Plant materials and growth conditions
The Arabidopsis thaliana ecotype Col-0 was used as the WT in this study. The mekk1 (salk_052557), mpk4, summ2 (summ2-8) and mekk2 mutants have been reported previously18,61, and the T-DNA insertion mutant (salk_022911C, sag101) was obtained from the Arabidopsis Biological Resource Center. The adr1 triple (adr1/L1/L2)62, nrg1 double (nrg1.1/1.2)63, adr1 triple/nrg1 double (adr1/L1/L2/nrg1.1/1.2)53, mekk1/summ2, mpk4/mekk2 ref. 32, pad4-1 ref. 64 and eds1-2 ref. 46 were previously described. The adr1/L1/L2/mekk1 or adr1/L1/L2/mpk4 mutants were obtained by crossing adr1/L1/L2 with mekk1 or mpk4 heterozygous mutants, respectively. The genotypes of the mutants were confirmed by PCR using the primers listed in Supplementary Table 1. The N. benthamiana eds1 ref. 65, pad4 ref. 65, epss (eds1a/pad4/sag101a/sag101b)66, nrg1-1 ref. 67 and nrc2/3/4 ref. 68 mutants were previously described.
The Arabidopsis and N. benthamiana plants were grown in soil (Metro Mix 366 for Arabidopsis and LP5 for N. benthamiana) in a growth room at 23 °C, 50% relative humidity, 75–100 µE m−2 s−1 light with a photoperiod of 12-h light–12-h dark. Seedlings were grown on plates containing half-strength Murashige and Skoog medium (1/2 MS) with 0.5% sucrose, 0.8% agar and 2.5 mM MES at pH 5.7 in a growth room with the same conditions as above.
Construction of plasmids
The virus-induced gene silencing (VIGS) constructs of pTRV-RNA1 and pTRV-RNA2 of pYL156-GFP (vector control) and pYL156-MEKK1 and plant expression constructs of MEKK2, HopAI1, SUMM2 and SUMM2ac have been previously reported18,61. The cDNA coding regions of ADR1-L2 (AT5G04720), EDS1 (AT3G48090), SAG101 (AT5G14930), PAD4 (AT3G52430) and RBA1 (AT1G47370) were amplified from Col-0 cDNA using the primers containing BamHI at the 5′ end and StuI at the 3′ end and ligated into a plant gene expression pHBT vector under the control of the CaMV 35S promoter with a HA, FLAG, GFP or mCherry tag at the C terminus. The genomic DNA fragment of gADR1-L1 (AT4G33300, 2,844 bp) was amplified from Col-0 genomic DNA and subcloned into the pHBT vector under the CaMV 35S promoter with a FLAG, GFP or mCherry tag at the C terminus using a one-step cloning kit (Vazyme Biotech). SUMM2G2A, SUMM2ac, SUMM2ac/G2A, SUMM2ac/L22A, SUMM2ac/D12N, SUMM2ac/E13Q and SUMM2ac/D12N/E13Q were generated by site-directed mutagenesis using Platinum Pfx DNA polymerase-mediated PCR.
