Single guide RNA design and cloning
The gRNA used for SGE was designed using Benchling’s CRISPR design tool to search the RNU4-2 locus, including upstream and downstream regions of low sequence homology to RNU4-1 and pseudogenes, identifying a candidate with high on-target and low off-target scores. The selected gRNA was not predicted to target RNU4-1, owing to eight mismatches occurring in the protospacer and PAM. The gRNA spacer sequence was ligated into the pX459 backbone as previously described33. In brief, complementary primers containing the spacer were ordered from IDT (Supplementary Table 3), phosphorylated, hybridized and ligated into the pX459 linearized backbone followed by PlasmidSafe DNase (Lucigen) digestion. Next, 2 µl of the ligation reaction were transformed in NEB Stable Competent Escherichia coli cells using the high-efficiency transformation protocol and 75 µl of transformant was plated on ampicillin-resistant plates and cultured overnight at 30 °C. Three colonies were then picked and grown overnight at 37 °C in 7 ml of Luria–Bertani medium supplemented with carbenicillin (100 µg ml−1). Plasmid DNA was extracted using the QIAprep Spin Miniprep kit (Qiagen) and verified using Plasmidsaurus whole-plasmid sequencing. The selected clone was then grown in 100 ml of Luria–Bertani medium at 37 °C in a shaking incubator supplemented with carbenicillin. The cells were then pelleted and the plasmid was extracted using a ZymoPure Maxiprep kit (Zymo Research), endotoxins were removed using EndoZero columns (Zymo Research) and the product was quantified with the Qubit double-stranded DNA (dsDNA) BR assay kit (Invitrogen).
HDR library cloning
An oligonucleotide library comprising RNU4-2 variants was manufactured by Twist Bioscience and subsequently cloned into a vector containing homology arms for RNU4-2 to make the HDR library for SGE.
To generate the vector with homology arms, a nested PCR was performed on genomic DNA (gDNA) extracted from HAP1 cells10 using primers designed to generate homology arms of 700–800 base pairs (bp) flanking RNU4-2 (Supplementary Table 3). The PCR was performed using the Kapa HiFi HotStart ReadyMix (Roche). The product was purified using AmpureXP (Beckman Coulter) magnetic beads at 1.2× volume and eluted in 12 µl of nuclease-free water. The amplicon containing RNU4-2 homology arms was then inserted in the linearized pUC19 backbone using In-Fusion HD cloning (Takara) and 2 µl of cloning reaction was transformed into NEB Stable cells following the manufacturer’s 5-min transformation protocol. Cells were plated on agar plates containing ampicillin and incubated at 30 °C overnight. The pUC19 plasmid containing RNU4-2 homology arms (pUC19-RNU4-2-HA) was purified and sequence-verified from a successfully transformed clone. pUC19-RNU4-2-HA was then diluted to 8.7 pg in a 50-µl PCR reaction and amplified with Kapa HiFi to obtain a linearized product with 17–18 bp complementarity to the RNU4-2 oligo library. A PAM-blocking mutation was introduced 27 nt upstream of the RNU4-2 sequence (chromosome 12:120291930-C-G) by means of primer overhang extension during PCR. The location of the PAM-disrupting edit was selected to minimize recutting by Cas9, converting a 5′-GGG PAM sequence to 5′-GCG. The PAM-disrupting edit had a CADD score of 4.20 (Phred) and a 100 vertebrates PhyloP score of 0.11. The reaction was treated with 1 µl of DpnI (NEB) for 30 min at 37 °C, gel extracted and quantified. Then, the RNU4-2 oligo library was amplified using Kapa HiFi and purified using AmpureXP (1.2×). The amplified library and linearized pUC19-RNU4-2-HA plasmid were then assembled using the In-Fusion HD cloning kit, and the product was transformed into NEB Stable cells using the high-efficiency transformation protocol. To quantify efficiency, 1% of cells in the transformation reaction were plated and the remainder were cultured in 100 ml of Luria–Bertani medium with carbenicillin overnight at 37 °C. Cells were then pelleted by centrifugation and the final RNU4-2 HDR library was extracted using the ZymoPure Maxiprep kit (Zymo Research) with endotoxin removal. The isolated HDR library was quantified with a Qubit dsDNA BR assay kit and sequence-verified by Plasmidsaurus.
