Ethics
This research complies with all relevant ethical regulations. The study protocol (NHSR-8429) was determined to be not human subject research by the Broad Institute Office of Research Subject Protection as all data analysed were previously collected and de-identified. Use of SPARK data for this research was approved by SFARI (project 3350.2).
UK Biobank, All of Us, and SPARK WGS data
All WGS data analysed in this work were generated in previous studies. For all cohorts, PCR-free methods were used in library preparation and sequencing was done on Illumina NovaSeq 6000 machines. WGS from UKB23 was performed on libraries prepared with NEBNext Ultra II PCR-free kit (New England Biolabs) using blood-derived DNA from 490,401 individuals at the deCODE facility in Reykjavik, Iceland and the Wellcome Sanger Institute (Sanger), Cambridge, UK. Sequencing reads were aligned to human reference build GRCh38 graph genome with Illumina DRAGEN Bio-IT Platform Germline Pipeline v.3.7.8. Samples were sequenced to an average coverage of 32.5×.
WGS from the NIH All of Us v8 cohort24 was performed on blood and saliva-derived DNA from 414,817 individuals (365,918 blood-derived and 48,899 saliva-derived samples). Sequencing reads from libraries prepared with PCR-Free Kapa HyperPrep library construction kit were aligned to human reference build GRCh38 with Illumina DRAGEN Bio-IT Platform Germline Pipeline v.3.4.12. Samples were sequenced to an average coverage of 37.9×.
WGS from the SPARK cohort of the Simons Foundation Autism Research Initiative (SFARI)25 was performed on saliva-derived DNA from 12,519 individuals. Sequencing reads from libraries prepared with Illumina DNA PCR-Free Library Prep kit were aligned to human reference build GRCh38 with BWA-MEM by the New York Genome Center (NYGC) using Centers for Common Disease Genomics project standards. Samples were sequenced to an average coverage of 42×. Details of saliva sample collection, DNA extraction, and sequencing were described in ref. 68 (which analysed data from sequencing waves WGS1–3 of the SPARK integrated WGS (iWGS) v.1.1 dataset; here we analysed WGS1–5, which included additional samples included in subsequent sequencing waves).
Genotypes of genome-wide human genetic variants (SNPs and insertion–deletions) were previously generated for all three datasets. For UKB, we analysed genotypes previously imputed into UKB SNP-array data55 using the TOPMed reference panel69 (as the final UKB WGS genotype call set was not available at the time of analysis). For AoU and SPARK, we analysed genotypes previously called from WGS using DRAGEN (AoU24) and DeepVariant (SPARK25).
Selection of viral reference genomes
We selected a panel of 31 viruses for which to profile viral DNA load (Supplementary Table 1) based on previous work that identified the most prevalent viruses observed in blood-derived DNA sequencing data from 8,240 individuals9. Additional viral reference genomes were included to more comprehensively represent viral families previously observed. For example, adenovirus was represented by Human adenovirus type 7 and human adenovirus E (type 4) reference genomes, as these are two of the most commonly observed in adults70, and polyomavirus was represented by a set of common types (1/BK, 2/JC, 3/KI, 4/WU, 5/MC, 6, 7)71,72. Undetected herpesviruses (HSV-2 and VZV) were added given the high general prevalence of EBV and HHV-7. To maximize representation of the diversity of anellovirus genomes while limiting the size of the viral reference panel, we selected a representative from each of the five recently described anellovirus clades with available NCBI reference genomes73 (Extended Data Fig. 1a).
The rationale for prioritizing a smaller set of viruses to profile (rather than attempting to comprehensively characterize viral diversity) was that our main goal was to identify effects of human genetics, age, sex and environmental exposures (such as smoking) on viral DNA load, and we only had statistical power to detect such effects for commonly observed viruses. Working with a smaller set of common viruses allowed us to perform careful QC on WGS-based quantifications of viral DNA load, which was important given the potential for read alignment artefacts.
