Sample collection
The first phase of the 1KCP project enrolled 1,379 participants aged 18–76 years (737 men and 642 women) from the Health Examination Center of the Second Affiliated Hospital of Wenzhou Medical University. Peripheral blood samples were collected for library preparation and sequencing. All samples underwent Illumina short-read WGS and RNA-seq. Among these participants, 55 samples were selected for high-coverage PacBio HiFi long-read WGS and Hi-C sequencing, and 1,099 were selected for modest-coverage PacBio long-read WGS. Self-reported ancestry information was not explicitly collected and was therefore not available for this study. Informed consent was obtained from all participants. The study was approved by the Ethics Committee of Westlake University (approval no. 20201127YJ001) and the Ethics Committee of Wenzhou Medical University (approval no. 2019-107).
Short-read WGS
gDNA was extracted from peripheral blood using a gDNA extraction kit and a HF24 nucleic acid purification instrument (Concert). DNA concentration was measured using a Qubit 4.0 fluorometer (Thermo Fisher). A total of 1 µg of DNA was treated with ribonuclease A (Takara) and purified using CleanNGS-R beads (Vdobiotech). Fragmentation, end-repair and A-tailing were performed using 5× WGS fragmentation mix and 10× fragmentation buffer (Enzymatics), followed by ligation with TruSeq index adaptors using T4 DNA ligase and 5× rapid ligation buffer (Enzymatics). Purification, size selection and PCR amplification produced a sequencing library with a target size of 450–550 bp and normalized to 3 ng µl–1. Short-read WGS was performed on an Illumina NovaSeq-6000 platform.
High-coverage long-read WGS
High-quality DNA samples (absorbance of A260/A280 = 1.8–2.0, A260/A230 > 2.0, concentration of ≥100 ng µl–1 and fragment size of ≥40 kb) were purified with Ampure PB beads and fragmented using a Megaruptor2 system (Diagenode). Libraries were prepared using a SMRTbell Prep kit 2.0 with steps similar to modest-coverage sequencing, targeting a library size of 15 kb. Long-read WGS was performed on a PacBio Sequel IIe platform using 8M SMRT Cell and Sequencing plates (v.2.0), with a pre-extension time of 4 h and a video time of 30 h. Primary CCS reads (accuracy >0.88) were generated using CCS (v.6.2.0; https://github.com/PacificBiosciences/ccs) with the –all option. Subreads were aligned to their corresponding primary CCS reads using actc (v.0.60; https://github.com/PacificBiosciences/actc). Finally, HiFi reads (accuracy >0.99) were generated using DeepConsensus (v.1.2.0)60.
Modest-coverage long-read WGS
DNA quality was assessed using a Nanodrop spectrophotometer (Thermo Fisher), a Qubit 4.0 fluorometer (Thermo Fisher) and Mapper XA Chiller system (Bio-Rad). Samples meeting quality criteria (A260/A280 = 1.8–2.0, A260/A230 > 2.0, concentration of ≥70 ng µl–1 and fragment size of ≥40 kb) were purified with Ampure PB beads and fragmented using an Eppendorf centrifuge. Libraries were prepared using a SMRTbell Prep kit 2.0 (PacBio), which involved DNA damage repair, end-repair, adapter ligation and size selection. The libraries were then bound with Sequencing Primer v.3 and Bind Polymerase 2.0, followed by purification with 1.2× AMPure PB beads. Long-read WGS was performed on a PacBio Sequel IIe platform using 8M SMRT Cell and Sequencing plates v.2.0, with a pre-extension time of 4 h and a video time of 30 h. With the –all option, CCS generates one representative read for each zero-mode waveguide (ZMW), either by generating a consensus sequence when sufficient subreads are available or by selecting the median-length subread otherwise (Extended Data Fig. 1a). This process produces two types of read: HiFi reads and non-HiFi reads. Although non-HiFi reads have lower accuracy than HiFi reads, their inclusion resulted in a 2.7-fold increase in total sequencing coverage compared with using HiFi reads alone (Extended Data Fig. 1b).