HAP1 cell culture
HAP1 cells used for SGE (the HAP1-LIG4-KO line; herein referred to as ‘HAP1’) show increased rates of editing by HDR due to a frameshifting mutation in LIG4 (ref. 10). Frozen HAP1 cells were thawed at 37 °C in a water bath, then supplemented with 10 ml of prewarmed Iscove’s Modified Dulbecco’s Medium (IMDM) containing l-glutamine, 25 nM HEPES (Gibco), 10% FBS (Gibco), 1% penicillin–streptomycin (Gibco) and 2.5 μM 10-deacetyl-baccatin-III (DAB, Stratech), herein referenced to as IMDMc. Cells were centrifuged at 300g for 3 min. The supernatant was then aspirated and the cells were resuspended in fresh media, plated on a 10-cm dish and cultured at 37 °C with 5% CO2. The next day, the IMDMc media was replaced, and cells were cultured routinely from that point forward.
The HAP1 subculture routine included a 1:5 split every 48 h or 1:10 split every 72 h to prevent cells from exceeding 80% confluency. To split cells, the media was aspirated and the dish washed with 10 ml of room-temperature Dulbecco’s PBS (Gibco). Following Dulbecco’s PBS aspiration, the cells were treated with 1 ml of 0.25% trypsin–EDTA (Gibco) and incubated for 3 min at 37 °C. Next 14 ml of prewarmed IMDMc was then added and cells were collected and centrifuged at 300g for 5 min. Cells were then resuspended in 10 ml of IMDMc, counted and seeded on a 10-cm dish.
Generation of diploid HAP1 cells
Parental HAP1 cells were cultured for 9 days after thawing in IMDMc without DAB supplementation to allow for the spontaneous occurrence of diploid cells. On day 10, cells were stained with 5 µg ml−1 Hoechst working solution (Thermo Fisher Scientific) for 1 h at 37 °C, followed by fluorescence-activated cell sorting to select diploid cells using a BD FACSAria Fusion Flow Cytometer. Diploid cells were sorted on the basis of their G2/M peak (4n), with gates established using a monoclonal diploid HAP1 control population. Sorted diploid HAP1 cells were then expanded for 10 days in IMDMc without DAB supplementation before the subsequent SGE experiment.
Transfection and selection
The day before transfection, 12 million cells were seeded on a 10-cm dish for each replicate and 2 million cells were seeded on a six-well plate for the negative control sample. On the day of transfection (day 0), a transfection mix containing 10 µg of HDR library, 30 µg of the pX459 gRNA plasmid and 24 µl of Xfect polymer (Takara) in a final volume of 800 µl was prepared according to the manufacturer’s instructions for each replicate. For the negative control sample, a pX459 plasmid with a gRNA targeting HPRT1 (ref. 13) instead of RNU4-2 was used to prevent successful editing, and the transfection volume mix was scaled down eightfold. Following transfection, cells were incubated for 24 h at 37 °C and supplemented with prewarmed IMDMc with 1 µg ml−1 puromycin (Cayman Chemical). On day 4, half of the cells for each replicate were collected for gDNA extraction and stored as a pellet at −70 °C; the rest were kept in culture in 15-cm dishes supplemented with 15 ml of IMDMc. The negative control sample was collected when reaching 70% confluency at day 6. A second sample of 10 million cells per replicate was collected at day 14 and stored at −70 °C.
Sequencing library preparation
gDNA was extracted from cells using QIAshredder (Qiagen) columns followed by the Allprep DNA/RNA kit (Qiagen) according to the manufacturer’s instructions. Concentrations were determined using the Qubit dsDNA BR assay kit. The RNU4-2 locus was subsequently amplified using nested PCR to avoid amplification of plasmid DNA, followed by an indexing PCR, in total using three primer sets (Supplementary Table 3). For the first reaction, the total gDNA template from each condition was partitioned into separate reactions, each containing 1.25 µg of DNA in a 100 µl reaction volume, using NEBNext Ultra II Q5 master mix (NEB) supplemented with MgCl2 (Ambion) to a final concentration 4 mM. The amplification reaction was monitored by quantitative PCR (qPCR) using SYBR green (Invitrogen) and stopped before completion. The reactions for each sample were pooled and mixed before 50 µl of each product was purified using AmpureXP (1.2×) and eluted in 15 µl of nuclease-free water. Then 1 µl of purified product was loaded into the second qPCR reaction (50 µl final volume) and amplified using NEBNext Ultra II Q5. The reaction was again monitored using SYBR green and stopped before completion. The AmpureXP purification was then repeated, and a final qPCR (NEBNext Ultra II Q5) to incorporate sample indexes and sequencing adapters was performed using 1 µl of purified product as template in a 50 µl reaction for 8 cycles. Final products were purified and quantified with the Qubit dsDNA HS kit. The samples were then pooled for sequencing, aiming for 5 million reads per experimental replicate timepoint, 2 million reads for the negative control sample and 1 million reads for the HDR library. The pool was purified using AmpureXP (1×), quantified and loaded on a Novaseq X sequencer (Illumina).
