Tissue acquisition and processing for joint single-nucleus multiome profiling and spatial transcriptomics
Snap-frozen decidual and basal plate samples were obtained from the existing placenta tissue banks at Stanford University and University of California, San Francisco (Supplementary Tables 1 and 4). All samples were collected with written informed consent. Tissues were derived from women undergoing elective termination of presumed normal pregnancies (first and second-trimester samples; no known or predicted fetal chromosomal abnormalities, Extended Data Fig. 1c) or after term delivery (≥37 gestational weeks). For term samples, clinical records were reviewed to exclude placenta-associated complications (for example, chorioamnionitis); cases with NICU admission and preterm premature rupture of membranes were also excluded. Fresh placental tissues were grossly inspected and dissected under a microscope (Leica Microsystems) by pathologists. Decidua basalis was micro-dissected on ice from the MFI and distinguished from decidua parietalis and capsularis on the basis of characteristic histological and morphological features (Extended Data Fig. 1a,b). Dissected tissues were sequentially washed to remove residual blood cells in DMEM/H-21 medium, supplemented with 12.5% FBS (Hyclone), 1% L-glutamine (Atlanta Biologicals), 1% penicillin/streptomycin and 0.1% gentamicin and cold 1× PBS (Gibco, Thermofisher). Samples used for single-cell or spatial transcriptomic profiling were flash-frozen in liquid nitrogen and stored at −80 °C until processing. RNA quality was assessed from adjacent cryosections using a Bioanalyzer or Tapestation. For fresh frozen samples, only samples with RIN ≥ 7.0 were included.
Isolation of single nucleus from snap-frozen tissues
Single nuclei were isolated from snap-frozen tissues as previously described61 with modification. In brief, tissues were ground on dry ice, and 30–50 mg was homogenized into a pre-chilled 7 ml PYREX dounce homogenizer (Corning Life Science). Tissue was homogenized in 2 ml ice-cold buffer (250 mM sucrose, 0.3% NP-40, 5 mM MgCl2, 25 mM KCl, 10 mM Tris-HCl pH 7.8) supplemented with protease inhibitors (Roche, cOmplete) and 0.6 U µl−1 Ribolock RNase inhibitor (thermofisher). Debris was removed by 40 μm filtration, and nuclei were purified by OptiPrep iodixanol gradient centrifugation (25%, 30%, 40%). After centrifugation in a swinging bucket centrifuge (Eppendorf 5810R) for 30 min at 3,000g, nuclei were collected from the 30–40% interface, washed, and assessed by trypan blue staining and microscopy to ensure nuclei integrity. Approximately 15,000 nuclei per sample were processed using the Chromium Next GEM Single Cell Multiome ATAC + Gene Expression platform (10x Genomics).
Tissue preparation and CODEX imaging
Placenta samples for CODEX were OCT-embedded, cryosectioned at 10 µm, mounted on poly-L-lysine-coated slides and stored at −80 °C. On the day of staining, sections were equilibrated, acetone-treated, rehydrated, fixed with 1.6% paraformaldehyde, blocked, and incubated with a barcoded antibody cocktail (200 µl/section) for 3 h at room temperature. Sections were then washed, post-fixed with 4% paraformaldehyde and cold methanol, stabilized using CODEX fixative reagent, and stored in storage buffer at 4 °C (≤2 weeks) before imaging. Multiplex imaging was performed on an Akoya CODEX microfluidic system coupled to an inverted fluorescence microscope using a 7-cycle protocol (including blank cycles for alignment) across 4 channels (DAPI, FITC, Cy3 and Cy5) with a 20×/0.75 NA objective. Images were processed using CODEX Analysis Manager. The antibody panel included Akoya-validated barcoded antibodies and custom-conjugated antibodies generated using Akoya oligo barcodes following the manufacturer’s protocol (Supplementary Table 6). Additional details are provided in the Supplementary Note.