To make binary vector constructs, gADR1 (AT1G33560, 2,787 bp), gADR1-L1, ADR1-L2, EDS1, RBA1 or SUMM2 mutant variants were subcloned into the binary vectors pMDC32-p2x35S::HA, pMDC32-p2x35S::FLAG or pMDC32-p2x35S::GFP by BamHI and StuI digestions or pTA7002-HA and pTA7002-GFP by XhoI and StuI digestions. To generate the binary vector pCB302-pSUMM2::SUMM2-GFP, the promoter of SUMM2 (2,685 bp upstream of ATG) was subcloned into the binary vector pCB302::GFP by XhoI and BamHI digestions, and SUMM2 was inserted into pCB302-pSUMM2::GFP by BamHI and StuI digestions. The fragment of SUMM2CC (1–480 bp) was subcloned into the binary vector pMDC32-p2x35S::GFP by BamHI and StuI digestions. To generate pCAMBIA2300-p35S::gADR1-L1-nYFP (1–174 amino acids) and pCAMBIA2300-p35S::gADR1-L1-cYFP (175–239 amino acids) plasmids, gADR1-L1 was amplified from pHBT-p35S::gADR1-L1-FLAG and subcloned into pCAMBIA2300-p35S::nYFP and pCAMBIA2300-p35S::cYFP using a one-step cloning kit by BamHI and SmaI digestions. To generate pCAMBIA2300-p35S::EDS1-nYFP and pCAMBIA2300-p35S::PAD4-cYFP plasmids, EDS1 or PAD4 was amplified from pHBT-p35S::EDS1-FLAG or pHBT-p35S::PAD4-FLAG and subcloned into pCAMBIA2300-p35S::nYFP and pCAMBIA2300-p35S::cYFP using a one-step cloning kit by BamHI and SmaI digestions. To generate pCAMBIA1300-p35S::gADR1-L1-TagRFP-HA, gADR1-L1 was amplified from pHBT-p35S::gADR1-L1-FLAG and subcloned into pCAMBIA1300-p35S::TagRFP-HA69 using a one-step cloning kit by BamHI or SpeI digestion. To construct the pCAMBIA1300 binary vector containing the native promoter-driven gADR1-L1 with a GFP tag at the C terminus, the promoter (2,036 bp upstream of the start codon) and genomic sequence were amplified from Col-0 genomic DNA using primers containing XbaI at the 5′ end and StuI at the 3′ end, and ligated into pCAMBIA1300-GFP to obtain the pCAMBIA1300-pADR1-L1::gADR1-L1-GFP construct. HopAI1 was subcloned into the binary vector pMDC32-p2x35S::HA by BamHI and StuI digestions. The SlNRC3 gene from tomato was synthesized into the pUC57 vector (GenScript). To generate pMDC32-p2x35S::SlNRC3-GFP, SlNRC3 was amplified from pUC57-SlNRC3 and ligated into the pMDC32-p2x35S::GFP vector using a one-step cloning kit by BamHI and StuI digestions. The construct of pBRI1::BRI1-GFP has been previously described70.
All the primer sequences have been listed in Supplementary Table 1. The sequences of all genes and mutations were verified by Sanger sequencing.
Agrobacterium-mediated VIGS assays
Agrobacterium tumefaciens strain GV3101 carrying pTRV-RNA1 or pTRV-RNA2 was first cultured in 2 ml LB liquid medium containing 50 µg ml−1 kanamycin and 25 µg ml−1 gentamicin in the 20-ml glass culture tubes overnight at 28 °C in a roller drum and then transferred to 200 ml fresh LB liquid medium containing 50 µg ml−1 kanamycin, 25 µg ml−1 gentamicin, 10 mM MES and 20 µM acetosyringone for overnight at 28 °C with 200 rpm shaking. Cells were pelleted by 1,300g centrifugation, resuspended in buffer containing 10 mM MgCl2, 10 mM MES and 200 μM acetosyringone, adjusted to optical density at 600 nm of 1.5, and incubated at 25 °C for at least 3 h. Bacterial cultures containing pTRV-RNA1 and pTRV-RNA2 derivatives were mixed at a ratio of 1:1 and inoculated into the first pair of true leaves of 2-week-old soil-grown plants using a needleless syringe.
Generation of transgenic plants and growth phenotype analyses
The binary plasmids were transformed into A. tumefaciens strain GV3101 and introduced into Arabidopsis using floral dipping. Transgenic plants were selected by glufosinate-ammonium (Basta, 50 μg ml−1) for the pCB302 vector and hygromycin (50 μg ml−1) for the pMDC vector. Transgenic lines were analysed by immunoblotting using an anti-HA or anti-GFP antibody for protein expression.
