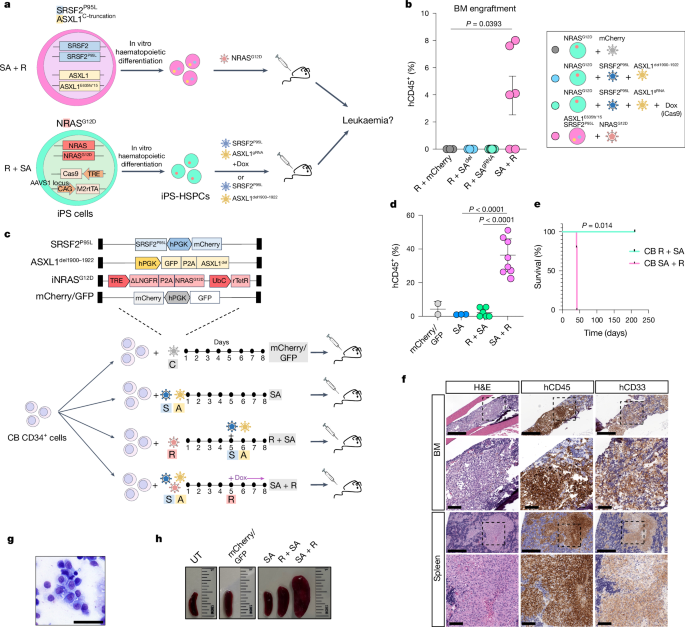
Gene editing of human iPS cells
We used the previously described normal iPS cell line N-2.12-D-1-1 as the parental line to generate all CRISPRâCas9-edited lines described in this study, unless otherwise specified36. For the edited lines described in Extended Data Fig. 2d, we used a patient-derived RUNX1-FPD line harbouring a heterozygous RUNX1 mutation (NM_00100189: c.533-1G>T) as parental line50,51. The gene editing strategies used to generate ASXL1Câterminus truncation, SRSF2P95L mutation, NRASG12D mutation and FLT3-ITD were described previously27,28,52. Several independent clones with each mutation were isolated after each gene editing step and, following genetic and preliminary phenotypic characterization to exclude potential outliers, one clone was selected for each subsequent editing step.
We used CRISPRâCas9-mediated homology-directed repair (HDR) to introduce the DNMT3AR882H mutation using co-delivery of a mutant and a WT donor template (Extended Data Fig. 2a) as previously described27. Nucleofection of a plasmid expressing the gRNA and Cas9 with mCitrine and clone selection by restriction fragment length polymorphism analysis were performed as previously described27. In brief, the N-2.12-D-1-1 iPS cell line was cultured in hESC medium containing 10âmM Y-27632 for at least 1âh before nucleofection. The cells were dissociated into single cells with accutase and 1âmillion cells were used for nucleofection with 5âµg of gRNA/Cas9 plasmid and 5âµg of each donor plasmid (WT and G12D) using Nucleofector II (Lonza). mCitrine+ cells were FACS-sorted 48âh after transfection and plated at clonal density. Single colonies were screened by PCR and restriction fragment length polymorphism analysis with DdeI restriction enzyme.
An NRASG12D iPS cell line (NRAS-66) was engineered to introduce a TRE-driven Cas9 and the M2rtTA in the two alleles of the AAVS1 locus (Extended Data Fig. 2f) by TALEN-mediated gene targeting, as described53.
Human iPS cell culture and haematopoietic differentiation
Derivation of the AML-9.9, AML-4.10, AML-4.24, N-2.12, AML-37.16 and AML-47.1 iPS cell lines has been described previously28,30,36. Human iPS cells were cultured on mitotically inactivated mouse embryonic fibroblasts as described previously52. Haematopoietic differentiation used a spin-EB protocol as described previously52. For monocytic differentiation, dayâ11â16 HSPCs were transferred to StemPro-34 SFM medium with 1% non-essential amino acids, 1âmM l-glutamine and 0.1âmM β-mercaptoethanol, supplemented with 100ângâmlâ1 macrophage colony-stimulating factor and 25ângâmlâ1 interleukin-3 for 3â30âdays with medium changes every 2âdays. In the end of the differentiation culture, the cells were collected and dissociated with accutase into single cells and used for flow cytometry, cytological analyses, VEN treatment or transplantation into immunodeficient mice.