Single-nucleus multiome library construction and sequencing
Single-nucleus Multiome libraries were prepared using the Chromium Next GEM Single Cell Multiome ATAC + Gene Expression kit (10x Genomics) following the manufacturer’s protocol, using one reagent kit per sample. Around 15,000 isolated nuclei per sample were encapsulated into Gel Bead-In Emulsions (GEMs) containing unique cell barcodes, where reverse transcription and transposition occurred, followed by library amplification, and separation of gene expression and chromatin accessibility libraries. Libraries were sequenced on an Illumina NovaSeq 6000 using paired-end reads, with sequencing depth selected on the basis of recommendations from 10x Genomics. On average, each sample yielded approximately 250 million paired-end reads for ATAC and RNA libraries. Raw BCL files were demultiplexed, aligned to the GRCh38 (v.3.0.0) reference genome, and processed for barcode assignment, UMI counting, and quality control using Cell Ranger ARC v.2.0.0 (10x Genomics) (https://support.10xgenomics.com/single-cell-geneexpression/software/pipelines/latest/advanced/references).
Single-nucleus multiome data processing
High-quality nuclei with paired snRNA-seq and snATAC–seq profiles were retained using the following criteria: RNA UMI counts 1,000–50,000; detected genes >400; mitochondrial reads <20%; ATAC fragment counts 1,000–100,000; transcription start site enrichment >1.0; and nucleosome signal <2.0. Doublets were identified and removed using Scrublet62 with prior set to 0.1. After filtering, 191,735 nuclei were retained with paired snATAC- and snRNA-seq data. For snATAC–seq, open-chromatin peaks were called per sample using MACS2 (v.2.2.7)63, and merged into a unified peak set after excluding ENCODE blacklist regions64. Peak-by-cell count matrices were integrated across samples using reciprocal latent semantic indexing (LSI) projection in Signac12. For snRNA-seq, gene expression count matrices were integrated using reciprocal principal components analysis projection in Seurat (v.4)39. Prior to integration, data were normalized, scaled and feature-selected following best practices recommended in Seurat/Signac workflows12.
Dimensionality reduction was performed using principal components analysis on the top 3,000 variable genes (RNA) and LSI on the top variable ATAC peaks observed in at least 10 cells. UMAP embeddings for RNA/ATAC spaces were projected individually for visualization. Clustering was performed using the Louvain algorithm in Seurat/Signac12,39, and clusters were annotated on the basis of canonical marker genes (Supplementary Table 2) using gene expression and gene activity scores. Concordance between RNA- and ATAC-based clustering was assessed using the Jaccard index and percentage of cells assigned to same cell type.
Cell-of-origin assignment was performed using Souporcell10 with default parameters. Variants were called using FreeBayes based on 1000 Genomes Project reference, and only informative heterozygous loci (≥5 cells with both reference and alternative alleles) were retained. Trophoblasts were used as fetal reference populations, whereas lymphatic endothelial cells and DSCs were used as maternal reference populations to inform Souporcell assignment. Assignment accuracy was validated using UTY expression in samples from male fetuses, in which 94.3% of UTY+ nuclei were correctly classified as fetal.
Transcriptional regulation and ATAC–seq footprinting analysis
Putative enhancer peaks were identified by intersecting ATAC–seq fragments with experimentally validated enhancers from FANTOM5 datasets13. An enhancer-by-cell accessibility matrix was constructed by counting overlapping fragments per cell. Cell-type–specific enhancers were identified using a log-ratio test, comparing chromatin accessibility in each cell type to all others; enhancers with enriched accessibility (FDR < 0.05) were considered cell-type-specific.
Transcription factor activity was inferred using chromVAR11, which quantifies motif-associated accessibility deviations while correcting for GC content. Motif annotations were obtained from the JASPAR2020 database65. Cell-type-enriched transcription factors were those with significantly elevated chromVAR deviation scores in each cell type, identified by Wilcoxon rank-sum test. To validate transcription factor activities at the DNA level, transcription factor footprinting was analysed using Signac12. For each candidate transcription factor, footprinting profiles were generated over corresponding binding motif(s) within accessible chromatin regions after correcting for Tn5 insertion bias.
