Sample preparation and exome sequencing
Exome sequencing was performed at the Regeneron Genetics Center using a custom automated sample preparation approach (Supplementary Information 9). Samples from MAYO-CLINIC and part of the CNCD (n = 25,580) cohorts were captured with Twist Comprehensive Exome probes and the remaining cohorts were captured with IDT xGen v1 and sequenced using the Illumina HiSeq 2500-v4 or Illumina NovaSeq instrument, with 75-bp paired-end reads and two index reads. For the MAYO-CLINIC cohort sequenced with Twist, probes also included the Twist Diversity SNP panel, for which multipoint refinement was conducted using GLIMPSE before further genotype QC and imputation61.
Study populations
This study included 1,020,833 participants from seven cohorts: 469,662 participants from the UKB, 173,585 participants from the GHS cohort, 140,996 participants from the MCPS cohort, 116,345 participants from the Mayo-Clinic Biobank, 44,970 participants from the CNCD cohort, 44,027 participants from the University of Pennsylvania Penn Medicine Biobank, 31,248 participants from the Mount Sinai cohort (Supplementary Table 10 and Supplementary Information 1).
Genetic relatedness analysis
To estimate the portion of identical-by-descent (IBD) genomic regions shared between pairs of individuals in each study, we first obtained a set of high-equality common SNPs from the exome variant set by excluding SNPs with minor allele frequency < 10% and genotype missingness >5% and all indels. Furthermore, variants with abnormal heterozygosity rates based on the expected (exp) versus observed (obs) heterozygosity calculations based on empirically determined cutoffs (obs − exp > 0.01 or exp − obs > 0.1) were excluded from further analysis. The asymmetry in the cut-off values was selected to account for the Wahlund effect. IBD estimates were calculated among individuals within the same ancestral superclass that was determined in ancestry predictions as mentioned above using PLINK with a minimum PI_HAT cut-off of 0.1875 to capture out to second-degree relationships, which generates ancestry–version IBD estimates. A separate IBD estimation was calculated among all individuals using a minimum PI_HAT cut-off of 0.3 to identify the first-degree relationships among all samples to generate first-degree family networks, which are connected components of individuals (nodes) and first-degree relationships (edges). Each first-degree family network was analysed using the prePRIMUS pipeline built into PRIMUS62 using the default settings to produce improved IBD estimates for the relationships within each family network and capture close relationships that span more than one ancestral superclass that were not captured in the ancestry–version IBD estimates. The two versions of IBD estimates were combined in the form of a PLINK .genome file and subject to summary analysis after removing overly related samples with more than 100 close relatives (PI_HAT > 0.1875) or 25,000 relatives (PI_HAT > 0.08). All of the samples in the predicted first- and second-degree relationships were removed to generate the maximum unrelated data set for further analysis.
Ancestry assignment
We used array data released by the UKB study to determine continental ancestry super-groups (AFR, AMR, EAS, EUR and SAS) by projecting each sample onto reference PCs calculated from the HapMap3 reference panel. In brief, we merged our samples with HapMap3 samples and retained only SNPs in common between the two datasets. We further excluded SNPs with minor allele frequency < 10%, genotype missingness > 5% or Hardy–Weinberg equilibrium test P < 1 × 10−5. We calculated PCs for the HapMap3 samples and projected each of our samples onto those PCs. To assign a continental ancestry group to each non-HapMap3 sample, we trained a kernel density estimator (KDE) using the HapMap3 PCs and used the KDEs to calculate the likelihood of a given sample belonging to each of the five continental ancestry groups. When the likelihood for a given ancestry group was greater than 0.3, the sample was assigned to that ancestry group. When two ancestry groups had a likelihood of greater than 0.3, we arbitrarily assigned AFR over EUR, AMR over EUR, AMR over EAS, SAS over EUR, and AMR over AFR. Samples were excluded from analysis if no ancestry likelihoods were greater than 0.3, or if more than three ancestry likelihoods were greater than 0.3.
These sample analysed in this study include 763,174 participants of EUR ancestry, 149,124 participants of AMR ancestry, 57,196 participants of SAS ancestry, 41,405 participants of AFR ancestry and 5,313 participants of EAS ancestry (Supplementary Table 10). Within the UKB cohort, repeats called from WGS data were available for 465,021 samples, of which 441,379 were of EUR ancestry, 848 were of AMR ancestry, 10,085 of SAS ancestry, 9,173 of AFR ancestry and 2,287 of EAS ancestry.
