ScFks1 expression and purification
ScFks1 was expressed and purified from S. cerevisiae BY4741 fks2 deletion strain18 harbouring the genomic copy of the fks1 gene engineered to have a C-terminal HRV3C protease site, ten amino acid linker, 3× Flag tag, and 10× His-tag. The fks1 gene was modified by homologous recombination by following a standard procedure42. To express ScFks1, yeast cells were grown in YPD medium and then collected by centrifugation. The resulting cells were used to prepare membrane fractions as described previously18. To purify ScFks1, the membrane fractions were passed through a French Press cell twice at 18,000 psi. Then, buffer A (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM EDTA, 10% glycerol) with 1.5% N-dodecyl β-d-maltoside (DDM) and 0.15% CHS (w/v) was added slowly while stirring then stirred overnight at 4 °C. The resulting mixture was clarified by centrifugation at 15,000g for 30 min at 4 °C, and the supernatant was then treated with anti-Flag affinity resin (Pierce). The slurry was mixed for 1 h, and the resin was then packed in a column and washed with 12.5 column volumes of buffer A with 1.5% DDM and 0.15% CHS (w/v). The resin was then washed with another 12.5 column volumes of buffer A with 0.05% LMNG and 0.005% CHS (w/v) or 0.1% CHAPS and 0.01% CHS (w/v) buffer. Bound proteins were eluted by washing with buffer containing 200 µg ml−1 Flag peptide. Eluates were concentrated to <0.5 ml, and purified with size-exclusion chromatography (Superose 6 Increase 10/300 GL column, GE Healthcare) equilibrated in buffer B (20 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM EDTA) with 0.005% LMNG and 0.0005% CHS (w/v) or 0.1% CHAPS and 0.01% CHS (w/v). The peak fractions were pooled and concentrated for cryo-EM sample preparation. For the structural and functional characterization of S643P-Fks1, the S643P mutation was introduced to the genomic copy of fks1 described above using the CRISPR–Cas9 genome editing using a reported protocol43. The mutant was expressed and purified following the same protocol as for wild-type ScFks1.
ScRho1 expression and purification
S. cerevisiae Rho1 (ScRho1) with N-terminal 6×His-tag and tobacco etch virus protease site was expressed in Escherichia coli Rosetta 2 (DE3) cells (Novagen) harbouring the pET28a plasmid with the rho1 gene codon optimized for expression in E. coli. The E. coli cells were grown in Luria Bertani medium with chloramphenicol (25 mg l−1) and kanamycin (50 mg l−1) at 37 °C to an optical density at 600 nm (OD600) of around 1.0. Protein expression was induced by adding 0.5 mM isopropyl β-d-1-thiogalactopyranoside, and the cultures were continued for 20 h at 15 °C. Cells were then collected by centrifugation and resuspended in buffer C (50 mM Tris, pH 7.5, 200 mM NaCl, 10% glycerol) containing 2 mM β-mercaptoethanol (BME) and lysed by passing them through a French Press cell operating at 18,000 psi. The lysate was then clarified by centrifugation at 25,000g for 25 min at 4 °C. The supernatant was then applied to a column packed with cobalt-iminodiacetate resin (Prometheus), the column was washed with 12.5 column volumes of buffer C with 5 mM imidazole, and Rho1 was eluted with 5 column volumes of buffer C with 250 mM imidazole. After adding 2 mM BME, the eluate was concentrated and buffer-exchanged using PD10 columns (GE Healthcare) equilibrated with 50 mM Tris, 200 mM NaCl, 2 mM BME and 30% glycerol. Protein concentrations were determined using the Bradford method, with bovine serum albumin as a concentration standard. For cryo-EM sample preparation, Rho1 was further purified by size-exclusion chromatography (Superdex 200 Increase, Cytiva) equilibrated in 20 mM Tris, 150 mM NaCl, 2 mM BME. The peak fractions were pooled and concentrated for cryo-EM sample preparation.