Gene regulatory network construction
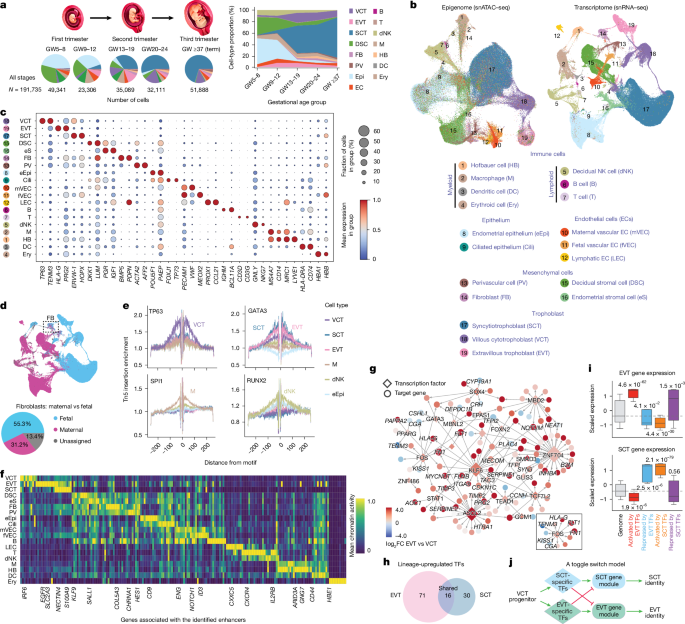
Cell type-specific GRNs were constructed using the CellOracle workflow15, integrating chromatin accessibility and gene expression data to model transcription factor–target gene interactions in a lineage-resolved manner. A base GRN scaffold was first constructed from snATAC–seq data via scanning transcription factor-binding motifs within proximal promoters and distal regulatory elements using GimmeMotifs (v.5.0)66. Proximal elements were annotated with HOMER67, and distal regulatory interactions were inferred on the basis of Cicero68 co-accessibility scores to link distal enhancers to putative gene targets. Using this scaffold, cell type-specific GRNs were inferred from snRNA-seq data via regularized Ridge regression implemented in CellOracle. Only statistically significant transcription factor–target gene interactions were retained. Among all transcription factors in the GRNs, DEG analysis identified 71 and 30 transcription factors significantly up-regulated (FDR < 0.01) in EVTs and SCTs, respectively, relative to VCTs. Regulatory interaction strength was evaluated by the absolute model-derived coefficients. Interactions with absolute coefficients exceeding 0.1 were visualized using Cytoscape69, and all identified regulatory interactions were included in our downstream analyses.
To identify lineage-specific regulatory rewiring, transcription factor–target gene pairs were classified into four categories: (1) EVT-specific activation (n = 224); (2) EVT-specific repression (n = 18); (3) SCT-specific activation (n = 234); and (4) SCT-specific repression (n = 48). Expression distributions of target genes in each category were compared with genome-wide background expression in EVT or SCT populations. For transcription factors up-regulated in both lineages, overlap between their target gene sets was quantified using the Jaccard index to assess shared regulatory programmes. Statistical comparisons were performed using the two-tailed Wilcoxon rank-sum test with Benjamini–Hochberg correction.
DEG and gene ontology analysis
DEGs between cell types or subtypes were compared using Wilcoxon rank-sums test by SCANPY70. A composite DEG score was derived for each gene that summarizes fold changes and the differences in the percentage of expressing cells between two cell types. DEGs were determined by FDR (FDR < 0.05). All non-ribosomal DEGs receiving the highest DEG scores were selected as signature genes to define: (1) aEC states (|DEG score| > 10); (2) eEVT, iEVT and pEVT (|DEG score| > 20); and (3) path A DSC and path B DSC (|DEG score|>20). These top ranked DEGs were included for the downstream gene ontology (GO) analysis using Enrichr71, and gene ontology terms with an FDR < 0.05 were considered statistically significant.
Pseudotime trajectory inference
For pseudotime analysis within arterial endothelial cells (spatial transcriptomics), trophoblasts (snRNA-seq) and DSCs (snRNA-seq), the initial and terminal cell states during cell differentiation were inferred by CellRank (v.1.5.1)72. Inferred initial states were selected as the developmental origin for pseudotime analysis by Palantir (v.1.1.0)32 with default settings. The fate probability to each identified terminal state was estimated for subsequent visualization. The major developmental branches identified by Palantir were visualized on UMAP projection. For trophoblast analysis, major branches with more than 500 cells were aggregated for stream plot.