Cytological analyses
Approximately 200,000 cells from liquid haematopoietic differentiation cultures were washed twice with PBS containing 2% FBS and resuspended in PBS. Cytospins were prepared on slides using a Shandon CytoSpinâIII cytocentrifuge (Thermo Electron). Slides were then air-dried for 30âmins and stained with the Hema 3 staining kit (Fisher Scientific). The slides were read on a Nikon Eclipse Ci microscope and digital images were taken with a Nikon DS-Ri2 camera and NIS-Elements D4.40.00 software.
Flow cytometry and FACS-sorting
The following antibodies were used: CD34-PE (clone 563, catalogue no. 550761, BD Pharmingen, 1:100 dilution), CD34-BV711 (clone 563, catalogue no. 740803, BD Biosciences, 1:100 dilution), CD45-APC (clone HI30; catalogue no. 555485, BD Pharmingen, 1:100 dilution), mCD45-PE-Cy7 (clone 30-F11, catalogue no. 552848, BD Pharmingen, 1:100 dilution), CD33-BV421 (clone WM53, catalogue no. 562854, BD Biosciences, 1:100 dilution), CD19-PE (clone HIB19, catalogue no. 561741, BD Biosciences, 1:100 dilution), CD19-BV650 (clone HIB19, catalogue no. 740568, BD Biosciences, 1:100 dilution), CD38-PE-Cy7 (clone HIT2, catalogue no. 980312, Biolegend, 1:100 dilution), CD123-BV421 (clone 7G3, catalogue no. 563362, BD Biosciences, 1:20 dilution), CD45RA-APC (clone MEM-56, catalogue no. MHCD45RA05, ThermoFisher Scientific, 1:100 dilution), CD68-PE-Cy7 (clone Y1/82âA, catalogue no. 565595, BD Pharmingen, 1:100 dilution), CD11b-BB515 (clone ICRF44, catalogue no. 564517, BD Biosciences, 1:100 dilution), CD11b-BV650 (clone ICRF44, catalogue no. 301336, Biolegend, 1:100 dilution), CD14-APC (clone M5E2, catalogue no. 555399, BD Biosciences, 1:100 dilution), CD14-BV421 (clone M5E2, catalogue no. 565283, BD Biosciences, 1:100 dilution) and CD271 (LNGFR)- APC-Cy7 (clone ME20.4; catalogue no. 345125, Biolegend, 1:2,000 dilution). Cell viability was assessed with 4,6-diamidino-2-phenylindole (DAPI; Life Technologies). Cells were assayed on a BD Fortessa or BD Symphony A5 SE and data were analysed with FlowJo software (Tree Star). Cells were sorted on a BD FACS Aria II.
iPS- and CB-derived HSPC culture and lentiviral transduction
CB CD34+ cells were purchased from AllCells and cultured in X-VIVO 15 medium with 1% non-essential amino acids, 1âmM l-glutamine, 0.1âmM β-mercaptoethanol and 20% BIT 9500 serum substitute (Stem Cell Technologies) and supplemented with 100ângâmlâ1 stem cell factor, 100ângâmlâ1 Flt3 ligand, 100ângâmlâ1 thrombopoietin and 20ângâmlâ1 interleukin-3 for 1â4âdays. Lentiviral vector packaging and cell transduction with viral supernatants in the presence of 4âµgâmlâ1 polybrene were performed as described previously36.