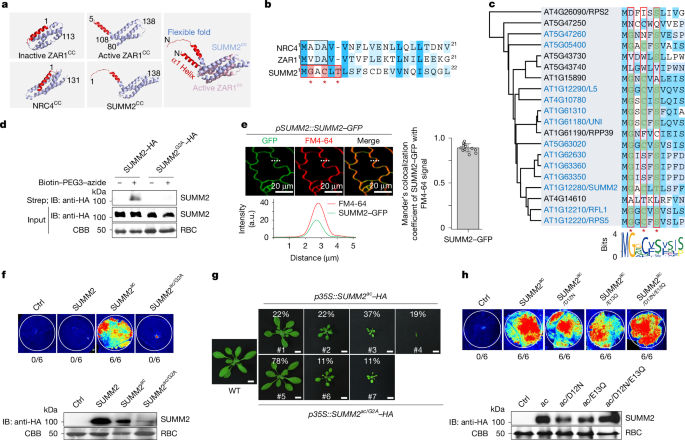
To determine the requirement of the myristoylation motif for the SUMM2 function, p35S::SUMM2ac-HA or p35S::SUMM2G2A/ac-HA was transformed into WT plants. Independent primary (T1) transgenic plants with resistance to hygromycin (50 μg ml−1) were screened by immunoblotting using an anti-HA antibody. On the basis of the severity of growth defects, 46 plants carrying p35S::SUMM2ac-HA in WT were grouped into four categories, with 19% (9 out of 46) of plants exhibiting severe dwarfism, 37% (17 out of 46) showing moderate dwarfism, 22% (10 out of 46) showing weak dwarfism and 22% (10 out of 46) showing no dwarfism. A total of 54 T1 transgenic plants carrying p35S::SUMM2G2A/ac-HA in WT were obtained and grouped into three categories, with 11% (6 out of 54) of plants exhibiting moderate dwarfism, 11% (6 out of 54) showing weak dwarfism and 78% (42 out of 54) showing no dwarfism.
To determine the requirement of ADR1/L1/L2 and EDS1 for overexpression of MEKK2-mediated cell death, p35S::MEKK2-HA was transformed into WT, the adr1/L1/L2 triple or eds1-2 mutant, and followed by screening for hygromycin resistance and immunoblotting using anti-HA antibody. A total of 32 T1 transgenic plants carrying p35S::MEKK2-HA in WT were obtained and further classified into three categories, with 34.4% (11 out of 32) of plants exhibiting severe dwarfism, 40.6% (13 out of 32) showing moderate dwarfism and 25% (8 out of 32) showing weak dwarfism. A total of 34 independent T1 transgenic plants expressing p35S::MEKK2-HA in the adr1/L1/L2 background were obtained, with 11.8% (4 out of 34) of plants showing moderate dwarfism, 50% (17 out of 34) showing weak dwarfism and 38.2% (13 out of 34) showing slightly smaller size than adr1/L1/L2. A total of 48 independent T1 transgenic lines carrying p35S::MEKK2-HA in the eds1-2 mutant with slight dwarfism were obtained.
To examine whether EDS1, PAD4 or ADR1/L1/L2 is required for SUMM2-mediated autoimmunity, p35S::SUMM2ac-HA was transformed into WT, eds1, pad4 or adr1/l1/l2 mutant followed by screening for hygromycin resistance and immunoblotting using an anti-HA antibody. About 50% (23 out of 46) of p35S::SUMM2ac-HA transgenic plants in WT showed severe dwarfism. Transgenic plants carrying p35S::SUMM2ac-HA in eds1-2 or pad4-1 showed weak dwarfism with 46.6% (21 out of 45) or 42.9% (18 out of 42), respectively. Among 42 independent transgenic plants expressing p35S::SUMM2ac-HA in adr1/L1/L2, 33.3% (14 out of 42) of plants showed weak dwarfism. Transgenic plants carrying p35S::gADR1-HA in WT or summ2 exhibited similar dwarfism, with 14.3% (5 out of 35) and 17.9% (7 out of 39), respectively.