Measurement of viral DNA presence and abundance in WGS samples
In the UKB (blood), AoU (blood and saliva) and SPARK (saliva) WGS datasets, we first extracted unmapped reads (that is, reads that did not align to the GRCh38 human reference genome) and reads that aligned to the chrEBV decoy contig. We realigned these reads to the reference panel of 31 viral genomes (merged into a single reference for alignment) using BWA-MEM74 (v.0.7.18) with 4 threads (-t 4). We took this read-mapping-based approach following Moustafa et al.9, who observed minimal additional detection of viral sequences from blood-derived WGS upon using de novo assembly followed by protein-based search.
After realignment, each virus’ genome was then scanned to identify regions with excessive numbers of alignments suggestive of accumulated misalignments originating from some other source of DNA (for example, mismapped human DNA) by first computing alignment coverage aggregated across all samples in each dataset (UKB, AoU blood, AoU saliva and SPARK, each analysed separately). To do so, for each sample, we used mosdepth75 (v.0.3.9) to compute depth-of-coverage in each 500-bp window of each viral reference genome (–by 500); we skipped per-base depth output (-n) and mate overlap/CIGAR corrections (–fast-mode), restricted to reads passing default filters on SAM flags (-F 1796, which excludes duplicate reads), and filtered to reads with mapping quality ≥5 (-Q 5) for which their mates mapped to the same reference genome with an insert size in the range 100–1,000 bp (-l 100 -u 1000). The results of these filters were not sensitive to the choice of the mapping quality threshold; using a more stringent threshold (-Q 20) affected only a few percent of reads attributed to common viruses and had a negligible effect on downstream genetic association analyses.
Upon computing the 500 bp-resolution coverage profile of each viral genome of each sample, we computed the coverage profile of each virus (that is, for each 500-bp bin, what fraction of samples had non-zero coverage) within each cohort (UKB, AoU blood, AoU saliva and SPARK). For each virus, for each cohort, we flagged a subset of 500-bp regions for exclusion based on having coverage exceeding the following threshold:
$$4\times (Q_3-Q_1)+Q_3+5$$
where Q1 and Q3 are the first and third quartiles, respectively, of the distribution of alignment coverage across all 500 bp regions of that virus’ genome in that cohort. This expression corresponds to a lenient ‘Tukey fence’, which is an outlier removal boundary defined based on adding a multiple of the interquartile range (Q3–Q1) to the third quartile (Q3). We used a lenient Tukey fence to retain regions with modest elevation of coverage (as such regions may still have a majority of alignments derived from the viral genome), and we added a constant offset of 5 to handle situations in which the first and third quartiles have the same value. Applying this bin-level filtering strategy per virus per cohort helped handle cohort-specific error modes of false positive viral alignments that might arise from heterogeneity in WGS data generation and processing (for example, details of how reads had previously been aligned to the human reference genome, which impacted which reads did and did not map to human chromosomes).
For association analyses of viral DNA load with biological and clinical phenotypes, these measures of viral DNA load were then converted into binary ‘viral DNA positivity’ indicators of presence or absence of reads from a given virus in each individual. For genetic association analyses of viral DNA load in blood, the number of viral read pairs mapping to each genome (obtained by summing mosdepth 500-bp bin depth values across non-excluded bins and multiplying by 500/300 = (bin size)/(bases per read pair)) was inverse-normal transformed to capture quantitative information about viral abundance while limiting the influence of outlier samples. In saliva samples, in which viral reads were often much more abundant, two phenotypes were generated for genetic association analyses: viral DNA positivity (binary presence or absence of reads), and a quantitative abundance metric in which we applied inverse-normal transform to non-zero values (that is, masking individuals with no reads from a given virus) after normalizing read pair counts for library size. This allowed for the possibility of observing effects on viral prevalence but not abundance and vice versa. For associations with sex (Fig. 2c,d), the cohort used for analysis (UKB or AoU for blood; AoU or SPARK for saliva) was chosen to maximize prevalence and/or representation of an age range with a larger sex difference; for SPARK, which used a family design, analyses of sex effects were restricted to parents.