Repeat expansion genotyping and QC
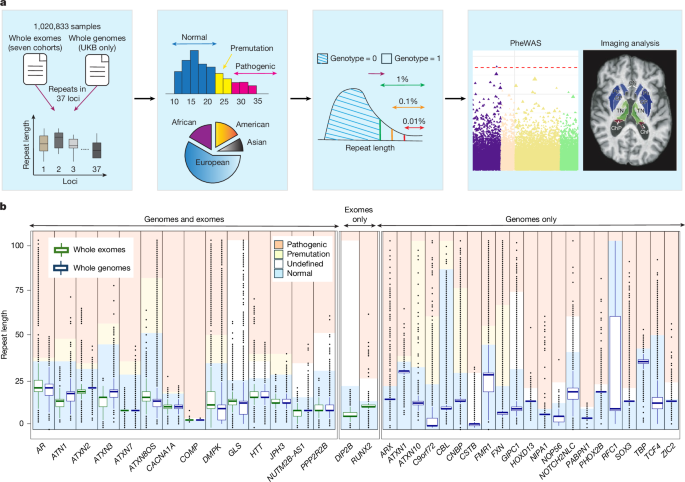
Repeats were called from whole exomes using GangSTR28 (v.2.5.0) from the CRAM files obtained by aligning the exome sequencing reads to the GRCh38 human reference genome. We used the list of 832,380 (version 13) repeat loci provided by GangSTR to curate a list of 7,046 loci that either overlapped with the exome capture probes or are one of the 34 loci associated with neurological disorders. We then ran GangSTR for each sample using the –targeted and –nonuniform options and provided insert size metrics as inputs. Both methods were run using DNANexus applets built to process samples in batches of 50 using AWS instances with dual core CPUs, 8 GB memory and 75 GB of storage. We curated a high-quality WES call set by identifying calls that either had quality score (Q score reported by GangSTR) of at least 95% or that met the following conditions, (1) be supported by read(s) that enclose the expansion; (2) read depth ≥ 10; (3) Q score ≥ 5%, and ($) maximum-likelihood (ML) estimate within 95% confidence interval (CI). That is, when the point estimate for ML falls outside the 95% CI, the authors of GangSTR deem such predictions to be unreliable and recommend discarding them. For the WGS calls, we used the ‘FILTER’ column provided by ExpansionHunter to remove calls with the ‘LowDepth’ flag and considered only calls with the ‘PASS’ flag for subsequent analyses. Moreover, we applied two filters at the locus level on both generated GangSTR calls and the obtained ExpansionHunter WGS calls (Table 1): loci were excluded (1) if after applying QC, they had fewer than 20% remaining samples; (2) or in cases in which the repeat distribution fell outside of the expected normal range based on literature thresholds. To identify the latter, we calculated for each locus the upper whisker of the repeat length distribution (that is, third quartile + [1.5 × interquartile range]) and excluded loci where the upper whisker fell over the expected pathogenic threshold. These QC criteria identified 16 and 35 loci with reliable distributions from WES and WGS, respectively, 14 of which overlapped between the two sources (Table 1).
STRs genotyped in UKB 500k Whole Genome data using ExpansionHunter (v.4)29 on the DRAGEN (v.3.7.8) platform were made available as part of the UK Biobank 500k Whole Genome Sequencing data release in November 2023. These STR genotypes were downloaded from the UKB Research Analysis Platform63 and subsequently used to estimate carrier frequencies and identify phenotypic associations. Spearman correlation between the repeat lengths of the longer allele estimated from WES and WGS samples for the 14 loci was done using the base R function cor.test64 (Supplementary Table 11).
As the repeat genotypes used in our study were generated from seven different cohorts, we assessed batch effects for three loci we report heavily in the manuscript (CACNA1A, DMPK and HTT). Specifically, we compared the entire repeat-length distribution across all cohorts (Supplementary Figs. 80–82) for these loci as well as the frequency of pathogenic carriers between cohorts and found evidence for batch effects likely arising from the variations in sequencing platform, capture kits and cohort ascertainment (Supplementary Information 10 and Supplementary Table 12).