Spatial transcriptomic library preparation and sequencing
The Stereo-seq (STOmics) spatial whole-transcriptome sequencing platform was applied to construct a high-resolution spatial transcriptomic atlas of the human MFI. Stereo-seq captures polyadenylated mRNA directly from tissue sections using spatially barcoded DNA nanoball (DNB) arrays, achieving subcellular resolution (~0.5 µm) with an effective spot diameter of ~220 nm. 16 basal plate samples (GW20–24) were embedded in Tissue-Tek OCT compound (4583, Sakura Finetek) and stored at −80 °C. The anatomical structure of cryosections was visually confirmed under a microscope by a pathologist on the day of the experiments. 10 µm cryosections were dissected using a Leica cryostat and mounted onto Stereo-seq transcriptomics T chips (Complete Genomics). Stereo-seq library preparation was performed per the manufacturer’s guidance (Supplementary Note). In brief, tissue sections were methanol-fixed and subjected to antibody or single stranded DNA (ssDNA) staining prior to tissue permeabilization. Released RNA was reverse-transcribed in situ, followed by tissue removal, cDNA amplification and SPRISelect bead purification. Libraries were sequenced on the DNBSEQ T7 platform (50 bp read 1, 100 bp read 2).
Spatial transcriptomic data analysis
Raw FASTQ files were processed using the STOmics SAW v.8.1 pipeline. Spatial barcodes (CIDs) were aligned to the chip coordinate grid, and high-quality reads were mapped to the human genome (GRCh38) using STAR73. Reads with identical CID–UMI pairs were collapsed to generate the final spatial gene expression matrix at bin1 resolution (0.5 nm × 0.5 nm). DAPI and ssDNA images were used to delineate cell boundaries using deep learning–based segmentation pipeline in the SAW workflow. To interrogate gene expression at single-cell resolution, read counts from all bins within a single cell’s boundary were aggregated to achieve CellBin resolution. Cells with low complexity (n_genes <100), excessive transcript counts (>10,000 MIDs or genes), or high mitochondrial gene content (>20%) were excluded. The count matrices in cells passing quality control were processed and normalized using Stereopy (v.1.6.1)27, in which the top 3,000 variable genes were selected for principal components analysis-based dimensionality reduction. Batch effects across donors were corrected by Harmony before UMAP visualization and clustering analysis74. Clusters were annotated on the basis of known cell-type- and subtype-specific gene signatures from snRNA-seq or known markers, and were then mapped to spatial coordinates for localization validation. Cell community and neighbourhood analysis were performed using Stereopy (v.1.6.1)27. Integrated visualization of cell segmentation, gene expression, and immunofluorescence signals was performed on StereoMap (v.4.1). Detailed experimental protocol and downstream spatial data analysis are described in Supplementary Note.
Spatially resolved cell–cell communication analysis
Intercellular signalling among spatially resolved cell types was analysed using CellChat (v.2)36. Analyses were performed with the CellChatDB (the human ligand–receptor database), following the standard workflow. Spatial coordinates were incorporated to constrain inferred intercellular interactions within a maximum Euclidean distance of 200 µm, consistent with the expected physical range of paracrine signalling. Ligand–receptor pairs were grouped into signalling pathways, and communication probabilities were aggregated to estimate pathway-level and overall interaction strengths between cell-type pairs. Communication scores were standardized for comparison. Statistical significance was assessed by label-shuffled permutation (n = 1,000) and interactions with Benjamini–Hochberg corrected P < 0.05 were considered significant.