Estimation of the number of EBV-derived reads expected to be present in blood WGS
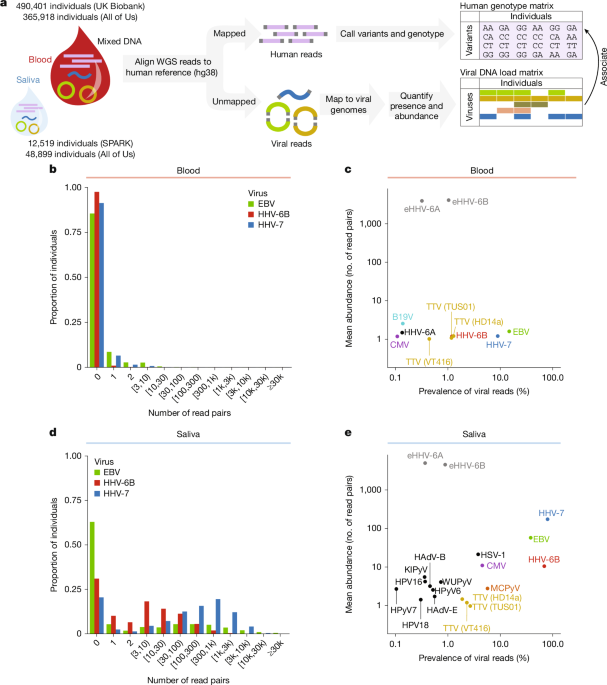
To assess the reasonableness of the distribution of viral read counts observed in blood WGS (typically 0 or 1 read pair per sample, even for near-ubiquitous herpesviruses; Fig. 1b), we roughly estimated the number of EBV-derived reads expected per WGS sample as follows. On average, EBV is present in 1 out of 100,000 B cells4 with roughly 100 episomes per infected cell67. Assuming B cells comprise roughly 5% of all white blood cells76 yields an expected 5 × 10−5 EBV genomes per white blood cell, or 2.5 × 10−5 EBV genomes per haploid human genome in blood-derived WGS. The EBV reference genome is 171,823 bp, so WGS at 30x coverage of the human genome by 2x150bp read pairs should produce an expected 0.4 read pairs from the EBV genome per sample.
Association of viral DNA prevalence with sample collection time
Collection time for blood samples in UKB was obtained from field 3166. For analyses of collection time for saliva-derived WGS in the AoU cohort, we excluded a large fraction of samples (62%) that we determined were likely to have been mailed; for such samples, recorded collection times corresponded to receipt of samples rather than time of saliva sampling. Specifically, samples were excluded if they lacked an in-person physical measurement for heart rate or had a heart rate measurement time separated by more than a day from WGS sample collection time. The recorded collection times for the 18,751 remaining samples were converted from UTC to local time by taking the modal time zone of the state containing the three-digit zip code for the corresponding individual.
P values for hour-of-day and month-of-year associations (Fig. 2e,f and Extended Data Fig. 3k–n) were calculated with ANOVA by comparing models with and without hour of the day or month of the year. Both hour of the day and month of the year were encoded as a series of indicator variables to allow for non-linear relationships with viral prevalence. All models included age, age squared, sex, assessment centre, and top genetic principal components as covariates (20 principal components for UKB; 16 principal components for AoU). Associations were confirmed with Kronos (v.1.0.0), which was run with default settings; to handle covariates, viral phenotypes were first adjusted for covariate effects estimated using harmonic regression.
Association of viral DNA prevalence with genetic ancestry
For UKB, genetically inferred ancestry was determined as described previously77. In brief, 20 genome-wide ancestry principal components were used to identify groups of individuals within a Euclidean distance radius from the centre of individuals within each self-reported ethnicity category, with the distance threshold chosen to include a large fraction of individuals in that self-reported ethnicity category.
For AoU participants, previously generated genetically inferred ancestry24 and ancestry admixture estimates from Rye78 were used.