ScFks1 activity assays
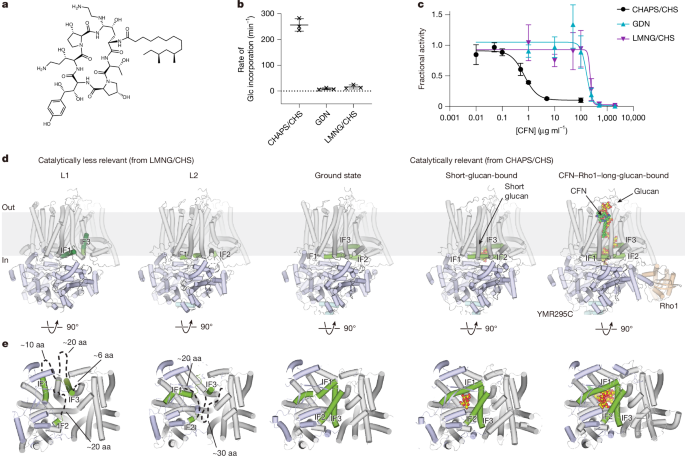
In vitro radioactive GS assays were performed as reported previously18. To test the effects of detergents on GS activity, ScFks1 was mixed with four times the final reaction concentration of detergents. The mixture was then mixed with an equal volume of 125 µM ScRho1 and incubated on ice for 1 h. Reactions were initiated by mixing with equal volumes of the ScFks1–Rho1 solution and a UDP-Glc (4 mM) solution as described previously18.
Cryo-EM sample preparation
All cryo-EM samples in this study were prepared on freshly glow-discharged UltrAuFoil R1.2/1.3 300 mesh grids (Quantifoil), using a Leica EM GP2 plunge freezer with the chamber set at 30 °C and 85% humidity. For the ScFks1 sample in LMNG/CHS, peak fractions from the size-exclusion chromatography in LMNG/CHS were concentrated to approximately 10 mg ml−1 and doped with 0.1% CHAPS/0.02% CHS. Then, 5 mg ml−1 CFN dissolved in water with 10% dimethylsulfoxide was mixed with ScFks1 at 1:20 (v/v) and incubated for 2 min. A 3-μl sample of ScFks1 was applied to the grid, incubated in the chamber for around 60 s then blotted for 1.5–2 s followed by plunge-freezing in liquid ethane cooled by liquid nitrogen. For the ScFks1 sample in CHAPS/CHS, peak fractions from the size-exclusion chromatography in CHAPS/CHS were concentrated to around 12–15 mg ml−1. Purified Rho1 was mixed with ScFks1 at a 1.5:1 molar ratio, and GTPγS was added at a 1.5-fold molar concentration of Rho1. 1.5 mM EDTA, 50 mM KF and 7.5% glycerol were further included in the reaction mixture. The sample was incubated with 12.5 mM UDP-Glc and 0.5 mg ml−1 CFN on ice for 30 min. During grid freezing, 3 μl of ScFks1 was applied to the grid, incubated in the chamber for approximately 60 s and blotted for 1.5–2 s followed by plunge-freezing in liquid ethane cooled by liquid nitrogen. The S643P sample was prepared similarly, with a longer reaction time (2 min).
Cryo-EM data collection
The LMNG/CHS doped with CHAPS/CHS sample was collected with a Titan Krios microscope (Thermo Fisher) operating at 300 kV and equipped with a K3 detector (Gatan) and a GIF energy filter with slit width of 20 eV in super-resolution mode using SerialEM (v.4.2)44. Videos were collected at a nominal magnification of ×105,000 with a super-resolution pixel size of 0.4135 Å per pixel at the specimen level. Each video contained 50 frames over a 2.9-s exposure time, using a dose rate of around 17.2 e− pixel−1 s−1, resulting in the total accumulated dose of approximately 50 e− Å−2. The nominal defocus range was set from −0.8 to −2.2 μm.
The wild-type CHAPS/CHS sample was collected with a Titan Krios microscope (Thermo Fisher) operating at 300 kV and equipped with a K3 detector (Gatan) in counting mode behind a BioQuantum GIF energy filter with a slit width of 20 eV, using the Latitude-S (v.3.3) automated data-acquisition program. Videos were collected at a nominal magnification of ×105,000 with a pixel size of 0.840 Å per pixel at the specimen level. Each video contained 40 frames over a 2.4-s exposure time, using a dose rate of around 25 e− pixel−1 s−1, resulting in the total accumulated dose of around 60 e− Å−2. The nominal defocus range was set from −0.8 to −2.2 μm.
The S643P sample was collected with a Titan Krios microscope (Thermo Fisher) operating at 300 kV and equipped with a K3 detector (Gatan) in counting mode behind a BioQuantum GIF energy filter with slit width of 20 eV, using the EPU (v.3.8.1) program. Videos were collected at a nominal magnification of ×105,000 with a pixel size of 0.820 Å per pixel at the specimen level. Each video contained 50 frames over a 2.7-s exposure time, using a dose rate of approximately 18 e− pixel−1 s−1, resulting in the total accumulated dose of around 50 e− Å−2. The nominal defocus range was set from −0.8 to −2.2 μm.