Variant frequency quantification
The fastq files were de-multiplexed using the bcl2fastq script and the variants were quantified as previously described13. In brief, paired-end reads were adapter trimmed and merged, and reads containing N bases were discarded. HDR editing rates were computed from fastq files directly as the fraction of reads containing the exact PAM-blocking mutation. Fastq files were then aligned to a reference RNU4-2 sequence and the frequency of each variant included in the library was determined.
Function score calculation
All variants were observed in the library and day 4 at a frequency higher than 10−4, and were therefore included in downstream analyses. Function scores for library variants were first calculated per replicate, computed as the log2 ratio of day 14 to day 4 variant frequencies, normalized by subtracting the median function score of negative control insertions from all scores. Final function scores were then calculated for each variant by averaging function scores across replicates, again normalizing to the median of negative control insertions such that the median final function score of control insertions equals 0. For each variant, P values were determined using the norm.cdf function in Python, defining a normal distribution from the mean and standard deviation of function scores for negative control insertions. The P values were corrected for multiple hypothesis testing using the multipletests function in Python (Benjamini–Hochberg procedure) to derive q values. Significantly depleted variants were defined as those with q < 0.01, corresponding to a function score below −0.302. We further classified depleted variants into two categories using an arbitrary function score threshold of −0.9 to include sufficient variants and individuals per category to assess for phenotypic differences.
Variant scoring with CADD and ViennaRNA
Variants were annotated as ReNU syndrome variants if they were reported in ref. 1 or classified as pathogenic or likely pathogenic in ref. 4. Variants were annotated with whether or not they were observed in the 490,640 genome sequenced individuals from the UK Biobank18 (DRAGEN pipeline) or in 414,840 individuals from All of Us V8. CADD v.1.7 (ref. 19) annotations were obtained by uploading a synthetic VCF to the online annotation tool (https://cadd.gs.washington.edu/score). As we preselected which insertions and deletions to include in the SGE assay (because of assay size limitations), we restricted analyses involving CADD to SNVs within the RNU4-2 transcript.
For variants assayed within the RNU4-2 transcript, predicted changes in U4/U6 interaction stability (ΔΔGbind) were computed using the ViennaRNA package34 (v.2.7.0). Minimum free energies (MFEs) were obtained by use of RNA.fold_compound() at 37 °C using default Turner RNA thermodynamic parameters. U4/U6 pairing was modelled with the ViennaRNA cofold grammar by providing sequences in the dimer format (u4(AGCUUUGCGCAGUGGCAGUAUCGUAGCCAAUGAGGUUUAUCCGAGGCGCGAUUAUUGCUAAUUGAAAACUUUUCCCAAUACCCCGCCAUGACGACUUGAAAUAUAGUCGGCAUUGGCAAUUUUUGACAGUCUCUACGGAGACUGA).
+ ‘&’ + u6(GUGCUCGCUUCGGCAGCACAUAUACUAAAAUUGGAACGAUACAGAGAAGAUUAGCAUGGCCCCUGCGCAAGGAUGACACGCAAAUUCGUGAAGCGUUCCAUAUUUU), and the intermolecular MFE was extracted using mfe_dimer(). Single-strand MFEs for U4 and U6 were computed independently using mfe().
Binding free energy was defined as:
$$\Delta {G}_{{\rm{bind}}}=\Delta {G}_{{\rm{complex}}}-(\Delta {G}_{{\rm{U}}4}+\Delta {G}_{{\rm{U}}6})$$
The same procedure was applied to RNU4-2 variant sequences, and differential stability was then calculated as:
$$\Delta \Delta {G}_{{\rm{b}}{\rm{i}}{\rm{n}}{\rm{d}}}=\Delta {G}_{{\rm{b}}{\rm{i}}{\rm{n}}{\rm{d}}.{\rm{v}}{\rm{a}}{\rm{r}}{\rm{i}}{\rm{a}}{\rm{n}}{\rm{t}}}-\Delta {G}_{{\rm{b}}{\rm{i}}{\rm{n}}{\rm{d}}.{\rm{r}}{\rm{e}}{\rm{f}}{\rm{e}}{\rm{r}}{\rm{e}}{\rm{n}}{\rm{c}}{\rm{e}}}$$
Positive ΔΔGbind values indicate predicted destablization of U4/U6 pairing.