Hi-C sequencing
Peripheral blood samples were treated with 4 ml of 37% formaldehyde solution (Sigma-Aldrich) for crosslinking, followed by centrifugation to remove the supernatant. The pellets were washed twice with DPBS (Solarbio) and treated with a red blood cell lysis solution (Miltenyi) to remove erythrocytes. After additional washes with DPBS, the samples were resuspended in deionized water. gDNA was released from the suspension using 10% SDS and 10% Triton-X100 (Sigma), followed by digestion with the MboI enzyme (NEB). The resulting DNA fragments were end-repaired using Klenow fragment (Enzymatics) with Biotin-16-dCTP (Trilink) and ligated to adjacent fragments with T4 DNA ligase (Enzymatics). The ligated product was de-crosslinked using proteinase K (Qiagen) and 10% SDS and purified with SPRIselect beads (Beckman Coulter). DNA concentration was quantified using a Qubit 3.0 fluorometer (Thermo Fisher). Approximately 500 ng of purified DNA was subjected to fragmentation, end repair and A-tailing using 5× WGS fragmentation mix and 10× fragmentation buffer (Enzymatics). The fragmented DNA was ligated to Truseq index adaptors and purified using CleanNGS-R beads (Vdobiotech). Biotin-labelled target fragments were captured with M270 magnetic beads (Invitrogen) and amplified using KAPA HiFi HotStart ReadyMix (Kapa Biosystems). The PCR product was purified and normalized to 3 ng µl–1, producing a sequencing library with a target size range of 500–600 bp. Hi-C sequencing was performed on an Illumina NovaSeq-6000 platform.
RNA-seq
Peripheral blood samples were treated with a red blood cell lysis solution (Tiangen) to remove erythrocytes. RNA was then extracted from the leukocyte precipitate using a PAXgene Blood RNA kit (Qiagen). The extracted RNA was assessed using an Agilent Bioanalyzer 2100, and only samples with a total RNA amount >1 µg and a RNA integrity number of >6.5 were used for library preparation. rRNA was depleted from the total RNA using a Ribo-Zero Plus rRNA Depletion kit (Illumina), leaving mRNA and long non-coding RNA intact for downstream processing. The sequencing library was prepared using a Hieff NGS Ultima Dual-mode mRNA Library Prep kit (Yeasen), which involved cDNA synthesis, end repair, A-tailing, adaptor ligation, purification, size selection and PCR amplification. The resulting PCR product was normalized to a concentration of 3 ng µl–1, producing a library with a target insert size of 450 bp. RNA-seq was performed on an Illumina NovaSeq-6000 platform.
PCA
To place the 1KCP samples in a global genetic ancestry context, we performed a PCA using SNV genotypes from the 1KCP samples combined with unrelated samples from the 1000 Genomes Project (1000G) and the Human Genome Diversity Project61. For the 1KCP samples, short-read data were aligned using BWA-MEM (v.0.7.17)62, and SNV genotypes were called using the GATK (v.4.2.6.1) workflow63. Genotypes at overlapping SNV sites across the three datasets were merged to create two analytical sets: (1) a global set with all samples, and (2) an East Asian-only set. For each set, we retained biallelic SNVs with MAF > 0.05, HWE P > 1 × 10–6 and genotype missing rate <10%. The remaining SNVs were further pruned to remove sites in LD using PLINK2 (ref. 64) (–indep-pairwise 50 5 0.5). PCA was then conducted on this pruned dataset using PLINK2.
Diploid genome assembly using the high-coverage dataset
High-coverage HiFi assemblies were generated using hifiasm (v.0.15.3)12, incorporating Hi-C data for haplotype phasing65. To improve mitochondrial genome assembly, we removed contigs aligned to mitochondrial DNA using minimap2 (v.2.24)66 (-cxasm5) and reassembled the mitochondrial genome with Unicycler (v.0.5.0)67.
To clean raw assemblies, PacBio adapter sequences were identified using minimap2 (-cxsr -f5000 -N2000 -secondary=yes) and masked with BEDTools (v.2.30.0)68. Non-human sequences (bacterial, viral, fungal and archaeal sequences) were detected by aligning to the NCBI RefSeq database using blastn (v.2.13.0)69 (-task megablast -word_size 28 -evalue 0.0001 -perc_identity 98.0) and removed.