Cryo-EM data processing
All datasets were processed following similar procedures. Beam-induced motion correction and dose-weighting were performed using Relion’s own implementation45. The motion-corrected micrographs were then imported into cryoSPARC (v.4.4)46 for contrast transfer function (CTF) estimation. Micrographs were selected subsequently based on CTF fit quality and CTF estimated resolution. An initial set of approximately 5,000 particles was picked manually and subjected to a reference-free two-dimensional classification, from which the classes with clear protein features were selected as references for template-based particle picking. Picked particles were extracted with a 320-pixel box size at bin4. Iterative two-dimensional classification was performed in cryoSPARC, and classes showing clear secondary structure features of ScFks1 were selected, re-extracted at bin2 for ab initio reconstruction with two classes. For the CHAPS/CHS samples, particles from the good two-dimensional class were used to train a TOPAZ (v.0.2.5)47 model, which was applied to pick through all the micrographs again. The particles from TOPAZ extract were subject to two-dimensional classification separately from the particles from template picking. Good particles from template picking and topaz picking were combined and duplicate particles removed before heterogeneous refinement.
Iterative heterogeneous refinement was performed with the selected reference from parallel ab initio reconstruction—one good and one junk reference—until the estimation resolution for the good class is better than 4.8 Å for wild type and 6.4 Å for S643P. Particles from the good class that shows clear protein density were re-extracted without binning for non-uniform refinement48. For wild-type samples, after non-uniform refinement, the particles from the consensus reconstruction were transferred to RELION 4.0 (ref. 45).
For the LMNG/CHS doped with CHAPS/CHS data, skip-align three-dimensional classification yielded classes corresponding to the L1 and L2 states. To further improve resolution, signal subtraction was performed on the L1 and L2 states separately to remove micelles, followed by two to three more rounds of skip-align three-dimensional classification. After each round, all classes were refined to evaluate resolution and conformational heterogeneity. The good classes were kept for the next round. During these rounds, only one class from the L1 or L2 state remained at high resolution, suggesting little conformational heterogeneity within each state. Finally, the best particles from each state were reverted to full particles and subject to Bayesian polishing. The polished particles were imported back to cryoSPARC and finalized with CTF refinement and non-uniform refinement.
For the wild-type CHAPS/CHS data, skip-align three-dimensional classification only yielded two good classes. One class had clear CFN density but weak Rho1 density. The other class showed no CFN or Rho1 density, corresponding to the ground state. Further subtraction of the micelle signal and the subsequent skip-align classification did not improve the map quality of either class. These two classes were subject directly to Bayesian polishing in RELION v.4.0 (ref. 45). The polished particles were transferred back to cryoSPARC for CTF refinement. After CTF refinement, the ground state class was subject to more iterations of heterogeneous refinement to improve resolution. The CFN-containing class was first subject to three-dimensional classification in cryoSPARC with a mask excluding the micelle signal, which yielded two classes containing similar numbers of particles. One class showed clear Rho1 and CFN density and corresponds to the CFN–Rho1–long-glucan-bound state. The other class did not have CFN or Rho1 density but had signal for three sugar units in the channel. This class corresponds to the short-glucan-bound state. Both classes were further cleaned up by heterogeneous refinement and finalized with non-uniform refinement.
For the S643P sample, one round of three-dimensional classification was performed in cryoSPARC first to further clean up the particle set. Only particles from the class that converges at 3.68 Å were transferred to RELION v.5.0. Signal subtraction of the micelle and the subsequent skip-align classification yielded only one good class. The particles were reverted to full particles and subject to Bayesian polishing. The polished particles were imported back to cryoSPARC and finalized with CTF refinement and non-uniform refinement.