To determine the requirement of ADR1/L1/L2 and SUMM2 in HopAI1-mediated cell death, p35S::HopAI1-HA was transformed into WT, adr1/L1/L2 or summ2-8 and followed by screening for hygromycin resistance and immunoblotting using anti-HA antibody. A total of 55 T1 transgenic plants carrying p35S::HopAI1-HA in WT were obtained and further classified into three categories, with 56.4% (31 out of 55) of plants exhibiting severe dwarfism, 38.2% (21 out of 55) showing moderate dwarfism and 5.4% (3 out of 55) showing weak dwarfism. A total of 26 independent T1 transgenic plants expressing p35S::HopAI1-HA in the adr1/L1/L2 background were obtained, with 15.4% (4 out of 26) of plants exhibiting severe dwarfism, 30.8% (8 out of 26) showing weak dwarfism and 53.8% (14 out of 26) showing slightly smaller size than adr1/L1/L2. A total of 36 independent T1 transgenic plants expressing p35S::HopAI1-HA in the summ2-8 background were obtained, with 13.9% (5 out of 36) of plants showing moderate dwarfism and 86.1% (31 out of 36) showing slightly smaller size than summ2-8.
To obtain ADR1-L1 transgenic plants in WT and mekk1 for protein subcellular localization assay, pADR1-L1::gADR1-L1-GFP or p35S::gADR1-L1-TagRFP was transformed into mekk1 (+/−) heterozygous mutant plants, followed by screening in 1/2 MS medium containing hygromycin (50 μg ml−1) under dark conditions for 7 days. Plants with elongated hypocotyls were transferred to 1/2 MS medium for another 10 days, and plants with severe dwarfism were subjected to imaging analysis followed by PCR analysis to genotype the mekk1 (−/−) homozygous background.
To obtain ADR1-L1 transgenic plants in WT and adr1/L1/L2, p35S::gADR1-L1-TagRFP-HA was transformed into WT and adr1/L1/L2-mutant plants, followed by screening in 1/2 MS medium containing hygromycin (50 μg ml−1) under dark conditions for 7 days. Plants with elongated hypocotyls were transferred to soil for another 7 days and were subjected to VIGS assays. The transgenic plants were confirmed with immunoblotting using anti-HA antibody.
To obtain ADR1-L1 transgenic plants in WT, eds1-2 and pad4-1 mutants, pADR1-L1::gADR1-L1-GFP was transformed into eds1-2-mutant or pad4-1-mutant plants, followed by screening in 1/2 MS medium containing hygromycin (50 μg ml−1) under dark conditions for 7 days. Plants with elongated hypocotyls were transferred to soil for another 7 days and were subjected to VIGS assays. The leaf samples were imaged with a confocal or TIRF microscope 2 weeks after infiltration.
Transient expression assays
The Arabidopsis protoplast isolation and transient assay followed a previously reported protocol71. In brief, for co-IP assays, 100 μl of paired plasmid DNA (2 µg µl−1) was mixed with 1 ml of protoplasts (2 × 105 cells per ml) for the polyethylene glycol (PEG)-mediated transfection. The transient assay in Arabidopsis plants has been previously reported72.
For transient assays in N. benthamiana, the indicated constructs were transformed into A. tumefaciens strain GV3010 by electroporation. A single transformant was transferred into the 2 ml LB liquid medium containing 50 µg ml−1 kanamycin and 25 µg ml−1 gentamicin for overnight incubation at 28 °C. Bacteria were harvested by centrifugation at 1,200g and resuspended in the induction buffer (10 mM MgCl2, 10 mM MES pH 5.7 and 200 µM acetosyringone). The Agrobacterium cultures were infiltrated into the leaves of 4-week-old N. benthamiana. Proteins were isolated 2–3 days after inoculation from the infiltrated area and subjected to immunoblotting analysis. Cell death was observed at 2 days or the indicated time after 50 µM dexamethasone infiltration, and pictures were taken under the UV light with the ChemiDoc system. The A. tumefaciens strains carrying Rpi-blb2 or AVRblb2 have been previously described73.