Transplantation into NSG and NSGS mice
All mouse studies were performed in compliance with Icahn School of Medicine at Mount Sinai laboratory animal care regulations and approved by an Institutional Animal Care and Use Committee. NSG (NOD.Cg-PrkdcscidIl2rgtm1Wjl/SzJ) and NSGS (NOD.Cg-PrkdcscidIl2rgtm1WjlTg(CMV IL3,CSF2,KITLG)1Eav/MloySzJ) mice were purchased from Jackson Laboratories and housed at the Center for Comparative Medicine and Surgery at Icahn School of Medicine at Mount Sinai. Female mice at 6â8âweeks of age were used and were assigned randomly to treatment and control groups. Numbers of mice per group were determined on the basis of historical observations. The smallest sample size estimated to provide more than 80% power to detect differences in leukaemic potential was used. Investigators were not blinded. At 1âday before transplantation, the mice were injected intraperitoneally with 30âmgâkgâ1 busulfan solution. Gene-edited iPS-cell-derived HSPCs from days 12â14 of haematopoietic differentiation, AML-iPS-cell-derived LSCs from days 14â16 of haematopoietic differentiation or cultured CB HSPCs were resuspended in StemPro-34 and injected through the tail vein using a 25G needle at 1âÃâ106 (for iPS-cell-derived cells) or 2â3âÃâ105 (for CB cells) per mouse in 100âµl. For Dox administration, mice were fed with Dox chow. For VEN administration, VEN was formulated for oral dosing in 60% phosal 50 propylene glycol, 30% polyethylene glycol 400 and 10% ethanol; 100âmgâkgâ1 was administered daily for 3âweeks. All mice were euthanized promptly once they showed signs of illness, according to Institutional Animal Care and Use Committee guidelines. Bone marrow was collected from the femurs and tibia. Bone marrow and spleen cells were haemolysed with ACK lysis buffer. Human engraftment was assessed by flow cytometric evaluation using hCD45-APC (clone HI30, BD PharMingen) and mCD45-PE-Cy7 (clone 30-F11, BD Biosciences) antibodies. Human cells were isolated using magnetic activated cell sorting (MACS), using CD45 microBeads (catalogue no. 130-045-801, Miltenyi Biotec) or mouse cell depletion kit (catalogue no. 130-104-694, Miltenyi Biotec), and cryopreserved for subsequent scRNA-seq analyses. For secondary transplantation, cells were obtained from the bone marrow of a primary NSGS recipient on weekâ6 post-transplantation (endpoint due to lethal disease). Following mouse cell depletion by means of MACS, 2âÃâ105 cells were injected intravenously in a secondary NSGS mouse, which was euthanized 7âweeks later.
Bulk RNA-seq
Three independent transductions of iPS-HSPCs for each of the four groups (Râ+âSA, Râ+âCtrl, SAâ+âR, SAâ+âCtrl; Fig. 2d) were performed. CD34+CD45+ HSPCs were obtained after MACS-sorting of CD45+ cells on a day of differentiation when all cells are CD34+ to obtain double positive CD34+CD45+ HSPCs using the MACS cell separation microbeads and reagents (Miltenyi Biotec). A total of 200,000 sorted cells were used for RNA extraction with the RNeasy mini kit (Qiagen) and 50,000 cells were used for ATAC-seq. PolyA-tailed mRNA was selected with beads from 1âμg total RNA using the NEBNext Poly(A) mRNA Magnetic Isolation Module (New England Biolabs). cDNAs were generated using random hexamers and ligated to barcoded Illumina adaptors with the NEXTflex Rapid Directional RNA-seq Library Prep Kit (Bioo Scientific); 75-nucleotide-long single-end reads were sequenced in a NextSeq-500 (Illumina).
ATAC-seq
A total of 50,000 MACS-sorted CD34+CD45+ cells from the same iPS-HSPC samples used for RNA-seq were processed as follows: nuclei were isolated by lysing with 50âµl of ATAC lysis buffer (10âmM Tris pHâ7.4, 10âmM NaCl, 3âmM MgCl2, 0.1% NP40, 0.1% Tween-20 and 0.01% Digitonin) and washing with 1âml of ATAC wash buffer (10âmM Tris pHâ7.4, 10âmM NaCl, 3âmM MgCl2, 0.1% Tween-20). Cell lysates were spun to obtain nuclear pellets, which were subjected to transposase reaction using the Illumina Nextera DNA Sample Preparation Kit according to the manufacturerâs instructions. The final libraries were quantified using the Agilent BioAnalyzer; 75-nucleotide-long paired-end reads were sequenced in a NextSeq-500 (Illumina).