For SPARK participants, genetic ancestry was inferred using Euclidean distance on 10 genome-wide ancestry principal components to cluster centres determined from the subset of individuals who self-reported race. For each ancestry group, a cluster centre was chosen to have the median principal component coordinate for each principal component among individuals who self-reported a corresponding race. As in UKB, all individuals that fell within a Euclidean distance radius that enclosed a large majority of individuals self-reporting that race were then assigned to that genetically inferred ancestry. For European ancestry, the radius was set to include 90% of individuals who self-reported as ‘white’ (n = 8,157). For African ancestry, the radius was set to include 90% of individuals who self-reported as ‘African American’ (n = 345). For American ancestry, the radius was set to include 75% of individuals who self-reported as ‘Hispanic’ or ‘Native American’ (n = 906). For East Asian ancestry, the radius was set to include 75% of individuals who self-reported as ‘Asian’ and were separated from the majority of samples on PC2 (n = 342). For South Asian ancestry, the radius was set to include 75% of individuals who self-reported as ‘Asian’ and were separated from the majority of samples on PC4 (n = 144).
HLA allele imputation in UK Biobank
The T1DGC reference panel for HLA allele imputation79 (n = 5,225) was first converted to VCF format with variants lifted over to hg19 and merged into multiallelic sites where appropriate. A small number of individuals (n = 136) with >2 alleles for at least one multiallelic site were excluded from the reference panel. Imputation of HLA alleles onto phased SNP-array haplotypes80 was done with BEAGLE81 (v.5.4) using default parameters, after which imputed alleles were converted back to biallelic variants for genetic association analysis and lifted over to hg38.
Genome-wide association analyses of viral DNA load phenotypes in UK Biobank
Abundances of reads aligning to reference genomes for EBV, HHV-6B, HHV-7 and anellovirus strains TUS01, VT416 and HD14a were inverse-normal transformed into quantitative phenotypes for GWAS. Individuals were excluded based on the following criteria: not having European genetic ancestry, not having available TOPMed-imputed genotypes (including for chromosome X), and/or having withdrawn, leaving 453,770 individuals for genetic association analyses (447,190 for HHV-6B after removal of individuals with endogenous HHV-6 integration). TOPMed-imputed variants for these individuals were filtered to require minor allele frequency >0.001 and INFO > 0.3. Linear mixed model association tests were performed with BOLT-LMM82 (v.2.5) to account for relatedness, using the following covariates: age, age squared, sex, genotype array, assessment centre and 20 genetic principal components. SNP array genotypes were used for model fitting, and linkage disequilibrium scores derived from European-ancestry 1KGP samples were used for test statistic calibration. GWAS of germline-inherited endogenous HHV-6A and HHV-6B carrier status were performed using linear regression with the same covariates in individuals of European genetic ancestry, excluding one from each pair of relatives with second-degree or closer relatedness83.
To identify index variants outside the MHC region (Supplementary Table 7), we first iteratively selected the strongest association and removed any variants within 1 Mb. Index variant pairs within 3 Mb were then evaluated to determine whether they represented independent associations using the following approximation of the association strength of the index variant i conditional on the more strongly associated index variant j:
$$\chi _j^2\approx \chi _i^2\left(1-r_ij\mathrmsign(\beta _i\beta _j)\sqrt\frac\chi _j^2\chi _i^2\right)^2$$
as previously described84. The less-strongly associated variant i was dropped if its approximate conditional association was no longer genome-wide significant (P < 5 × 10−8). Identified index variants were annotated with nearby genes using GENCODE 39 (ref. 85) definitions for protein-coding genes, long non-coding RNAs, and microRNAs. Index variants were annotated as expression quantitative trait loci (eQTLs) and splicing quantitative trait loci (sQTLs) using the v.10 release of GTEx86. Follow-up genetic association analyses of variants in the MHC region including both TOPMed-imputed variants and imputed HLA alleles were performed using linear regression with BOLT-LMM on unrelated individuals with European ancestry with the same covariates (Supplementary Table 8).