In planta Ca2+ influx assays
Agrobacterium strains carrying the indicated genes were syringe-infiltrated into 3-week-old transgenic GCaMP3 N. benthamiana leaves74. At 48 h later, leaf discs (0.5 cm in diameter) were collected and placed in 200 µl ddH2O in 96-well cell culture plates and equilibrated for 1 h at 23 °C. Followed by 50 μM DEX treatment, GCaMP3 fluorescence was recorded using VANTAstar Microplate reader (BMG LABTECH) with an excitation at 470 nm (15-nm bandwidth) and emission detection at 515 nm (20-nm bandwidth) with 20-µs integration time and 5-min cycle time. Absolute fluorescence values for each experiment were normalized to the untreated control value as F/Feq (where F was the measured fluorescence at a given time point and Feq was the averaged measurement for water-equilibrated control samples with DEX treatment at the final resting time point measured).
Ion leakage assays
Leaf discs (10 mm in diameter) were excised from N. benthamiana plants infiltrated with Agrobacterium at the indicated times. For protein expression under the DEX-inducible promoter, leaves were sprayed with 20 μM DEX before sampling. Leaf discs were washed in 15 ml Milli-Q water for 50 min, then transferred to a 24-well plate with 1 ml Milli-Q water per well, and incubated at room temperature under light. Ion leakage was measured using a conductometer LAQUAtwin EC-33 conductivity meter (Horiba).
Detection of N-myristoylation using click chemistry
The click chemistry-based approach to detect N-myristoylation was carried out based on the previously described protocol with modifications29. In brief, N. benthamiana leaves were infiltrated using a needleless syringe with Agrobacterium carrying the binary vector expressing SUMM2–HA or SUMM2(G2A)–HA. At 12 h later, 10 μM cell-permeable ω-alkynyl fatty acid analogue Alk12 (alkynyl myristic acid; Cayman Chemical) was infiltrated for in vivo metabolic labelling of the myristoylated proteins, followed by a second round of infiltration at 30 h after Agrobacterium infiltration. After 6 h, tissues were collected, and total proteins were extracted with RIPA buffer (1× PBS, pH 7.4, 1% Triton X-100, 0.5% sodium deoxycholate, 0.1% SDS and protease inhibitor cocktail from Roche), and treated with or without biotin–PEG3–azide for click reaction following the manufacturer’s protocol (VectorLabs). Streptavidin affinity purification was used to enrich biotinylated proteins, and SUMM2 and SUMM2(G2A) were detected by immunoblotting using anti-HA antibody.
Trypan blue staining for cell death and DAB staining for H2O2
Detached leaves were submerged in trypan blue staining solution (2.5 mg ml−1 trypan blue dissolved in lactophenol containing an equal volume of lactic acid, glycerol, liquid phenol and ddH2O) or 3,3′-diaminobenzidine (DAB) solution (1 mg ml−1 DAB dissolved in ddH2O, pH 3.8) for overnight incubation. Samples were transferred into trypan blue destaining solution containing lactophenol and ethanol in a ratio of 1:2 or DAB destaining solution containing glycerol, acetic acid and ethanol in a ratio of 1:1:3 and incubated at 23 °C with gentle shaking until completely destained. Samples were observed and recorded under a dissecting microscope.
Co-IP assays in Arabidopsis protoplasts and N. benthamiana
Proteins were expressed overnight in Arabidopsis protoplasts or in N. benthamiana leaves for 2 days. Protoplasts were lysed, and leaves were ground in the extraction buffer (100 mM NaCl, 1 mM EDTA, 20 mM Tris-HCl, pH 7.5, 2 mM NaF, 2 mM Na3VO4, 1 mM dithiothreitol, 0.5% Triton X-100, 10% glycerol and protease inhibitor cocktail from Roche). After centrifugation at 12,500g at 4 °C for 15 min, 250 µl of extraction buffer was added to dissolve pellets, and 20 μl of supernatant was collected for input controls, and the remaining was incubated with anti-FLAG agarose (Sigma-Aldrich) or anti-HA agarose (Sigma-Aldrich) at 4 °C for 2 h with gentle shaking. Beads were collected and washed three times with washing buffer (20 mM Tris-HCl, pH 7.5, 100 mM NaCl, 1 mM EDTA and 1% Triton X-100), and once with 50 mM Tris-HCl, pH 7.5. Proteins were eluted by 2× SDS–PAGE loading buffer and boiled at 94 °C for 5 min. Immunoprecipitated and input proteins were analysed by immunoblotting using the indicated antibodies.
