sections were blocked with 10% donkey serum in PBS with Tween detergent (PBST, 0.05% Tween-20) and incubated with fluorophore-conjugated primary antibodies diluted in PBST + 3% donkey serum at 4 °C overnight. After incubation, the sections were washed twice in PBST and once in PBS, each for 5 min, incubated with DAPI solution (1 µg ml−1), and mounted in ProLong Glass Antifade Mountant (P36980, Thermo Fisher Scientific). The images in Fig. 1f were taken using the Zeiss LSM710 Confocal Microscope, and images in Figs. 2h and 6d, Supplementary Fig. 2 and Extended Data Figs. 3c and 12e were taken using the Zeiss LSM 980 Confocal Laser Scanning Microscope (Carl Zeiss Microscopy), with a Plan-Apochromat ×20/0.8 NA air objective lens, and a 32-channel gallium arsenide phosphide (GaAsP) detector. Low-power full views of tumour sections were obtained through tile scanning and stitched using Zeiss ZEN Blue v.3.8.2. Raw images were processed using Fiji/ImageJ111, using functions for background subtraction (25 px) and despeckling. Cell classification and quantification were done using QuPath110. For Fig. 1f, the NMDAR antibodies used were 003-102 conjugated to Alexa Fluor 488 and IgG2 conjugated to PerCP-Cy5.5, with conjugation performed using the Lightning-Link kit (Abcam). For Fig. 6d and Supplementary Fig. 2, the NMDAR antibodies were 003-102 conjugated to Alexa Fluor 488 and IgG2 conjugated to Alexa Fluor 594, both used at 1:150 dilution. For all other images, the antibodies used were CoraLite Plus 555 anti-Mouse CD45R (CL555-65139, Proteintech), 1:100 dilution; Alexa Fluor 647 anti-mouse CD3 (100209, BioLegend), 1:100 dilution; and Alexa Fluor 750 anti-pan-cytokeratin (NBP1-48348-AF750, Novus Biologicals), 1:200 dilution.
Quantification and statistical analysis
Data are expressed as the mean ± s.e.m. unless otherwise stated in figure legends. Statistical tests refer to unpaired two-tailed Student’s t-tests unless otherwise stated in figure legends. Wilcoxon rank-sum tests adjusted for multiple comparisons were used for gene expression comparisons between gene expression clusters. All measurements were taken from distinct samples unless otherwise stated in figure legends. Analyses were performed in R (v.4.4.0).
Material availability
On reasonable request, 4T1-NMDAR cells will be available under a material transfer agreement with CSHL. Ighj-knockout mice can be obtained from Cyagen.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.