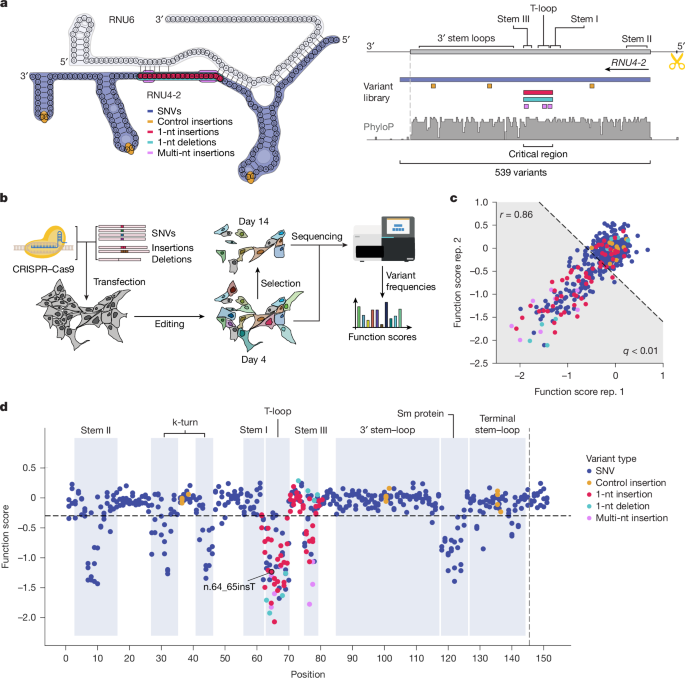
Variants were mapped to the following structural regions of RNU4-2: Stem II (n.3 to n.16), k-turn within the 5′ Stem loop (n.27 to n.35 and n.41 to n.46), Stem I (n.56 to n.62), T-loop (n.63 to n.70), Stem III (n.75 to n.79), 3′ Stem loop (n.85 to n.117), Sm protein (n.118 to n.126) and terminal Stem loop (n.127 to n.144).
ROC area under the curve (AUC) values were calculated by assigning a 1 label to ReNU syndrome SNVs and a 0 label for SNVs observed in UK Biobank or All of Us. The labels and corresponding function scores were used to compute false positive and true positive rates (using Python’s roc_curve function), as well as ROC-AUC values (using the roc_auc_score function). This analysis was also restricted to SNVs only.
Assigning evidence codes to variants based on function score
We followed established guidelines8 to calibrate function scores from SGE experiments in haploid cells to evidence strengths for classification of ReNU syndrome variants. To do so, we defined a gold standard set of pathogenic, dominantly inherited variants as the 17 previously reported4 as ‘pathogenic’ or ‘likely pathogenic’ for which we derived function scores. Few RNU4-2 variants have been deemed benign in ClinVar, so we instead used reported allele counts in the UK Biobank and All of Us studies to define a neutral set of variants. This included all 45 assayed variants with a combined allele count of more than 100 between the two studies. A two-component Gaussian mixture model was then fit from the function score distributions of these variant sets, using the ‘Mclust’ package in R. This model was then used to determine the probability of pathogenicity for each variant in the CR based on function score. The resulting posterior probabilities were then converted to OddsPath values using a uniform prior of 0.5, and evidence codes were assigned according to established OddsPath thresholds8 with the exception that PS3 evidence was capped at strong (+4 points), in line with the limited number of gold standard variants available for calibration. We did not apply the model to variants outside the CR on account of there being no known pathogenic variants for ReNU syndrome in these regions.
Phenotype severity and clustering
Categorical data for 44 clinical features from 143 patients with pathogenic and likely pathogenic RNU4-2 variants4 were transformed into a 0–1 scale, with 0 indicating a more favourable phenotype and 1 a more severe presentation. Principal component analysis was generated after imputing missing data with 0 and performing variable scaling. UMAP representation was created using the umap package in R. Two-sided Fisher’s tests with Bonferroni adjustment to account for four tests were used to compare clinical features between SGE function score variant categories (strong versus moderate) in Extended Data Table 1.