Model building and refinement
The ScFks1 structure at the L2 state was built de novo in Coot (v.0.98)49. During model building, residue registration was guided by the presence of large aromatic side chains. All other wild-type ScFks1 models used the L2 structure as the template. The S643P model used the L1 state as the template. In the CFN–Rho1–long-glucan-bound state, the ScRho1 model from PDB 3A58 was first docked into the Rho1 density. For YMR295C, a polyalanine model was first built de novo into the density in the CFN–Rho1–long-glucan-bound state. After we identified this segment as YMR295C, side chains were registered based on the AlphaFold50 model and the presence of large aromatic side chains. The entire structure was then inspected manually and refined in Coot with ideal geometry restraints. The YMR295C model was used as the template for the YMR295C density in other states. The restraints for lipids and ligands, including PE, UDP, CFN and GTPγS were generated in the Grade2 server51 and optimized in eLBOW (as implemented in Phenix v.1.20 (ref. 52)) using the REEL QM2 method if necessary. Ligand restraints were inspected and adjusted manually to ensure correct stereochemistry before being fitted into the cryo-EM maps in Coot. The glucan polymer was modeled from the reducing end towards the non-reducing end with an ascending numeric order of Glc units (1–17), whereas all the subscripted Glc numbers (0–16) in the main text refer to Glc positions from the non-reducing end towards the reducing end based on distance from the active site. The MolProbity(v.4.4)53 server was used to identify problematic regions in the models, which were then adjusted manually in Coot. The final refinement was performed using the phenix-real_space_refine function with global minimization and secondary structure restraints as implemented in the Phenix suite52. The Fourier shell correlation of the half- and full-maps against the model, calculated in Phenix, was in good agreement, indicating that the models were not over-refined. Structural analyses and illustrations were performed using PyMOL (v.3.1) (Schrödinger), University of California San Francisco (UCSF) Chimera (v.1.7) and ChimeraX (v.1.10)54. Structure alignments and cryo-EM density map alignments were performed by Fit-In-Map in UCSF Chimera.
Estimation of the ScFks1–ScRho1 affinity by ScRho1 titration assay
CHAPS/CHS purified ScFks1 were mixed with four times the desired concentration of purified ScRho1 and a 1.1× concentration of GTPγS. This mixture was incubated for 1 h on ice, and reactions were performed as above. Additional GTPγS (20 µM) was added in the reaction mix in case low GTPγS concentrations affected activity. To determine the EC50 value of ScFks1 activation by ScRho1, the observed GS activity was plotted against ScRho1 concentration and fit to the four-parameter logistic equation (GraphPad Prism v.8.0).
Yeast spot assay
ScFks1 mutants were prepared by site-directed mutagenesis using p423 GAL1 plasmid vector harbouring ScFks1-HRV3c-3xFlag-10×His gene as template. The resulting plasmids were introduced into S. cerevisiae BY4741 ΔFks1 (Horizon Discovery) using the lithium acetate/ssDNA carrier/polyethylene glycol transformation protocol42. Transformed cells were plated on Synthetic Defined media lacking Histidine (SD-His), and single colonies were re-streaked. Colonies were grown overnight in SD-his with 2% raffinose, and the OD600 of each culture was adjusted to 2.0. This adjusted culture was serially diluted six times, and 5 µl of each serial dilution was plated on SD-His +2% raffinose plates with or without 1 µg ml−1 FK506 and 0.24 or 0.5 µg ml−1 CFN. Plates were incubated at 30 °C, and imaged after 48, 72 and 96 h using an Azure 400 Imager (Azure Biosystems).
Western blot analysis
To verify expression, selected mutants were grown in SD-His +2% raffinose at 30 °C and harvested at an OD600 of around 3–7. Cells were lyophilized and lysed with beads-beating, after which membrane fractions were prepared as described previously18. The membrane was solubilized with 1.5% DDM/0.15% CHS overnight in 50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM EDTA, 10% glycerol. Total protein content of the clarified lysates was determined using A280. Lysates containing approximately 15 mg total protein was incubated with 30 μl Flag resin for 2 h at 4 °C. The resin was washed three times with 50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM EDTA, 10% glycerol, 10 mM DDM/1 mM CHS and resuspended to 125 μl, 25 μl of which was loaded onto SDS–PAGE. Following SDS–PAGE (Bio-Rad), protein was transferred onto 0.45-μm polyvinylidene difluoride membranes, blocked with 5% bovine serum albumin in TBS and probed with 1,000× diluted monoclonal mouse anti-Flag M2 antibody (Sigma Aldrich), then 5,000× polyclonal IRDye 800CW donkey anti-mouse antibody (LICORbio) and detected using the Odyssey M Imaging System (LICORbio).