Statistical analysis
Association analyses between repeat length genotypes and binary phenotypes were performed for the three largest cohorts with ICD-10 based diagnoses (GHS, UKB and Mayo-Clinic) using REGENIE (v.3.2+), which adjusts for relatedness and population structure65. We used the following covariates: age, age2, sex, age-by-sex, and age2-by-sex, and the first ten common-variant derived genetic PCs (calculated separately for each cohort). Five genotypes for each of the 37 loci that passed QC were constructed by binarizing the repeat length of the longer allele based on (1) premutation and pathogenic repeat length cut-offs from literature; and (2) the lengths that corresponded to the 1, 0.1, and 0.01 percentiles obtained from the repeat-length distribution of each locus. The percentile cut-offs for each locus were calculated independently within each of the three cohorts (Supplementary Table 7). Repeats on the longer allele that were expanded beyond these thresholds were encoded as 1, and 0 otherwise. The encoded genotypes were then converted to the PLINK 1.9 format (.bed, .bim and .fam files) using the R package named genio (v.1.1.2)66 and subsequently to PLINK 2.0 format (.pgen, .psam and .pvar) using the –make-pgen command via PLINK (v.2.0)67. For the phenome-wide association analysis, diagnoses spanning thousands of ICD-10 traits were considered. Specifically, we analysed the associations between repeat length genotypes in 37 loci and 6,140 traits from GHS, 5,657 traits from the UKB and 5,318 from Mayo-Clinic. Subsequent meta-analysis of these associations were run using the fixed-effect inverse-variance-weighted approach using METAL68 for the 3,890 traits found in all three cohorts as well as the 1,663 traits found in two of the three cohorts. The study-wide statistical significance threshold was determined to be 1.7 × 10−7 based on the 7,671 phenotypes and 37 loci considered for the PheWAS (significance threshold = 0.05/(7,671 × 37)). In addition to this threshold, to ensure the quality of associations, we considered only the subset of associations supported by at least ten repeat carriers who are cases for the phenotype.
For the analysis of enrichments/depletions within specific ancestry groups, we considered the statistical significance threshold to be 3.3 × 10−5 based on 37 loci, two tested thresholds (premutation and pathogenic), two data sources (WES and WGS) and 10 pairwise comparisons per locus (significance threshold = 0.05/(37 × 2 × 2 × 10)). In addition to this threshold, to ensure the quality of associations, we considered only the subset of tests supported by at least five repeat carriers.
Linear regression models
Association analysis for the brain imaging traits was run using linear regression with repeat status as a categorical predictor, rank inverse normal transformed brain volume as the outcome and the following covariates: the first 10 array PCs, first 20 exome rare variant PCs, UKB sequencing batch, sex, age, age2, age × sex, UKB assessment centre, position of brain within the scanner, and head size from SIENAX. An analogous analysis was performed with NfL levels in the plasma. To test for a linear association between repeat length and the outcome variable, we also ran the same analysis with min(0, L − T + 1) as a predictor, where L is the repeat length and T is the premutation repeat-length threshold. Quoted P values in the main text are for the coefficients on repeat status/length in these models.
Penetrance calculation
Penetrance was calculated as follows: the calculation considered the age at diagnosis for cases and the age at the last encounter for controls. In the case of a recorded death, we considered age at death as the most recent age. The samples were further divided into repeat expansion threshold groups based on their length. The calculation of the proportion of cases was cumulative, considering all cases and controls up until the tested age bin. The age-dependent penetrance was calculated as follows:
penetrance = (case carriers of repeat length>t threshold who are younger than age a)/(case and control carriers of repeat length > t threshold who are younger than age a).
For example, at a = 50 and a threshold of t = 35 repeats, the penetrance was calculated as
penetrance = (case carriers of repeat length > 35 who are younger than 50)/(case and control carriers of repeat length > 35 who are younger than 50). At the age of 80, this calculation included all samples younger than 80 years of age.
Brain imaging and plasma protein data from the UKB
T1-weighted brain images and plasma protein data were obtained from the UKB for 66,258 and 54,572 individuals, respectively. The raw structural T1-weighted brain images were processed through an internal pipeline similar to that presented previously69. In brief, the raw .zip files were accessed from the UKB Research Access Platform and converted to NIfTI using dcm2niix70. The image was then brain-extracted71, rigidly transformed to MNI152 space using FLIRT72 (FSL v.6.0.7.8) followed by a deformable registration to MNI152 space using FNIRT73. The rigid and deformable transformations are combined into a single transformation. The inverse of this transformation is then obtained, and is used transform the MNI152 atlas to the structural space, which is then used as a mask for a final brain extraction. Tissue-type segmentation was then performed using FSL’s FAST74, which also provides the intensity bias, which is used to obtain the final brain-extracted and intensity-bias-corrected T1-weighted image. This final image was then segmented using FastSurfer75 (v.2.3.0), and cortical thicknesses and other morphological traits were obtained using FreeSurfer (v.7.3.2). Cerebellum volumes were obtained from CerebNet76.
Samples with repeat information for HTT (n = 62,963) were split into four groups based on repeat length: normal (<27 repeats), intermediate (27–35; n = 4,049), premutation (36–39; n = 122) and pathogenic (\(\ge \)40; n = 9). Similarly, individuals with repeat information for CACNA1A (n = 63,960) were split into two groups: normal (<19) and premutation + pathogenic (≥19; n = 11).