Diploid genome assembly using the modest-coverage dataset
We used the PIGA workflow19, an integrated framework incorporating multiple tools20,62,63,66,68,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97, to generate diploid genome assemblies using population-scale hybrid sequencing data. A detailed step-by-step description of the PIGA workflow is provided in Supplementary Note 1. In brief, this workflow involves the following steps:
-
1)
Performing population-level SNV calling by integrating long-read and short-read sequencing data.
-
2)
Performing SNV haplotype phasing by leveraging long-read and population information.
-
3)
Generating personalized references by modifying the reference genome with homozygous variants genotyped from the external pangenome and partitioning long reads into haplotypes.
-
4)
Producing draft diploid assemblies based on partitioned long reads.
-
5)
Constructing the pangenome using draft diploid assemblies and simplifying it.
-
6)
Reconstructing the final diploid assemblies by inferring the diploid paths from the constructed pangenome.
Assembly evaluation
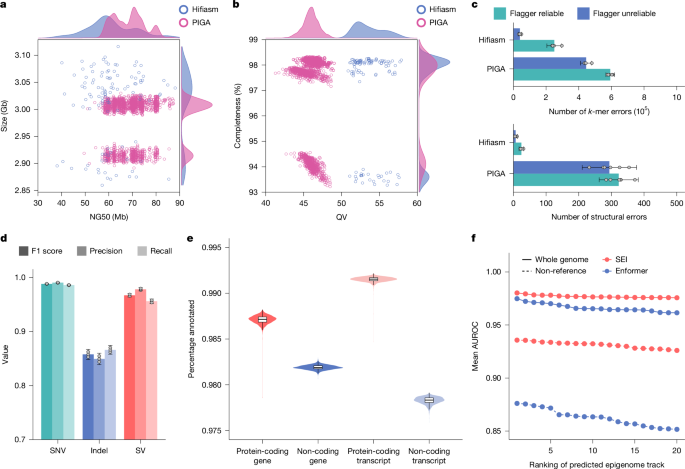
The quality of the 1KCP assemblies was evaluated using multiple approaches. We assessed assembly completeness and QV using Illumina short-read k-mers. A 21-mer database was built using Meryl (v.1.4.1), and completeness and QV of the assemblies were estimated using Merqury (v.1.3)20. Structural errors were detected using high-coverage HiFi reads with Inspector (v.1.0.1)21. Moreover, unreliable assembly regions in both PIGA and hifiasm assemblies of the three benchmark samples were identified using high-coverage HiFi reads with Flagger (v.0.2.0)2. The unreliable regions (including collapsed, duplicated and error regions) in the PIGA assemblies were lifted over to both GRCh38 and CHM13 references, using minimap2 (-cxasm20) for alignment and rustybam (v.0.1.34; https://github.com/mrvollger/rustybam) for coordinate transformation. Reference regions marked as unreliable in more than two haplotypes were defined as common unreliable regions.
Variant calling and haplotype phasing performance of the PIGA assemblies were evaluated using the three benchmark samples. Evaluation callsets were generated from PIGA assemblies with Dipcall (v.0.3)98. Benchmark small variant callsets were constructed by combining variant calls obtained independently using GATK, DeepVariant (v.1.4.0)71 and Dipcall from high-coverage short-read, long-read and hifiasm assemblies. Variant genotypes supported by at least two methods were considered true. Small variant calling performance was evaluated by comparing the evaluation callset to the benchmark callset in Dipcall-confident regions using hap.py (v.0.3.14)99 with the vcfeval engine. SV calling performance was evaluated by treating Dipcall SV calls from hifiasm assemblies as the benchmark set. The evaluation callset was compared with the benchmark set in Dipcall-confident regions using Truvari (v.4.2.2)28 bench (–pick ac –sizemin 50), with harmonization performed using Truvari refine (–recount -u -U). Haplotype phasing performance was evaluated by comparing SNV haplotypes from Dipcall datasets of PIGA and hifiasm assemblies using WhatsHap (v.1.4)73.