Liquid chromatography tandem mass spectrometry proteomics sample processing, data collection and data analysis
Samples were brought to 5% SDS, reduced for 15 min at 80 °C with 10 mM dithiothreitol and alkylated with 20 mM iodoacetamide for 30 min at room temperature. Samples were then supplemented with a final concentration of 1.2% phosphoric acid and 375 µl of S-Trap (Protifi) binding buffer (90% MeOH/100 mM TEAB). Proteins were trapped on the S-Trap micro cartridge, and digested using 20 ng µl−1 sequencing grade trypsin (Promega) for 1 h at 47 °C. The cartridge was washed using 50 mM TEAB, followed by water with 0.2% formic acid, and the peptides were eluted using 50% acetonitrile in water with 0.2% formic acid. The resulting samples were then lyophilized to dryness and resuspended in 20 μl of 1% trifluoroacetic acid and 2% acetonitrile in water with 12.5 fmol µl−1 of yeast alcohol dehydrogenase as an internal standard.
Liquid chromatography with tandem mass spectrometry (MS/MS) was performed using an EvoSep One ultra performance liquid chromatography coupled to a Thermo Orbitrap Astral high-resolution tandem mass spectrometer (Thermo). In brief, 2 μl of each sample was loaded onto an EvoTip and the peptides were eluted onto a 1.5-µm EvoSep 150um ID × 15 cm performance (EvoSep) column using the SPD30 gradient at 55 °C. Data collection on the Orbitrap Astral mass spectrometer was performed in a data-independent acquisition (DIA) mode of acquisition with r = 240,000 (at m/z 200) full mass spectrometry scan from m/z 380–980 in the Orbitrap with a target automatic gain control (AGC) value of 4 × 105 ions. Fixed DIA windows of 4 m/z from m/z 380 to 980 DIA MS/MS scans were acquired in the Astral with a target AGC value of 5 × 104 and maximum fill time of 6 ms. A high-energy collision dissociation energy setting of 27% was used for all MS/MS scans. Data were imported into Spectronaut v.19 (Biognosis). Relative peptide abundance was measured based on MS/MS fragment ions of selected ion chromatograms of the retention time-aligned runs. The MS/MS data were searched against a SwissProt Yeast database (downloaded in March 2025 from https://www.uniprot.org/uniprotkb?query = %28taxonomy_id%3A559292%29), a common contaminant/spiked protein database (bovine albumin, bovine casein, yeast alcohol dehydrogenase, and so on), and an equal number of reversed-sequence ‘decoys’ for false discovery rate determination. A Direct DIA+ library-free approach within Spectronaut was used to perform the database searches. Database search parameters included fixed modification on Cys (carbamidomethyl) with variable modification on Met (oxidation), Ser/Thr/Tyr (phosphorylation) and Protein N-term (acetylation). Full trypsin enzyme rules were used, with 10 ppm mass tolerances for precursor ions and 20 ppm for product ions.
Molecular dynamics simulation
We constructed five distinct simulation systems: CFN–Rho1–long-glucan (Glc0–16)-bound Fks1 (System 1), Rho1–long-glucan-bound Fks1 (System 2), CFN–Rho1–no-glucan-bound Fks1 (System 3), Rho1–trimmed-glucan (Glc0–3)-bound Fks1 (System 4) and the experimental short-glucan (Glc1–3)-bound Fks1 (with the gate closed) (System 5). All systems featured a membrane composed of DOPC, DOPE and ergosterol at a molar ratio of 68:17:15. The CHS molecules in the structures were replaced manually with ergosterol in Coot49 before system construction. The systems were assembled using the CHARMM-GUI Membrane Builder55,56,57,58 and neutralized with 0.15 M Na+ and Cl− ions. Simulations were performed using the CHARMM36m force field for lipids and proteins59,60,61, along with the TIP3P water model62. We ran 1-ns constant particle number, volume and temperature simulations and 10-ns constant particle number, pressure and temperature simulations to equilibrate the systems preliminarily. The CFN force field parameters were generated using the CHARMM General Force Field63. Production runs were conducted under NPT conditions at 303.15 K and 1 bar, with hydrogen mass repartitioning64 to enable a 4-fs timestep. All simulations were executed in GROMACS65 with input files generated by CHARMM-GUI. Production simulations of five replicas were performed for 1 μs for Systems 1–4 and 500 ns for System 5. VMD (v.1.9)66 was used for visualization of molecular dynamics simulation results; Python 3 and MDAnalysis (v.2.7.0)67 were used to analyse molecular dynamics results and generate molecular dynamics-related figures.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.