HTT PCR validations
The AmplideX PCR/CE HTT Kit (49657; Asuragen) was used to PCR amplify the HTT trinucleotide CAG fragment starting from 2 μl of purified genomic DNA. The samples were prepared for the PCR with a master mix from Asuragen containing HTT PCR Mix (5.0 μl), HTT forward and reverse primer mix (3.0 μl). Tubes of master mix were vortex-mixed, centrifuged and transferred into a thermal cycler. Thermal cycling was performed on Eppendorf Nexus Gradient Mastercycler (Eppendorf) using the following cycle: 95 °C for 5 min, 10 cycles of 97 °C for 35 s, 64 °C for 35 s, 68 °C for 4 min, 18 cycles of 97 °C for 35 s, 64 °C for 35 s, 68 °C for 4 min plus 20 s per cycle, and final extension at 72 °C for 10 min, 4 °C hold. Then, 2 μl of PCR product was mixed with 11 μl Hi-Di Formamide (Applied Biosystems) and 2 μl ROX 1000TM Size Ladder (Asuragen). The samples prepared were then denatured at 95 °C for 2 min and cooled at 4 °C and held until ready for analysis by capillary electrophoresis (3730xL Genetic Analyzer, Applied Biosystems). The FAM-labelled amplicons were detected using the following fragment analysis protocol: 50 cm capillary, 2.5 kV, 20 s injection and 15 kV run for 4,800 s. Data analysis and interpretation was conducted using the fragment analysis software GeneMapper v.6 and an Excel-based analysis tool, AmplideX PCR/CE HTT Macro (Asuragen). The AmplideX PCR/CE HTT Kit Macro can determine size and mobility correction, and repeat size was determined using a linear fit adjustment of the ROX ladder size peaks to the PCR/CE control sample alleles. Alleles are reported as integer CAG repeats. The largest allele size determines genotype category: normal, intermediate, reduced penetrance or expanded. The alleles up to 200 CAG repeats are reported; the alleles with >200 repeats are identified as ‘>200’.
Validation of pathogenic expansions using Integrated Genome Viewer
Integrated Genome Viewer (v.2.19.4), a read stack viewer, was used to visually inspect the alignment and document evidence of expansion within the repeat region of HTT, CACNA1A and DMPK (Supplementary Information 6, Supplementary Table 13 and Supplementary Figs. 83–85).
Validation of interruptions using REViewer
REViewer (v.0.2.7), a tool for visualizing alignment of reads, was used to validate interruptions identified in HTT and ATXN1 using Expansionhunter39. Interruptions were called from both WES and WGS samples from the UKB. Comparison of interruptions in 419,186 samples that passed QC in both WES and WGS UKB data showed interruptions called from WES data to be more reliable than WGS due to higher read depth in coding regions (Supplementary Information 8 and Supplementary Figs. 86–88). Specifically, validations showed that >93% of interruptions in HTT and ATXN1 called by ExpansionHunter from WES data were accurate (Supplementary Information 7 and Supplementary Table 14).
Ethical compliance
Ethical approval for the UKB was previously obtained from the North West Centre for Research Ethics Committee (11/ NW/0382). The work described here was approved by UKB under application number 26041. Approval for GHS MyCode analyses was provided by the Geisinger Health System Institutional Review Board under project number 2006-0258. Informed consent was obtained for all of the study participants. Appropriate consent for the University of Pennsylvania Penn Medicine BioBank was obtained from each participant regarding storage of biological specimens, genetic sequencing and genotyping, and access to all available EHR data. This study was approved by the Institutional Review Board of the University of Pennsylvania and complied with the principles set out in the Declaration of Helsinki. All individuals participating in the Mayo-RGC Project Generation provided informed consent for use of specimens and data in genetic and health research and ethical approval for Project Generation was provided by the Mayo Clinic IRB (09-007763). All research performed in this study used de-identified data (without any Protected Health Information data) with no possibility of reidentifying any of the participants. For participants in the MCPS cohort, approval for the study was given by the Mexican Ministry of Health, the Mexican National Council of Science and Technology (0595 P-M) and the Central Oxford Research Ethics Committee (C99.260) and the Ethics and Research commissions from the Medicine Faculty at the National Autonomous University of Mexico (UNAM) (FMED/CI/SPLR/067/2015). All of the study participants provided written informed consent. The study participants were recruited from the BioMe Biobank Program of the Charles Bronfman Institute for Personalized Medicine at Mount Sinai Medical Center from 2007 onward. The BioMe Biobank Program (Institutional Review Board 07-0529) operates under a Mount Sinai Institutional Review Board-approved research protocol. All of the study participants provided written informed consent.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.