Assembly annotation
We developed an integrative assembly annotation pipeline to identify repeats, genes and epigenomic elements in the 1KCP assemblies. For repeat annotation, we used Tandem Repeats Finder (v.4.09)100 (parameters: 2 7 7 80 10 50 15 -l 25 -h) to identify TRs and used RepeatMasker (v.4.1.1)101 with RMBlast to annotate other types of repeat elements, such as LINEs, SINEs and long terminal repeats.
Gene annotation was carried out in multiple steps. First, we lifted the GENCODE (v.40)102 annotation from GRCh38 to individual assemblies with a 95% identity threshold using Liftoff (v.1.6.3)103 (-sc 0.95 -copies -polish). We then aligned RNA-seq reads to the assemblies using HISAT2 (v.2,2.1)104, and performed transcriptome assembly with StringTie (v.2.2.1)105, using Liftoff annotations as guidance. StringTie annotations were filtered to exclude single-exon transcripts or those with transcripts per million (TPM) < 0.5. The filtered StringTie annotations were compared with the Liftoff annotations using gffcompare (v.0.12.9)106 to identify novel transcripts, including intronic, opposite-strand and non-overlapping transcripts. These novel transcripts were further classified as coding or non-coding using CPC2 (v.1.0.1)107. For novel coding transcripts, gene sequences were predicted with GeneMarkS-T (v.5.1)108 and further validated using InterProScan (v.5.66)109 with several databases (for example, CDD, Gene3D, PANTHER, Pfam and PIRSF). Novel non-coding transcripts, specifically long non-coding RNAs, were annotated if their length exceeded 200 bp and had no putative coding sequence longer than 100 bp. Finally, the novel coding and non-coding genes were merged with Liftoff annotations to create the final annotations.
To annotate epigenomic elements, we compared the performance of SEI and Enformer (v.0.8.8) for epigenome prediction in 128-bp steps on both PIGA and hifiasm assemblies from the three benchmark samples. Non-reference sequence regions in each assembly were identified using PAV13 and defined as insertion records of ≥50 bp. ATAC-seq peaks were identified on the assemblies of the matched samples using MASC3 (v.3.0.0)110 with a FDR < 0.01 and served as the ground truth for model evaluation. Predicted regions were labelled as true positive if they overlapped with the ground truth peaks and as false positive otherwise. The AUROC for each predicted track was calculated to evaluate the performance of SEI and Enformer. For large-scale annotation, SEI predictions were performed in 300-bp steps, balancing speed and accuracy (Supplementary Fig. 15). The SEI sequence classes were used to annotate epigenomic elements across all the 1KCP assemblies.
Pangenome construction
Given the computational challenges of integrating such a large dataset, we developed a pangenome construction pipeline that combines features from Minigraph-Cactus111 and PGGB91 while implementing subgraph parallelization to enhance efficiency (Supplementary Fig. 16). We first constructed the pangenome containing only SVs using Minigraph (v.0.21)89, which iteratively integrates assemblies into the pangenome. The order of integration was determined on the basis of the k-mer-based distance of assemblies to the CHM13 reference, estimated using Mash (v.2.3)112. We then realigned the assemblies to the Minigraph pangenome to obtain assembly-to-graph alignments, which were filtered using gaffilter (https://github.com/ComparativeGenomicsToolkit/cactus-gfa-tools) (-r 5.0 -m 0.25 -q 5 -b 250000 -o 0 -i 0.5). To accelerate subsequent steps, we divided the pangenome into approximately 20 Mb subgraphs by removing edges in border regions not part of the bubbles, using the API of odgi (v.0.8.6)113. Alignments and assembly sequences were split accordingly for each subgraph. We then induced the base-level pangenomes for subgraphs using seqwish (v.0.7.4)90 and refined them through three rounds of smoothxg (v.0.7.9)91 with partial order alignment parameters for 0.1% divergence (-p 1,19,39,3,81,1) and target lengths of 1,400, 1,800 and 2,200. The pangenome was then normalized with GFAffix (v.0.1.5; https://github.com/marschall-lab/GFAffix) to collapse shared affixes. Complex regions of the pangenome were filtered by clipping assembly regions masked by dna-brnn (v.0.1; https://github.com/lh3/dna-nn) or unaligned by the Minigraph path, with a size greater than 100 kb.