Bulk RNA-seq data processing and analysis
FastQC (v.0.11.8, RRID:SCR_014583) was used for quality control. Trim Galore! (v.0.6.6, RRID:SCR_011847) was used to trim the adapter sequences with a quality threshold of 20. The human reference genome GRCh38 and GENCODE release 36 was used as the transcriptome reference (RRID:SCR_014966). Alignment used STAR aligner (v.2.7.5b, RRID:SCR_004463). Gene-level read counts were obtained using Salmon (v.1.2.1, RRID:SCR_017036) for all libraries. Sample normalization was carried out using the median-ratios normalization method from DESeq2 R package (v.1.30.1, RRID:SCR_015687), and differential expression analysis used DESeq2. Genes with fewer than five reads in total across all samples were filtered out. A gene was considered differentially expressed if the BenjaminiâHochberg adjusted Pâvalue was less than 0.05 and the absolute log2FC was greater than 1. Heatmaps were prepared using pheatmap (v.1.0.12) with hierarchical clustering. Barplots were prepared with ggplot2 (v.3.4.3). Over-representation for âRAS-late genesâ was analysed using the clusterProfiler R package (v.3.16.0).
Bulk ATAC-seq data processing and analysis
FastQC (v.0.11.8, RRID:SCR_014583) was used for quality control. Trim Galore! (v.0.6.6, RRID:SCR_011847) was used to trim the adapter sequences with default parameters. For each individual sample, paired-end 75-base-pair reads were aligned to the human reference genome (GRCh38/GENCODE release 36, RRID:SCR_014966) using Bowtie2 (v.2.1.0, RRID:SCR_016368) with default parameters and âX 2000. Reads were sorted using SAMtools (v1.11, RRID:SCR_002105), and mitochondrial and pseudo-chromosomal alignments were removed. Picard (v2.2.4, RRID:SCR_006525) was used to remove duplicates (Picard Toolkit 2019). To generate a universe of regions, all samples were merged using SAMtools merge function, followed by peaks calling using MACS (v.2.1.0, RRID:SCR_013291) with parameters ânomodel ânolambda âslocal 10000. Reads for each sample at the universe of regions were quantified using BedTools multicov with the corresponding filtered bam files (v.2.29.2, RRID:SCR_006646). Sample normalization was carried out using the median-ratios normalization method from DESeq2 R package (v.1.30.1, RRID:SCR_015687). Regions with fewer than 750 normalized reads in total across all samples were filtered out and differential peak analysis was carried out using DESeq2 (adjusted Pâvalueâ<â0.05 and absolute log2FCââ¥â2. Coverage tracks (Bigwig files) were generated from filtered BAM files for individual replicates using deepTools (v.3.2.1, RRID:SCR_016366) bamCoverage with parameters ânormalizeUsing RPKM âbinsize 1.
Cell-type-specific regulatory elements were obtained from ref. 29 The liftOver function from rtracklayer package (v.1.60.1) was used to convert hg19 coordinates to hg38. Distal elements specific to cell types of interest were plotted in heatmap format using deepTools (v.3.2.1) computeMatrix and plotHeatmap functions. Transcription factor motifs were analysed with the Homer (v.4.10) findMotifsGenome function. Data were visualized with ggplot2 (v.3.4.3) and dcCompareCurves function from deepStats (v.0.4). The CI threshold for bootstraps was set to 0.95.
Gene set enrichment analysis
GSEA was carried out on all 6,495 C2 curated gene sets from the Molecular Signatures Database (MSigDB, http://www.broadinstitute.org/msigdb) using the âfgseaâ R package (v.1.22 RRID:SCR_020938). Genes were ranked on the basis of log2FC multiplied by âlog10FDR (false discovery rate). GSEA Pâvalues were adjusted to control for FDR using the BenjaminiâHochberg method. Gene sets with FDRâ<â0.05 were considered to show significant enrichment.