Partitioning of heritability between MHC and non-MHC variation
To partition heritability between common variants within and outside the MHC region of the human genome, BOLT-REML87 (v.2.5) was run on SNP-array genotypes for unrelated UKB participants with European ancestry with variants within the range chr. 6:24000000 to chr. 6:35000000 assigned to one component (MHC region) and all other variants assigned to a second component, with the –remlNoRefine option set. Age, age squared, sex, assessment centre, genotyping array, and the top 20 genetic ancestry principal components were included as covariates.
Rare variant association analyses in UK Biobank
Gene-level burden masks were generated as previously described88 using genotypes of rare protein-coding variants in the UKB DRAGEN WGS dataset and genotypes of copy number variants previously ascertained from UKB whole-exome sequencing data89. The specific burden masks analysed here for association with viral DNA load used a minor allele frequency threshold of MAF < 0.001 and included missense variants with PrimateAI-3D90 scores >0.7 merged with loss-of-function SNPs, insertion–deletions and copy number variants, using only the MANE Select transcript91. Association tests were performed using linear regression (implemented in BOLT-LMM) on unrelated European-ancestry individuals with age, age squared, sex, assessment centre, genotyping array and the 20 genetic principal components included as covariates. Genes in the MHC region (chr. 6:24000000 to chr. 6:35000000) were excluded given the potential for linkage disequilibrium with strong common-variant associations, leaving 17,589 gene burden masks and a Bonferroni threshold of 5.7 × 10−7 (adjusting for five viruses included in these analyses: EBV, HHV-7 and 3 TTVs).
Genome-wide association analyses of viral DNA load phenotypes in All of Us
GWAS was performed on AoU participants with European ancestry for the following phenotypes: inverse-normal transformed EBV, HHV-6B, HHV-7 and anellovirus strains TUS01, VT416 and HD14a abundances in blood WGS (n = 201,168 for EBV, 198,059 for HHV-6B, and 200,091 for other viruses), EBV, HHV-6B, HHV-7 and MCPyV DNA positivity (that is, presence of any EBV reads) in saliva WGS (n = 33,164 for EBV, 32,711 for HHV-6B, and 33,050 for other viruses), and inverse-normal transformed EBV (n = 16,282) and HHV-7 (n = 29,989) abundances in DNA-positive saliva WGS samples. Variants from the allele count/allele frequency (ACAF) threshold call set present in the TOPMed-r2 imputation panel (to exclude those in regions of poor mappability) were filtered to those with minor allele frequency >0.1% and allele count ≥40 in the subset of European ancestry samples used. BOLT-LMM was run with SNP-array genotypes (minor allele frequency >1%, missingness <10% in European-ancestry samples) as model SNPs, with the following covariates: age, age squared, sex, sequencing site, and 16 genetic principal components. For EBV GWAS, samples without a sex call of XX or XY were assigned indicator variables, whereas they were excluded from GWAS for other viruses due to a minor change in the analytical pipeline during the course of the project.
GWAS was also performed on AoU participants with African ancestry for inverse normal transformed EBV and HHV-7 abundances in blood WGS (n = 77,573). Variants from the ACAF threshold call set with minor allele frequency >0.5% in gnomAD v.4.1 African-ancestry samples were filtered to those with minor allele frequency >0.1% and allele count ≥40 in the subset of African-ancestry samples used. BOLT-LMM (using the –lmmInfOnly flag, as the non-infinitesimal mixed model provided a negligible increase in statistical power) was run with SNP-array genotypes (minor allele frequency >1%, missingness <10% in African-ancestry samples) as model SNPs, with the following covariates: age, age squared, sex, sequencing site and 16 genetic principal components.