For quality control, we developed a k-mer-based filtering method to remove non-reference nodes and edges in the pangenome that are unsupported by read k-mers. The procedure included the following steps. (1) For each individual, we used GenomeScope (v.2.0.1)114 to estimate a reliable k-mer coverage range, defining the lower bound as the 0.05th percentile of the single-copy heterozygous coverage distribution and the upper bound as the 99.5th percentile of the single-copy homozygous coverage distribution. (2) A node or edge was considered supported in a haplotype if the proportion of its associated k-mers with coverage between the lower and upper bounds exceeded 20% of all associated k-mers with coverage below the upper bound. (3) Nodes or edges supported by <50% of their carrying haplotypes were filtered out from the pangenome. (4) Graph tips introduced by this filtering were further removed to eliminate residual artefacts.
The integrated pangenome, which encompasses not only the 1KCP assemblies but also publicly available genome assemblies (HPRC and CPC) and reference genomes (GRCh38 and CHM13), was used for quality assessment. For further analysis, we created a 1KCP pangenome that contains only the 1KCP assemblies and reference genomes. To enhance the annotation resolution of the pangenome, segments were divided into lengths of ≤20 bp, as most functional elements exceed this size (Supplementary Fig. 19a).
Path-guided pangenome annotation
We developed a path-guided pangenome annotation method to detect genomic elements in pangenome segments. First, we mapped the pangenome segments to the assembly coordinates guided by the assembly paths. We then compared the overlap between segments and assembly annotations in each assembly. Segments with >80% overlap with specific genomic elements were annotated. The annotation results from all assemblies were integrated into the final pangenome annotation. To ensure robust annotations, we only annotated non-singleton segments and retained annotations supported by >80% of haplotypes (Supplementary Fig. 19b).
Variant detection
We identified reference variant sites by detecting allele traversals in pangenome bubbles using VG (v.1.59.0)78 deconstruct. Variant sites with maximum allele sizes >100 kb or redundant representations were filtered using vcfbub (v.0.1.0; https://github.com/pangenome/vcfbub) (-a 100000 -l 0). The retained variant sites were then classified on the basis of the size difference between the longest and shortest alleles: SNVs (size difference = 0), indels (size difference ≥ 1 and <50) and SVs (size difference ≥ 50).
To simplify the representations of multiallelic reference variant sites, we decomposed them into constituent biallelic variants by clipping shared affixes between reference and alternative alleles using a modified script of PanGenie97. Small variant alleles were identified by directly decomposing the original variant sites. For detecting SV alleles, similar alleles (≥70% size and sequence similarity) from original SV sites were first merged using Truvari collapse (-k common -r 0 -p 0.7 -P 0.7) and then decomposed.
To identify nested variant sites, we discovered additional bubbles in a reference-free manner using VG snarls24. Variant sites not traversed by the reference allele were classified as nested variant sites. The alleles of nested variants were further resolved by extracting substrings of allele traversals from reference variant sites based on the flanking segments.
We characterized TR sites by analysing their motif sequences using vamos (v.2.1.5)29 with the v.2.1 e0.1 motif set and the STRchive35 disease-associated TR set. Single-base motifs were excluded, as they are commonly associated with long-read sequencing errors. The individual-level TR calls were then integrated into the population-level callset using tryvamos combineVCF.
For quality control, we filtered out variants with >20% of their length overlapping common unreliable regions, treating the positions of nested variants as those of their parent variant sites. The quality control steps reduced the error rate of our callset (Supplementary Fig. 22). For TRs, we also removed loci for which inferred sequences exhibited an average identity of <80% to the corresponding assembled sequences across haplotypes.
Variant assessment
We used a Ti/Tv ratio-based approach to estimate the FDR of the SNV callset. Using the gnomAD (v.4.1) database as a reference, SNVs were classified as known (present in gnomAD) or novel (absent from gnomAD). Let x denote the Ti/Tv ratio of known SNVs, representing the expected biological signal, and y denote the Ti/Tv ratio of novel SNVs, thereby reflecting a mixture of true rare variants and random errors (assuming a random error Ti/Tv ratio of 0.5). The FDR among novel SNVs was calculated as FDRnovel = 3(x – y)/[(2x − 1)(y + 1)]. In the 1KCP callset, we identified 25.4 million known SNVs (x = 2.27) and 9.1 million novel SNVs (y = 1.34). Substituting these values produced an estimated FDR of 33.7% for novel SNVs. The overall genome-wide FDR was then calculated by weighting the novel FDR by the proportion of novel SNVs in the total SNV set, which resulted in an estimated overall FDR of 8.9%.