GSEA was also applied to gene sets derived from ref. 14, corresponding to the populations LSPC-Quiescent, LSPC-Primed, LSPC-Cycle, GMP-like, ProMono-like, Mono-like and cDC-like.
scRNA-seq
Chromium 10x Genomics 3â² protocol (v.3.0) was used for scRNA-seq in cells from MACS-sorted iPS-cell-derived xenografts and on FACS-sorted CB HSPCs.
scRNA-seq data quality control and preprocessing
The FASTQ files were aligned, filtered, barcoded and unique molecular identifier (UMI) counted using CellRanger Chromium Single Cell RNA-seq by 10x Genomics (v.7.1.0 or v.5.0.1), with GRCh38 database (v.2020-A) as the human genome reference. Each dataset was filtered to retain cells with at least 1,000 UMIs, at least 1,000 genes expressed and less than 15% of the reads mapping to the mitochondrial genome. UMI counts were then normalized so that each cell had a total of 10,000 UMIs across all genes, and these normalized counts were log-transformed with a pseudocount of 1 using the âLogNormalizeâ function in the Seurat package. The top 2,000 most highly variable genes were identified using the âvstâ selection method of âFindVariableFeaturesâ function and counts were scaled using the âScaleDataâ function. Datasets were processed using the Seurat package (v.4.0.3)54.
scRNA-seq data dimensionality reduction and integration
Principal component analysis was carried out using the top 2,000 highly variable features (âRunPCAâ function) and the top 30 principal components were used in the downstream analysis. Diffusion maps were generated as implemented in the destiny (v.3.4.0) R package55 with default parameters and using 10,000 subsampled cells from each integrated dataset. Datasets for each patient were integrated separately by using the âRunHarmonyâ function in the harmony package (v.0.1.0). K-nearest neighbour graphs were obtained by using the âFindNeighborsâ function, whereas the UMAPs were obtained by the âRunUMAPâ function56. The Louvain algorithm was used to cluster cells on the basis of expression similarity. Cell density estimations were performed using the stat_density_2d function of the ggplot2 (v.3.3.5) package.
scRNA-seq data cell type annotation
Differential markers for each cluster were identified using the Wilcox test (âFindAllMarkersâ function) with adjusted Pâvalueâ<â0.01, absolute log2FCâ>â0.25 and greater than 10% of cells expressing the gene in both comparison groups using 1,000 random cells to represent each cluster. The top upregulated genes and curated genes from the literature were used to assign cell types to the clusters. Metaclusters were obtained by merging the manually annotated cell types into groups. Cell type frequencies between samples were compared using logistic regression (GLM R function, stats package v.4.3.1).
Genotyping of transcriptomes
A bone marrow mononuclear cell sample was obtained from a patient with AML with written informed consent under a protocol approved by a local Institutional Review Board at Memorial Sloan-Kettering Cancer Center. The sample was FACS-sorted to deplete lymphoid cells and enrich blast populations (DAPIâCD45+CD3âCD20âCD19âCD34+CD117+) and processed using the 10x Genomics 5â² V1 Gene Expression protocol. The FASTQ files were aligned and the cell-by-gene count matrix was generated with CellRanger v.5.0.1 with GRCh38 as the human genome reference using default parameters. The data were filtered to retain cells with less than 20% of reads mapping to the mitochondrial genome, at least 200 genes detected, and at least 4,000 UMIs. Downstream analyses were performed using Seurat v.4 in R 4.0.
Full length cDNA from the 10x library preparation was set aside for GoT. Amplicon libraries for NRAS G12D were generated and sequenced on an Illumina MiniSeq 300 cycle mid-output kit in 147:8:16:147 configuration. Resulting FASTQ files were processed as described above for the gene expression library. Mutation status of single cells was determined by performing a pileup of both gene expression and GoT data at the NRASG12 genomic location. Cells were assigned as NRAS MT if one or more mutant UMIs was detected, and as WT if two or more UMIs with only WT alleles were detected. This increased stringency for WT cells was used to account for allelic dropout. The genotyping efficiency was 8.3%, with 576 cells genotyped as NRAS-MT and 423 as NRAS-WT.
Association of SAR mutations with phenotypic AML features in a patient cohort
Bulk RNA-seq data from 599 adult patients with AML from the Alliance Cohort33, for whom FAB subtype clinical annotations and bulk targeted DNA sequencing data were also available, were used to analyse cell type composition. Cell type fractions were determined using cell-type-specific gene expression profiles derived from a scRNA-seq AML dataset (GSE230559) using dampened weighted least squares57.