Genome-wide association analyses of viral DNA load phenotypes in SPARK
As an auxiliary analysis, we also performed GWAS of viral DNA load phenotypes from SPARK saliva WGS. Abundances of reads aligning to reference genomes for HHV-7, HHV-6B, and Merkel cell polyomavirus were transformed into up to two GWAS phenotypes for each virus: a binary phenotype encoding the presence/absence of any viral reads (HHV-6B, n = 9,081 after sample exclusions; MCPyV, n = 9,209) and a quantitative phenotype comprising inverse normal transformed non-zero values (HHV-6B, n = 6,258; HHV-7, n = 7,360). Individuals with non-European genetic ancestry were excluded using a more permissive ancestry definition based on genomic PC1 and PC2 to maximize power, leaving 9,209 individuals for genetic association analyses. For HHV-6B, carriers of eHHV-6B were also excluded. Variants called by DeepVariant were filtered and used to generate ancestry principal components as previously described83. Linear mixed model association tests were performed using BOLT-LMM to account for relatedness, using the following covariates: age, square root of age, age squared, sex, sequencing batch, percentage of mapped reads and ten genetic principal components.
We observed that GWAS power in SPARK saliva WGS was much lower than in AoU saliva WGS, as expected given the much smaller sample size. We therefore chose not to meta-analyse saliva viral DNA load GWAS results across AoU and SPARK because the potential power gain was modest (given the much smaller size of the SPARK cohort) and might be negated by the age heterogeneity of SPARK (mostly children) versus AoU (only adults).
Meta-analysis of GWAS results from UK Biobank and All of Us
Associations with EBV, HHV-6B, HHV-7 and anellovirus strains TUS01, VT416, and HD14a DNA load in blood in AoU were meta-analysed with those from UKB using METAL92 (v.2020-05-05) in standard error mode (SCHEME STDERR), restricting to variants with minor allele frequency greater than 0.1% (for EBV and HHV-7) or 1% (for HHV-6B and anelloviruses) in both cohorts (ADDFILTER MAF > 0.001 or 0.01), and applying genomic control correction within input studies (GENOMICCONTROL ON). One significantly associated locus in UKB that disagreed in direction of effect between cohorts (and between EBV presence phenotypes generated from left and right halves of the EBV genome; Supplementary Note 5) was filtered.
To compute genetic correlation between viral phenotypes in UKB, AoU, and SPARK, LDSC93 (v.2.0.0) was run with standard settings and pairs of viral summary statistics as input.
Generation of lymphocyte percentage phenotype in All of Us
A lymphocyte percentage phenotype was generated from the ‘Lymphocytes/100 leukocytes in blood by automated count’ phenotype (OMOP Concept Id: 3037511). Only entries with ‘percent’, ‘percent of white blood cells’, ‘percent’ and ‘percentage unit’ as units were kept, discarding entries with other or missing units as well as values outside the range [0,100]. For individuals with multiple valid measurements, we took the median value. This left 170,196 people with lymphocyte percentage values, among whom 24,789 individuals with with African ancestry had blood-derived WGS available and were used to evaluate the extent to which the Duffy-null effects on EBV and HHV-7 DNA load were mediated by effects on lymphocyte percentage.
Measurement of EBV type 1- and type 2-specific alignments
In both UKB and SPARK, unmapped reads and reads that aligned to the chrEBV decoy contig were realigned to the EBV type 1 (NC_007605.1) and EBV type 2 (NC_009334.1) reference genomes using BWA-MEM (providing both reference genomes simultaneously and using 4 computational threads). Read alignments were filtered to those for which both the read and its mate were mapped (samtools view -F 12) and were then collated within 500 bp bins of each of the two reference genomes using mosdepth with the same parameters as above, with the exception that the filter on insert size (-l 100 -u 1000) was dropped, as insert size was undefined in situations in which a read mapped to EBV type 1 and its mate to EBV type 2 or vice versa (for example, if only one read in the pair fell within a type-informative region, and its non-type-informative mate was mapped arbitrarily to type 1 or type 2). A 1 kb offset was added to alignment bin coordinates in the type 1 genome starting at position 85000 to account for differences between the two reference genomes in the EBNA3A–EBNA3C region. Alignments to 500 bp bins within the viral genomic regions 35501–38000 and 79001–92500 (corresponding to EBNA2 and EBNA3A–EBNA3C) were considered to be type-specific, and the numbers of type 1-specific and type 2-specific reads for each sample were computed by summing 500 bp depths across these regions (separately for the two EBV genomes), multiplying by 500/150 = (bin size)/(bases per read), and rounding to the nearest integer. In UKB, samples with at least two type 1-specific reads were designated as positive for EBV type 1, and analogously for type 2.