Plant RNA isolation and RT–qPCR analyses
Total RNA was isolated from 2-week-old seedlings grown on 1/2 MS using TRIzol reagent (Invitrogen). One microgram of total RNA was treated with RNase-free DNase I (NEB) and then was reverse transcribed to synthesize the first-strand cDNA with M-MuLV reverse transcriptase (NEB) and oligo (dT) 18 primer. Quantitative PCR with reverse transcription (RT–qPCR) analysis was performed by iTaq Universal SYBR green Supermix (Bio-Rad) with a Bio-Rad CFX384 Real-Time PCR System (Bio-Rad) using primers listed in Supplementary Table 1. The expression of indicated genes was normalized to UBQ10. The data analysis was performed using one-way ANOVA followed by Tukey test for multiple comparisons.
Confocal fluorescent microscopic analyses
N. benthamiana leaves were infiltrated with Agrobacterium carrying the suppressor of silencing HC-Pro and different combinations of fluorescent protein-tagged constructs, as shown in the figures. In brief, gADR1-L1–nYFP/gADR1-L1–cYFP or gADR1-L1–TagRFP was co-expressed with DEX::SUMM2ac–HA or DEX::SUMM2–HA for 2 days in N. benthamiana, followed by 50 μM DEX treatment for the indicated time before imaging analysis using a laser scanning fluorescence confocal microscope-Leica TCS SP8 (Leica).
For Arabidopsis transgenic plants carrying gADR1-L1–TagRFP driven by the 35S promoter or gADR1-L1–GFP driven by the native promoter in the WT or mekk1-mutant background, cotyledons of 2-week-old soil-grown transgenic lines were screened by Leica DMi8 with appropriate filters and light sources for detecting TagRFP and GFP fluorescence, and further imaged using a Leica Stellaris 5 confocal microscope. For plasma membrane staining, leaves were infiltrated with FM4-64 staining solution (10 μM).
The excitation wavelength was 488 nm for GFP, 514 nm for YFP, 554 nm for TagRFP, 559 nm for FM4-64 and 587 nm for mCherry. Emission was detected at 493–530 nm for GFP, 510–540 nm for YFP, 570–630 nm for TagRFP and 570–670 nm for FM4-64. In all multicolour imaging, signals were acquired using line-sequential scanning. The pinhole was set at 1 Airy unit. Leica LAS X software was used to adjust brightness and contrast and generate maximum Z-projection and 3D images. Fluorescence intensity profiles from selected regions of interest were generated using the Plot Profile plugin in Fiji. The fluorescence intensity was normalized to the maximum intensity observed for each fluorophore and presented as arbitrary units (a.u.). The number of ADR1-L1–TagRFP puncta was quantified in Fiji using the Analyze Particles function with parameters set to a size range of 0.08–1 μm2 and circularity of 0.1–1.
TIRF microscopic analyses
TIRF microscopy was conducted using a Leica DMi8 Infinity system equipped with an HC PL APO ×100/1.47 numerical aperture oil immersion TIRF objective (Leica Microsystems). The critical angle was adjusted to maximize the signal-to-noise ratio at the plasma membrane. Images were processed using the THUNDER computational clearing algorithm in LAS X software (Leica Microsystems), with the adaptive strategy, refractive index 1.33 and water as the mounting medium. This approach improves image contrast by suppressing out-of-focus blur while preserving the diffraction-limited lateral resolution (approximately 200–250 nm). Kymographs were generated in Fiji using the Multiple Kymograph plugin from TIRF image sequences consisting of 80 frames acquired over 10 s (120 ms per frame; exposure time of 95 ms). Laser 488 was used for GFP and YFP excitation, and laser 561 for TagRFP excitation. Emission for GFP and YFP was detected using a filter cube GFP for TIRF; for TagRFP, it was detected using a filter cube mCherry for TIRF. 3D projections were created from z-stack images acquired with a z-step of 0.05 µm using the LAS X software and colour coded according to depth to highlight the vertical distribution of the structures.