For evaluating the SV callset, HWE P values were calculated for SV alleles using the fill-tags plugin in BCFtools (v.1.20)70. SVs with HWE P > 1 × 10–6 were considered to be in HWE. The FDR of the SV callset was estimated based on SV calls derived from the pangenome constructed using both PIGA and hifiasm assemblies of the three benchmark samples, together with the GRCh38 reference and other 1KCP assemblies. A given SV in the PIGA assemblies was considered a true positive if validated by at least one of the following: (1) a direct match to the corresponding hifiasm assembly; (2) a match identified using Truvari bench and refine; or (3) read-based validation using high-coverage HiFi data aligned with GraphAligner (v.1.0.20)77 and genotyped using VG call79. The SV callset was also compared against other SV callsets, including public callsets, as well as the 1KCP long-read Sniffles2 callset (jointly called using Sniffles2 (v.2.6.2)86) and the 1KCP short-read callset (generated with Manta (v.1.6.0)115 and merged using SURVIVOR (v.1.0.6)116) (Supplementary Table 15), using Jasmine (v.1.1.5)87 (–allow_intrasample).
For evaluating the TR callset, we also called TRs from short-read data using GangSTR (v.2.5)117 and from long-read data using vamos. The PIGA and long-read TR callsets from the three benchmark samples were compared against the hifiasm TR callset by using edlib118 to calculate edit distances between repeat unit representations of paired TR alleles at corresponding TR sites. Alleles from different callsets were paired to minimize the total edit distance across all possible allele pairs. Furthermore, TR alleles from these callsets were normalized using vcfwave (v.1.0.14)119 and benchmarked using Truvari bench (-s 5) and refine (-u) against the hifiasm Dipcall callset, which was restricted to GIAB-defined TR regions120.
Variant annotation
For SV sites, the longest allele was considered the representative allele, and its sequence properties were characterized on the basis of the pangenome annotations. The proportion of each repeat type was calculated according to the cumulative size of segments annotated with that repeat. SV sites with more than 50% of their length annotated as a specific repeat were classified as specific repeats. SV sites with more than 50% annotated as multiple repeat types, but without a dominant repeat annotation, were classified as mixed repeats. SV sites with less than 50% annotated as repeats were labelled as non-repeats.
For nested variant sites, annotation was performed on the basis of their context in the pangenome. If a flanking segment was part of a specific genomic element, the nested variant site was considered to be located in that element.
Definition of MRG sets
Given that different medical genetic databases curate gene lists with varying scopes and purposes, we defined MRGs specifically for each downstream analysis. For gene-altering SV analysis, MRGs were defined as genes catalogued in the OMIM31 and COSMIC32 databases. For TR expansion analysis, MRGs were defined as known disease-associated TR genes catalogued in the STRchive35 database. For HLA allele analysis, MRGs were defined as genes catalogued in the IPD-IMGT/HLA121 database.
TR expansion detection
As TR expansion events can be regarded as outliers in the TR size distribution in the population, we used the non-parametric outlier detection algorithm DBSCAN to identify TR expansions. Following the settings of a previous study38, we used the log2 of the number of haplotypes as the minimum number of points (minPts) and the mode size of a TR site as the maximum distance (ε). TR alleles with sizes exceeding the longest clustered TR alleles were identified as TR expansion events.
Methylation rate estimation
To analyse the methylation status of expanded TR alleles, HiFi reads from samples with expanded alleles were mapped to the GRCh38 reference genome. Methylation status was estimated using MeLoDe122, and haplotypes of HiFi reads were assigned with WhatsHap. Methylation rates for each CpG site around the TR sites were separately calculated for each haplotype.
Gene cluster variation detection
To identify gene cluster variations, we aligned the protein sequences of GENCODE canonical isoforms to the assemblies using miniport (v.0.12)123 (–outs=0.97 –no-cs -Iu). Next, a gene graph was constructed using pangene (v.1.1)41, and gene cluster variations were detected by identifying bubbles in the graph using the pangene.js call.