To evaluate the risk of misclassification between type 1 and type 2, we examined the distribution of type 2 versus type 1 reads in SPARK saliva samples, making use of the fact that saliva samples frequently have hundreds of EBV reads that should typically come from only one EBV type (depending on whether the individual is infected with a type 1 or type 2 EBV strain). This analysis showed that as expected, nearly all samples had read counts heavily skewed to either type 1 or type 2 (typically >99% type 1 or >99% type 2), indicating a low rate of misclassification of reads.
Variation in EBV type 1 versus type 2 frequency by birthplace of UK Biobank participants
For analyses of individuals born in the UK, geographic boundaries for nine regions in England, Wales, Scotland and Northern Ireland were obtained as a GeoJSON file corresponding to ‘NUTS, level 1 (January 2018) Boundaries UK BFC’ (for EBV type analyses) or ‘NUTS, level 2 (January 2018) Boundaries UK BFC’ (for total EBV analyses) from the Open Geography portal from the Office for National Statistics (ONS) (https://geoportal.statistics.gov.uk). Birth coordinates for individuals born in the UK (fields 129 and 130) were assigned to regions with these boundaries using the R package sf (v.1.0-20), and the proportion of participants positive for EBV type 2 (among EBV-positive individuals) was estimated as the ratio of the number of samples determined to be EBV type 2-positive to the total number of samples determined to be either EBV type 1-positive or type 2-positive. For country-level analyses, fields 1647 and 20115 were used.
Genetic association analyses of MHC variants with EBV-type DNA load phenotypes
Variants in the MHC region of the human genome (including imputed HLA alleles) were tested for association with three binary EBV DNA load phenotypes derived from EBV type-specific read alignments. The first two phenotypes coded EBV type 1-positive individuals (respectively, type 2-positive individuals) as cases and all other individuals as controls. The third phenotype, used in case-case association analyses of EBV type 1 versus type 2 positivity, coded individuals who were type 2-positive and lacked type 1-specific alignments as cases (n = 1,366), and those who were type 1-positive and lacked type 2-specific alignments as controls (n = 9,817); a small number of individuals determined to be positive for both type 1 and type 2 were excluded (n = 126). Association tests were performed using linear regression (implemented in BOLT-LMM) on unrelated individuals with European ancestry with age, age squared, sex, genotype array, assessment centre and 20 genetic principal components as covariates.
Association analyses of viral DNA load with biological and clinical phenotypes in UK Biobank
Binarized viral DNA positivity phenotypes were tested for association with binary disease phenotypes in ICD-10 categories derived from UKB ‘first occurrence’ data fields (fields under category 1712, which merged data from electronic health records and self-report) and cancer registry data (field 40006). For each virus, we tested only binary disease phenotypes for which at least 5 cases were expected among viral DNA-positive individuals for that virus (comprising 493–1,413 tests for each of eight common viruses tested: EBV, HHV-6B, HHV-7, three TTVs and eHHV-6A and eHHV-6B). Bonferroni correction was applied to the full set of tests performed across all eight viruses. Association tests were performed using logistic regression with Firth correction using the Wald approximation (pl = FALSE) as implemented in the logistf R package (v.1.26.0) on participants with European ancestry with age, age squared, sex, assessment centre and the 20 genetic principal components as covariates.
Viral DNA positivity phenotypes were also tested for association with quantitative blood phenotypes in UKB: blood counts (category 100081), blood biochemistry (category 17518), and NMR metabolomics (category 220). Association tests were performed using linear regression on individuals with European ancestry with the above covariates, and Bonferroni correction was applied considering all pairwise tests of quantitative blood phenotypes with viral DNA positivity phenotypes.