Model-based prediction of EVT invasiveness
Gene signatures defining EVT invasiveness potential were identified via a supervised machine learning framework using L1-regularized regression (LASSO) based on spatial transcriptomic profiles. All identified EVTs excluding those associated with spiral arteries from sixteen sections were split into training and testing datasets at a 1:1 ratio. EVTs were ordered by their distance to the MFI. We then grouped ten cells with similar MFI distances within a narrow depth window to generate pseudobulk profiles, thereby matching spatial context across samples and reducing sparsity and batch effects during model training. A LASSO regression model was trained on the averaged gene expression as predictors and distance to the MFI as the response variable. The feature space comprised 3,192 genes enriched in EVTs relative to other cell types from the independent snRNA-seq data and analysis. Gene expression from training and test datasets were independently standardized while preventing information leakage from the training data into the test data. The model agnostically identified 54 predicting EVT invasiveness potential, with nonzero coefficients, where positive and negative coefficients indicated pro-invasive and anti-invasive effects, respectively. The trained model was applied to single-cell data to generate predicted invasion scores (iScores). Model performance was evaluated by Spearman correlation between predicted iScores and observed distance of each EVT (the held-out test dataset) to the MFI. iScores were normalized within each tissue section for cross-sample comparisons.
To benchmark iScores, the trained model was applied to two independent EVT scRNA-seq datasets, including second-trimester smooth chorion and placental villi (GSE198373)40, term placental tissues from placenta accreta spectrum (PAS) patients and healthy controls (GSE212505)41. Gene expression matrices were log-normalized, z-score standardized, and used as input to the trained model. Predicted iScores were compared across tissue contexts and disease states to evaluate differences in the predicted EVT invasiveness. To assess the effects of DSC subtypes on neighbouring EVTs, iScores of EVTs in direct spatial proximity to annotated DSC subtypes (within five neighbouring tiles on grids, by Squidpy75) were analysed and compared with depth-matched EVTs that were not in spatial proximity to any DSCs. Statistical comparisons were performed using the two-tailed Wilcoxon rank-sum test with Benjamini–Hochberg correction for multiple testing.
Immunofluorescence
Immunofluorescence was performed as previously described40,76. Placenta tissues were fixed in 3% paraformaldehyde in PBS for 30 min, followed by cryoprotection in a sucrose gradient (5%, 10%, 15% in PBS). Tissues were embedded in OCT (Tissue-Tek, Sakura Finetec), sectioned at 10 µm using a Leica cryostat, and mounted on poly-l-lysine-coated slides (Electron Microscopy Sciences). The tissue sections on coverslips were permeabilized using 0.3% Triton X-100 for 10 min and blocked in 1% BSA in PBS for 30 min. Sections were incubated with primary antibodies (Supplementary Table 6) in blocking buffer for 2 h at room temperature, washed four times with PBS, and incubated with Alexa Fluor 488- or 647-conjugated secondary antibodies (1:200; Jackson ImmunoResearch) for 1 h. After PBS washes, nuclei were counterstained with DAPI. Slides were mounted using ProLong Gold Antifade Mountant and stored at 4 °C in the dark prior to imaging. Images were acquired using a Leica DM5000 B microscope under optimized settings.
Single-molecule fluorescent in situ hybridization with RNAscope
Freshly dissected basal plate samples were fixed in 4% paraformaldehyde for 30 min at room temperature, embedded in OCT, and cryosectioned at 15 μm. Sections were permeabilized and subjected to target retrieval by protease prior to probe hybridization. In situ hybridization was performed using the RNAscope Multiplex Fluorescent Reagent Kit v.2 (Advanced Cell Diagnostics) following the manufacturer’s instructions. Sections were hybridized with a GPC5-pecific probe for 2 h at 40 °C in a humidified chamber, followed by signal amplification and detection using TSA-conjugated fluorophores. After hybridization, sections were washed in PBS and processed for immunofluorescence staining with a FITC-conjugated anti-CD94 antibody (BD Biosciences, 555888). Nuclei were counterstained with DAPI. Sections were mounted with ProLong Gold Antifade Mountant (Invitrogen) and stored at 4 °C prior to imaging. Images were acquired using a STELLARIS 5 Confocal Microscope (Leica Microsystems) with consistent acquisition settings across samples. Multiple representative fields were captured per section from three biological replicates. Quantification of GPC5+ cells was performed in QuPath using automated cell segmentation within defined regions of interest (syncytial aggregates or membrane), with manual verification. Results were reported as the percentage of GPC5+ cells among total DAPI+ nuclei. Statistical significance was assessed using two-tailed Student’s t-test.