Single-particle tracking and diffusion analysis
Single-particle tracking was performed on TIRF timelapse image sequences acquired at 8 frames per second with an exposure time of 95 ms. Fluorescent particles were detected and tracked using the TrackMate plugin in Fiji75. Spot detection used the Laplacian of Gaussian detector with an estimated particle diameter of 0.3 µm, and trajectories were reconstructed using the linear assignment problem tracker with a maximum frame-to-frame linking distance of 0.4 µm. Only trajectories with a minimum of 10 consecutive frames were retained for further analysis.
MSD analysis and diffusion coefficient (D) estimation were performed using a custom Python pipeline. Time-averaged MSDs were calculated for each trajectory, and D values were extracted by fitting the first five points of the MSD curve to a linear model, MSD(t) = 4Dt + offset. Trajectories with a poor goodness of fit (R2 ≤ 0.8) were excluded from downstream analyses. Diffusion coefficients were log10 transformed and visualized as frequency distributions, and Gaussian fits were applied to determine peak log(D) values per cell. Average MSD curves were generated from pooled high-quality trajectories.
All computational analyses and visualizations were performed in Python (v3.10), using the NumPy76, Pandas77, SciPy78 and Matplotlib79 libraries. This pipeline conceptually follows established methods for single-molecule motion analysis in plants80, adapted here to a Python-based workflow.
Fluorescence intensity quantification of single particles
Fluorescence intensity of ADR1-L1–TagRFP particles was quantified from raw, unprocessed TIFF images acquired by TIRF microscopy (exposure time of 95 ms). To facilitate particle localization, images were converted to 8-bit format for compatibility with the TrackMate plugin in ImageJ75. Spots were detected using the Laplacian of Gaussian detector with an estimated particle diameter of 0.3 µm and a quality threshold of 20. Sub-pixel localization and median filtering were enabled for spot detection. Fluorescence intensity values were extracted from the original raw TIFF images in ImageJ using the coordinates of the detected particles. Background fluorescence was estimated by measuring the mean grey value within a representative region of interest (ROI) placed in a signal-free area of each image, and this value was subtracted from all particle measurements to obtain background-corrected fluorescence intensities.
Spatial density analysis of ADR1-L1–TagRFP particles
To assess the spatial organization of ADR1-L1–TagRFP particles at the plasma membrane, single-particle coordinates were extracted from TIRF microscopy images using ImageJ. Particle centroids were identified using the TrackMate plugin75, with sub-pixel localization enabled. Spatial clustering was visualized by applying a 2D Gaussian kernel density estimator to the extracted coordinates using the stat_density_2d_filled() function from the ggplot2 package81 in R (v4.3.1). Heatmaps depict local particle density, with warmer colours indicating regions of higher ADR1-L1–TagRFP concentration. Individual particles are shown as overlaid white points.
To test deviations from complete spatial randomness in ADR1-L1–TagRFP particle distributions, quadrat count analysis was performed using the R package spatstat82. The ROI for each cell was subdivided into a 9 × 8 grid of equal-area quadrats. Observed particle counts per quadrat were compared with the expected Poisson distribution under complete spatial randomness using a chi-squared (χ2) goodness-of-fit test. Distributions with P < 0.05 were considered significantly clustered.
Colocalization analyses
Colocalization of fluorescently tagged proteins was analysed using the JACoP plugin83 in ImageJ. ROIs were manually defined, and signal thresholds were individually adjusted for each fluorescence channel to minimize background. Colocalization was quantified using Mander’s overlap coefficient, and results were averaged across multiple biological replicates, as indicated in the figure legends.