RNA sequencing cluster analysis
RNA sequencing from cultured lymphocytes was performed following the protocol described in ref. 4 for RNU4-2 and rMATS-turbo (v.4.3.0)35 was run on 19 ReNU samples and 20 control participants (excluding one individual previously deemed a control in ref. 4 who was here found to be a recessive RNU4-2 case); 101 significant alternative non-canonical 5′ splice sites (A5SS) events (false discovery rate less than 0.1, ΔPSI > 0.05) were retained. Then rMATS-turbo was rerun on the 19 ReNU samples, the 20 control participants, without statistical or ΔPSI filtering. The A5SS output was filtered on the 101 retained events and the PSI values were extracted to perform the principal component analysis.
Association testing in UK Biobank
We extracted phenotypes associated with educational attainment from UK Biobank following an approach published previously36. Fluid intelligence scores (field ID 20016) were retrieved for all participants. Where many scores were recorded, the median value was taken. Age left education was calculated as the maximum value in age completed full time education (field ID 845). Diagnosis with childhood developmental disorder was defined using the ICD codes for intellectual disability (ICD-10: F70–F73, F78, F79; ICD-9: 317, 318, 319), epilepsy (ICD-10: G40), global developmental disorders (ICD-10: F80–F84, F88–F95, R62, R48, Z55; ICD-9: 299, 312, 313, 314, 315) and congenital malformations (ICD-10: Q0–Q99, ICD-9: 740–759).
We identified UK Biobank participants with: (1) depleted variants in the 18-bp RNU4-2 CR (n = 6), (2) depleted variants outside the CR (n = 50) and (3) participants with non-depleted SNVs outside the CR (n = 12,132). We performed multiple linear regression on fluid intelligence scores and age left education, and multiple logistic regression on childhood developmental disorder for variant groups (2) and (3) defined above, compared with all individuals without any variants in any of the three groups. Age at recruitment (field ID 21022), age2 (age at recruitment × age at recruitment), sex (field ID 31) and first ten genetic principal components (field ID 22009) were included as covariates. P values were false discovery rate-corrected using the Benjamini–Hochberg method.
Investigating RNU4ATAC variants in ClinVar
Variants in RNU4ATAC with classifications of pathogenic, likely pathogenic, pathogenic or likely pathogenic, benign, likely benign or benign or likely benign were downloaded from the ClinVar37 website on 4 March 2025. Two regions of RNU4-2 and RNU4ATAC with identical structures were defined, mapping to the k-turn (RNU4-2 nucleotides 26–52; RNU4ATAC nucleotides 31–57) and the Sm protein binding site (RNU4-2 nucleotides 115–126; RNU4ATAC nucleotides 113–124). Variants at the same nucleotide in the structure and where the reference bases in RNU4-2 and RNU4ATAC are identical, were marked as ‘equivalent’.
Identifying biallelic variants in cohorts
We searched rare disease cohorts for individuals with biallelic variants in RNU4-2. These cohorts included the Genomics England 100,000 Genomes Project and NHS Genomic Medicine Service datasets accessed through the UK National Genomic Research Library38, the SeqOIA and Auragen clinical cohorts in France (PFMG2025), the Undiagnosed Disease Network, the Broad Institute Center for Mendelian Genomics and GREGoR (Genomics Research to Elucidate the Genetics of Rare Diseases)39 Consortium cohorts. We only included individuals with homozygous variants with function scores less than −0.302, or compound heterozygous variants in which both had function scores less than −0.302 (n = 20). All individuals had previous genome analysis including investigation of variants in known NDD genes and large structural variants. One individual (individual 17) had a reported likely pathogenic variant in GLI3; however, this variant did not explain all of their reported phenotypes (see ref. 21 for more details).
Ethics
Informed consent was obtained for all participants included in this study from their parent(s) or legal guardian, with the study approved by the local regulatory authority. The 100,000 Genomes Project Protocol has ethical approval from the Health Research Authority Committee East of England Cambridge South (Research Ethics Committee ref. 14/EE/1112). This study was approved by Genomics England under Research Registry Projects 354. Health related research in UK Biobank was approved by the Research Ethics Committee under reference 20/NW/0274 with this research conducted under application number 81050.
We received an exception to the Data and Statistics Dissemination Policy from the All of Us Resource Access Board to report questionnaire response data for the single individual with a homozygous depleted variant as well as variant counts below 20 for all variants in RNU4-2.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.