GO enrichment analysis
GO enrichment analysis was performed using the clusterProfiler (v.4.12.6)124 on a given gene list. Redundant enriched GO terms were filtered using the GOSemSim (v.2.30.2)125 with a similarity cut-off value of 0.7.
Pangenome visualization of complex loci
For the visualization of complex loci, subgraphs of regions of interest were generated based on the GRCh38 coordinates using odgi extract (-d 50000). These subgraphs were visualized using Bandage (v.0.9.0)126, whereby segments were coloured according to pangenome annotations.
Comparison with the GWAS catalogue
Trait associations from the GWAS catalogue127, downloaded on 15 October 2024, were linked to 258,641 unique variants. SV alleles in strong LD (r2 > 0.8) with these trait-associated variants were identified using PLINK2 (v.2.0.0) for LD calculation.
HLA gene analysis
Four-field alleles of HLA genes were called from assemblies using HiFiHLA (v.0.3.1; https://github.com/PacificBiosciences/hifihla) with the ‘call-contigs’ option, and one-field to three-field alleles were derived from the four-field alleles. The 39 typed HLA genes were categorized into 8 classical HLA genes (HLA-A, HLA-B, HLA-C, HLA-DRB1, HLA-DQA1, HLA-DQB1, HLA-DPA1 and HLA-DPB1) and 31 non-classical HLA genes.
For quality control, we applied a filter to retain only allele typing results with >99% sequence identity and >95% alignment coverage to their best-matching reference alleles. We further excluded three HLA genes (HLA-U, HLA-K and HLA-W) in which >20% of alleles did not pass these criteria. We also performed independent HLA typing from short reads using HLA-HD (v.1.7.1)48 to validate the HiFiHLA typing results. HLA genes with concordance <95% were excluded for LD analysis.
To assess the haplotype structure of these HLA genes, we used eLD (v.1.0)49 to calculate the entropy-based LD measurement index (ε), which reflects the multiallelic LD relationships between the HLA genes. The analysis was performed separately using three-field and four-field HLA alleles with AF > 0.05.
eQTL analysis
To quantify gene expression levels, RNA-seq reads were aligned to GRCh38 using STAR (v.2.7.9)128 with GENCODE (v.40) annotation. Gene read counts and TPM values were generated with RNA-SeQC (v.2.4.2)129, and samples with fewer than 10 million reads were excluded. Gene expression outlier samples were identified by calculating the average correlation of each sample with all others and labelling those with notably lower correlations130 (median absolute deviations of <0.85). Genes with >0.1 TPM and ≥6 reads in at least 20% of samples were retained. Expression levels were TMM-normalized using edgeR (v.3.22.5)131 and inverse-normal transformed. To account for the hidden factors influencing expression variation, probabilistic estimation of expression residuals132 (PEER) was calculated. Following the GTEx Consortium guidelines50, the number of PEER factors was selected based on sample size. A total of 60 PEER factors, along with sample age and sex, were used to adjust gene expression levels.
Non-TR variants with MAF > 0.01 and TRs with heterozygosity >0.98, both having a HWE P > 1 × 10–6, were retained. We included 8.03 million SNVs, 4.17 million indels, 56,242 SVs, 0.89 million nested variants with MAF > 1% and 0.46 million multiallelic TR variants (0.15 million TR length variants and 0.31 million TR motif variants) for the subsequent analysis. After excluding samples with genetic relatedness >0.05, 1,101 samples were included in the following analysis. Cis-eQTL mapping (within 1 Mb upstream and downstream of the TSS) was performed using tensorQTL (v.1.0.9)133, with 20 genotype principal components (PCs) and assembly methods included as covariates. Significant eGenes were identified at a 5% FDR using adaptive permutation mode with 10,000 permutations134. Nominal P values were then computed to detect significant gene–variant associations. Fine-mapping analysis was performed using the SuSiE algorithm implemented in tensorQTL to identify credible sets of eQTL signals. Stepwise conditional regression, implemented in tensorQTL, was then used to identify independent eQTL signals for each eGene.