Clinical trial
The analysis presented in this manuscript included patients with AML who received frontline therapy with DEC and VEN on a prospective clinical trial at the University of Texas MD Anderson Cancer Center, Houston, TX (NCT03404193). Patients were included if they were 60âyears old or older, or unfit to receive intensive chemotherapy. Patients with European LeukaemiaNet (ELN) favorable risk cytogenetics and previous BCL2 inhibitor exposure were excluded. DEC was dosed at 20âmgâmâ2 for 10âdays for induction, followed by 5âdays after achievement of a response. VEN was dosed at 400âmg daily or equivalent in conjunction with azole antifungals. Endpoints and outcomes were assessed per the ELN 2017 or ELN 2022 (extended cohort) guidelines58. The full protocol has been described previously19. Monocytic differentiation was determined by flow cytometric assessment, esterase positivity, myelomonocytic (FAB M4) or monoblastic/monocytic (FAB M5) morphology35. Measurable residual disease was assessed using multiparametric flow cytometry with sensitivity of 0.1% (ref. 59). All studies were conducted with informed consent in accordance with Declaration of Helsinki ethical guidelines.
VEN treatment and viability assay
iPS-cell-derived HSPCs, iPS-cell-derived monocytes and CB-derived HSPCs were plated on 96-well tissue culture-treated clear flat-bottom plates (Corning, catalogue nos. 3903 or 3603, respectively) at a density of 20,000 per well. VEN was purchased from Selleckchem and dissolved in DMSO for stock solutions at a concentration of 10âmM and subsequently diluted in StemPro medium and added to a total volume of 100âμl of medium per well at a final concentration of 6âμM in triplicate wells. RASi was obtained from Chemed (catalogue no. C-1418) and was added to the cultures at a final concentration of 100ânM in triplicate wells. After 3âdays, cell viability was measured using the CellTiter-Glo Luminescent Cell Viability Assay (Promega, catalogue no. G7570) per the manufacturerâs suggested conditions. Per cent viability at each compound concentration was calculated as: (Signal)/(DMSO control)âÃâ100. Half-maximal inhibitory concentration value calculations and generation of half-maximal inhibitory concentration curves were performed using the Prism v.8 software (Graphpad, RRID:SCR_002798).
Western blotting
A total of 1â2âÃâ105 iPS-cell- or CB-derived HSPCs were lysed with SDS sample buffer (ThermoFisher Scientific) supplemented with protease inhibitor and phosphatase inhibitor. Protein concentration was determined by bicinchoninic acid assay (Pierce Biotechnology Inc.) and 1â2âμg of protein from each extract was diluted in Laemmli SDS sample buffer, resolved by electrophoresis on Bolt 10% Bis-Tris precast gels (Invitrogen) and blotted on nitrocellulose membranes. The membranes were blocked with PVDF blocking reagent (TOYOBO) and incubated with primary antibodies: P-p44/42 MAPK (ERK1/2, clone D13.14.4E, catalogue no. 4370S, Cell Signaling Technologies, 1:5,000 dilution), p44/42 MAPK (ERK1/2, clone L34F12, catalogue no. 4696S, Cell Signaling Technologies, 1:5,000 dilution,), BCL2 (clone 124, catalogue no. M0887, DAKO, 1:5,000 dilution), MCL1 (clone D35A5, catalogue no. 5453S, Cell Signaling Technologies, 1:1,000 dilution), BCL-xL (clone 54H6, catalogue no. 2764S, Cell Signaling Technologies,1:5,000 dilution), β-actin (clone 13E5, catalogue no. 5125S, Cell Signaling Technologies, 1:10,000 dilution). After washing, blots were incubated with horseradish peroxidase-conjugated secondary antibody and developed using Western Blotting Detection Reagent (Western HRP Substrate, Millipore). For each antibody, independent blots were used from the same cell lysate with identical loading conditions.
Statistical analysis
Statistical analysis used GraphPad Prism software. Pairwise comparisons between different groups were performed using a two-sided unpaired unequal variance t-test (Stats R package v.4.3.1), unless stated otherwise. For all analyses, Pâ<â0.05 was considered statistically significant. Investigators were not blinded to the different groups.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.