Binary immunosuppressive drug phenotypes were generated by aggregating synonymous terms from self-reported medication data (category 100075). Specifically, methotrexate was generated from the union of individuals reporting ‘methotrexate’ or ‘mtx – methotrexate’; cyclosporin from ‘cya – cyclosporin’, ‘ciclosporin product’, ‘csa – cyclosporin a’, ‘cya – cyclosporin a’, ‘cyclosporin’, ‘cyclosporin product’ or ‘ciclosporin’; and corticosteroids from ‘prednisone’, ‘prednisolone’, ‘methylprednisolone’, ‘prednisolone product’, ‘dexamethasone’, ‘fludrocortisone’, ‘hydrocortisone’, ‘cortisone product’, ‘hydrocortisone product’ or ‘cortisone’.
Quantitative smoking phenotypes in UKB were generated from the pack-years and cigarettes per day phenotypes by encoding never smokers (and, for cigarettes per day, former smokers) as 0 values. These smoking phenotypes were tested for association with viral DNA positivity phenotypes using linear regression on individuals with European ancestry with the above covariates.
Mendelian randomization to identify causal effects of EBV DNA load on disease
Instrument variables for Mendelian randomization analyses were identified as the 44 lead variants at non-MHC loci from the GWAS meta-analysis of EBV DNA load in UKB and AoU. The MHC region was excluded from Mendelian randomization analyses given the likelihood of linkage disequilibrium generating pleiotropic effects of MHC haplotypes on many immune-related phenotypes. To minimize the impact of sample overlap between cohorts used in the exposure GWAS (EBV DNA load in UKB+AoU blood WGS) and outcome GWAS (disease phenotypes in FinnGen plus UKB plus MVP), we regenerated GWAS summary statistics for EBV DNA load in UKB after restricting to a control-only sub-cohort94. Specifically, we reran BOLT-LMM after removing UKB participants with the following ICD-10 phenotypes: G35, C81, C82, C83, C84, C85, C91, M05, M06 and M32. GWAS results from this control-only UKB sub-cohort were then meta-analysed with AoU associations to generate effect sizes and standard errors for use in Mendelian randomization.
GWAS summary statistics for outcome phenotypes of interest were downloaded from https://mvp-ukbb.finngen.fi/ after applying for access. In these GWAS, FinnGen phenotypes had been harmonized over ICD-8, ICD-9 and ICD-10, cancer-specific ICD-O-3, (NOMESCO) procedure codes, Finnish-specific Social Insurance Institute (KELA) drug reimbursement codes and ATC-codes collected from various registries. MVP phenotypes had been defined by ICD-9 and ICD-10 codes from electronic health records grouped into corresponding phecodes, with case status defined as having two or more phecode-mapped ICD-9 or ICD-10 codes. Meta-analyses had been performed by identifying phenotypes with concordant endpoints, as described at https://finngen.gitbook.io/documentation/methods/meta-analysis. We used the MendelianRandomization R package95 (v.0.10.0) with default settings to generate estimates of causal effect sizes of EBV DNA load on disease phenotypes using the weighted median, inverse-variance weighted (IVW), MR–Egger, and contamination mixture (ConMix) approaches. For weighted median, IVW, and MR–Egger, robust regression with penalized weights was used to account for invalid instrument variables96.
We caution that effect size estimates from Mendelian randomization (here, the increase in log-odds of disease per s.d. increase in EBV reads; Fig. 5e) are expected to be overestimated when the exposure variable (here, EBV DNA load) is measured with high noise. For example, were the exposure phenotype to be randomly permuted in a subset of samples, this would leave the s.d. of the exposure unchanged, but it would shrink down the effect sizes (betas) of the instrumental variables. The betas of instrument variables are used as independent variables in the regression analysis used to estimate Mendelian randomization effect sizes (x axis of Fig. 5f), such that shrinking the betas of the instrument variables would then increase the slope of the Mendelian randomization regression, causing the estimated effect size from Mendelian randomization to increase. This behaviour contrasts with how measurement noise in the exposure variable impacts direct analyses of association with the outcome variable (Fig. 5g): in such analyses, noise reduces the observed association.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.