In vitro decidualization and mAEA treatment
The HuF cells were collected from human term placenta the decidualized in vitro as described48. In brief, the decidual tissues were micro-dissected from the chorionic layer and enzymatically dispersed to reach over 95% purity. The cells are cultured in RPMI medium supplemented with 2% fetal bovine serum and antibiotics. The cells were decidualized for a continuous 7 days by adding medroxyprogesterone acetate (1 μM), oestradiol (10 nM) and prostaglandin E2 (1 μM) after reaching 90% confluence. On day 5, 500 nM mAEA or ethanol was added to the culture for 72 h. The conditioned media were collected 24 h after the medium replacement. In parallel experiments, the cells were collected by trypsin digestion on day 7 for 10× Chromium 3′ single-cell RNA sequencing. To identify enriched pathways in decidualized stromal cells following mAEA treatment, gene set enrichment analysis was performed. Genes were ranked by their log-fold changes from differential expression analysis, and pathways were considered significantly enriched at a threshold of FDR < 0.25, as recommended by gene set enrichment analysis guidelines.
Transwell invasion assay
Transwell invasion assays were performed using 24-well inserts with 8 µm pore-size polycarbonate membranes (Corning Costar, 3422). Membranes were coated with 10 µl of Matrigel (Corning) diluted 1:1 in serum-free DMEM/H-21 supplemented with 2% Nutridoma, 1% Penicillin-Streptomycin, 1% HEPES and 0.1% gentamicin. Trophoblasts isolated from second-trimester basal plate were resuspended in serum-free medium and seeded into the upper chamber at 2.5 × 105 cells per insert. The lower chamber contained conditioned medium from in vitro decidualized HuFs treated with vehicle, mAEA (methanandamide), rimonabant (SRI141716) or mAEA plus rimonabant. Cells were incubated at 37 °C with 5% CO2 for 48 h to allow invasion through the Matrigel-coated membrane. Following incubation, inserts were fixed and permeabilized in ice-cold methanol for 20 min, washed with PBS, and stained with DAPI for 5 min at room temperature. Membranes were mounted on slides and imaged at 40× magnification under a fluorescence microscope (Leica Microsystems), with at least 5 non-overlapping fields acquired per insert. Each condition was tested on primarily isolated cytotrophoblasts from three independent placentas. Invaded cells stained with DAPI were counted manually and normalized to the total number of DAPI-positive cells per field. Comparisons between conditions were performed using a two-tailed Student’s t-test, with P values < 0.05 considered statistically significant.
Single-cell risk map for pregnancy complications
Single-cell trait relevance scores (TRS) were computed using SCAVENGE by integrating snATAC–seq with GWAS summary statistics of pregnancy complications50. Cell-type-specific ATAC peaks were identified using MACS2 (v.2.2.7.1; default)63, and cell-by-peak matrices were generated as input for recommended SCAVENGE workflow. Analyses were performed using GWAS datasets for pre-eclampsia (maternal and fetal genomes; EGAD00010001984 and EGAD00010001986)51, spontaneous preterm birth (maternal genomes)52,53, and sporadic miscarriage (maternal genomes)54. Risk-associated cells were defined as cells receiving top 5% TRS scores by permutation testing within SCAVENGE50. Genetically vulnerable cell types were determined by the two-tailed Fisher’s exact after Benjamini–Hochberg correction, in which cell types with fewer than 200 cells were excluded. As negative controls, identical analyses using pre-eclampsia GWAS data were performed on adult and prenatal brain single-cell datasets56,57 (adult: GSE204684; fetal: https://assets.nemoarchive.org/dat-oiif74w), in which the percentage of genetically vulnerable cells in each cell type was normalized for visualization.
Ethics statement
All human pregnancy tissue samples were obtained with informed consent and processed in accordance with the Declaration of Helsinki. Study protocols were approved by the Stanford institutional review board (31552, 34745, 48255 and 46584) and University of California, San Francisco institutional review board (11-05530 and 10-00350). All samples were de-identified before processing.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.