Bimolecular fluorescence complementation and FRET-FLIM assays
For bimolecular fluorescence complementation assays, protoplasts were transfected or N. benthamiana leaves were infiltrated with Agrobacterium carrying the suppressor of silencing HC-Pro and different pairs of bimolecular fluorescence complementation assay constructs, as shown in the figures. Fluorescence signals were examined 12 h later using a Leica TCS SP8 laser scanning confocal microscope (Leica). The excitation wavelengths of YFP and autofluorescence of chlorophyll were 514 nm and 630 nm, respectively. The emission wavelengths for YFP and chlorophyll were 490–530 nm and 640–700 nm, respectively. The pinhole was set at 1 Airy unit. Imaging analyses were performed using Leica Application Suite X (LAS X) software.
The FRET-FLIM assays were performed as previously described84. In brief, protoplasts were transfected with different pairs of GFP and mCherry constructs, as indicated in the figures. Fluorescence signals were examined 12 h after transfection under a Leica TCS SP8 confocal microscope. The FRET-FLIM was analysed using LAS X software. The excitation wavelengths of GFP and mCherry were 488 nm and 588 nm, respectively. The emission wavelengths for GFP and mCherry were 490–530 nm and 590–620 nm, respectively. The GFP fluorescence lifetime (τ) in a specific ROI was measured by Leica LAS X software, shown in the figures as an average of 14 randomly measured protoplasts for each pair of proteins. The FRET efficiency (E) was calculated by the formula [E = 1 − (τDA/τD)] (where τDA represents the GFP lifetime of the donor in the presence of the acceptor and τD represents the GFP lifetime of the donor alone). The statistical analysis was performed using one-way ANOVA followed by Tukey test for multiple comparisons.
BN-PAGE assays
Total proteins from protoplasts or N. benthamiana leaves were extracted with 1× native PAGE sample buffer (Invitrogen) containing 1% digitonin and protease inhibitor cocktail. Protein samples containing 0.25% Coomassie G-250, along with protein molecular weight marker NativeMark unstained protein standard (Thermo Fisher Scientific), were loaded and run on a native PAGE 3–12% Bis-Tris gel (Invitrogen). Proteins were then transferred to polyvinylidene difluoride membranes (Bio-Rad) using NuPAGE transfer buffer with a Trans-Blot Turbo Transfer System (Bio-Rad), following the manufacturer’s instructions. Proteins were fixed to the membranes by incubating with 8% acetic acid for 15 min, washed with water and left to dry. Membranes were subsequently reactivated with methanol to visualize the unstained native protein marker. Membranes were immunoblotted with the indicated antibodies.
Protein structure prediction and analysis
The 3D structures of the type II CNL proteins and ADR1s were predicted using AlphaFold2 (refs. 23,85). Subsequently, the predicted structures were calculated at the spatial positions of membrane boundaries via the Positioning of Proteins in Membranes server24,86,87. The visualization of the protein structures was performed by PyMOL88.
Antibodies used in this study
The following antibodies and dilutions were used in this study: anti-HA-peroxidase (1:2,000; Roche); anti-FLAG-peroxidase (1:2,000; Sigma-Aldrich); anti-GFP (1:2,000; Roche); anti-mouse lgG, horseradish peroxidase (HRP)-linked (1:10,000; Cell Signaling Technology); anti-pan myristoylation (1:1,000; MyBioSource); anti-PR1 (1:2,000; Agrisera); anti-rabbit IgG, HRP-linked (1:10,000; Cell Signaling Technology); anti-MYC-HRP (1:2,000; BioLegend); and HRP-labelled streptavidin (10,000; Thermo Fisher Scientific).
Quantification and statistical analysis
Data for quantification analyses are presented as mean ± s.e.m., s.d. or minimum–maximum boxplot. Boxplots show the first and third quartiles as bounds of box, split by the medians (lines), with whiskers extending to the minimum and maximum values. The statistical analyses were performed by Student’s t-test or one-way ANOVA test, followed by the Tukey test. Group comparisons of fluorescence intensity data were assessed using a generalized linear model with a gamma distribution and log link function using the lme4 package89 in R (v4.3.1). Pairwise differences between treatments were evaluated using Tukey-adjusted post-hoc tests. Data analyses and figure generation were conducted using GraphPad Prism 8 (v8.0.1) or R (v4.3.1). The number of biological replicates is indicated in the figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.