Heritability estimation
We used GCTA (v.1.94.1)53,54,55 GREML to estimate the cis-heritability of gene expression, partitioned by variant type. Genetic relationship matrices (GRMs) for non-TR variant types (SNV, indel, SV and nested variant) were constructed using PLINK2. A GRM for TRs was constructed using dgrm (https://github.com/PeixiongYuan/dgrm). These five GRMs were fitted into a GREML multicomponent model, with the 20 genotype PCs and assembly methods as covariates. The parameter –reml-no-constrain was used to obtain unbiased heritability estimates.
Enrichment of eVariants
To assess the enrichment of eVariants with PIP > 0.9 across variant types, we performed 100,000 resampling iterations. For each resampling, an equal number of variants with matching distance to TSS as the eVariants with PIP > 0.9 were randomly selected to generate the null distribution. Fold-enrichment and empirical P values were then calculated for each genomic element based on the resampling results.
To assess the enrichment of eNested variants in genomic elements, we performed 100,000 resampling iterations. For each resampling, an equal number of nested variants with matching MAF and distance to TSS as the eNested variants were randomly selected to generate the null distribution. Fold-enrichment and empirical P values were then calculated for each genomic element based on the resampling results.
Colocalization analysis
We leveraged eQTLs of 15,699 significant eGenes in the 1KCP dataset and GWAS data for 215 traits from BioBank Japan to perform the eQTL–GWAS colocalization analysis. Genetic variants common to both eQTL and GWAS datasets were selected for further analysis. Colocalization analyses were conducted using the coloc (v.5.2.3)57 (in ABF mode) and SMR (v.1.3.1)56 with HEIDI test. eQTL–GWAS signal pairs were considered colocalized if they met either of the following conditions: (1) coloc PP4 > 0.9 or (2) PSMR < 3.18 × 10–6 and PHEIDI > 0.05.
Imputation panel construction and assessment
We constructed the 1KCP imputation reference panel based on the GRCh38 linear reference, incorporating a comprehensive range of variant types in biallelic form, including SNVs, indels, SVs, nested variants, TR variants and HLA alleles. For HLA alleles, the typing results were converted into variant representations, with the variant positions defined at the centre of the corresponding HLA genes. Variants with a missing rate >10%, HWE P < 1 × 10–6 or allele count <2 were excluded from the reference panel.
To evaluate imputation performance, we performed leave-one-out imputation using Minimac4 (v.4.1.6)135, whereby each of the 55 high-coverage samples was sequentially excluded as the imputation target sample. The SNV sites used for imputation were restricted to those accessible by the Infinium Omni2.5 array or short-read WGS. For non-TR variants, the imputation accuracy was assessed by calculating the squared correlation between the true genotypes and the imputed dosages across individuals. For TRs, the imputed length and motif copy numbers at multiallelic loci were derived as dosage-weighted sums. Specifically, the length copy number was calculated as the sum of genotype dosage multiplied by the repeat unit count for each allele, whereas the motif copy number was calculated as the sum of genotype dosage multiplied by the count of specific motifs for each allele. The squared correlation was calculated by comparing the true and imputed TR copy numbers across individuals.
For comparison with the existing panels, we used 70 East Asian samples from CPC, HPRC and HGSVC3 to construct the truth callsets and then performed imputation benchmarking. The SV callset was generated using PAV and further merged using Jasmine. The HLA callset was generated using HiFiHLA. As the 1000G-ONT SV panel contains only SNV sites restricted to the UKB array, imputation for SV comparison was performed using the UKB pseudo-array genotypes as input. For TR and HLA comparisons, we used the Omni 2.5 pseudo-array genotypes as input. Imputations using the 1000G-ONT SV panel and the EnsembleTR panel were conducted with Minimac4. Imputation with the four-digit multiethnic HLA v.2 panels was performed on the Michigan online imputation server135. For SV benchmarking, we used Truvari bench (–pick ac -p 0.7 -P 0.7 -s 50) to match SVs across different callsets. For TR benchmarking, we assessed the performance of imputed TR alleles using Truvari bench (-s 5) and refine (-u) against the PAV callset, restricted to GIAB-defined TR regions.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.