Animals
For neuroanatomical tracing experiments, most of the animals used were C57BL/6J male mice aged 2 months (n = 95; Jackson Laboratories). Mice were housed in a temperature-controlled (21–22 °C), humidity-controlled (51%) and light-controlled (12 h–12 h light–dark cycle; lights on, 06:00; lights off, at 18:00) room. Food and water were given ad libitum. After arrival at the facility, the mice were allowed a minimum of 1 week to adapt to the housing environment before surgeries were performed. All of the experiments were conducted according to the standards set by the National Institutes of Health Guide for the Care and Use of Laboratory Animals and the institutional guidelines of the University of Southern California (USC) and University of California Los Angeles (UCLA).
Male MORF3 mice (n = 20; aged 2 months), were also used for neuroanatomical tract tracing. The MORF3 mouse line (C57BL/6-Gt(ROSA)26Sortm3(CAG-sfGFP*)Xwy/J), generated by X. W. Yang’s laboratory at UCLA25, is a Cre reporter mouse line that uses a mononucleotide repeat frameshift (MORF) as a translation switch for cell labelling in vivo. MORF3 mice express a Cre-dependent tandem ‘spaghetti monster’ fluorescent protein with 20 V5 epitopes (smFP-V5) preceded by a polycytosine repeat (C22) MORF switch under the control of a CAG promoter. Cre recombination combined with a spontaneous frameshift results in sparse and stochastic labelling of neural cells.
Two other transgenic mouse lines, including VGLUT2 Cre (B6J.129S6(FVB)-Slc17a6tm2(cre)Lowl/MwarJ) and CRH-Cre (B6(Cg)-Crhtm1(cre)Zjh/J) were purchased from JAX and breeding colonies for these mouse lines were established at UCLA. CRH-Cre mice were crossed with Ai14 mice to generate CRH-Cre Ai14 mice for some of the experiments. A total of 5 male VGLUT2 Cre, 5 CRH-Cre and 4 CRH-Cre Ai14 mice aged 2 months were used for the neuroanatomic tracing experiments. Animal information for non-tracing studies is included in their corresponding subsections below.
Sample sizes for each of the tracing and morphology experiments were chosen based on previous experience. Randomization was not performed for the neural tracing experiments as animals were not assigned to different groups. Randomization is necessary for reducing bias and controlling variability. Instead, the data were validated in different ways (see the ‘Data reproducibility’ section). Randomization was also not necessary for morphology experiments. Owing to the characteristics of MORF3, we achieve random labelling of neurons and then reconstruct the traceable neurons. For neuroanatomic tracing and morphology experiments, blinding also was not necessary because the animals were not assigned to different groups.
Ethics statement
We are committed to promoting ethical research practices and ensuring the welfare of all animals involved in our studies. All procedures adhered to regulatory standards as delineated in the National Institutes of Health Guide for the Care and Use of Laboratory Animals, as well as institutional guidelines set forth by the Institutional Animal Care and Use Committees at the University of Southern California (USC) and the University of California, Los Angeles (UCLA).
Tracers
Phaseolus vulgaris leucoagglutinin (PHAL; 2.5%; Vector Laboratories) was used as the main chemical anterograde tracer. Chemical retrograde tracers included cholera toxin subunit b (CTB Alexa Fluor 647 conjugate, 0.25%; Invitrogen) and FG (1%; Fluorochrome). FG was also injected through the tail vein (2%, 30 μl) to label neuroendocrine cells. Each anterograde and retrograde tracer has distinct characteristics and exhibits varied neurotropism that can meaningfully affect connectivity results. For example, anterograde AAVs label fibres of passage, while PHAL does not. These differences underscore the importance of data validation. In this work, the connectivity data were validated in multiple ways to ensure the reliability of the results. Retrograde tracers were placed in regions of anterograde terminations, while anterograde tracers were placed in regions of retrogradely labelled cells. For example, Extended Data Fig. 10 shows retrograde tracer injection in the DPs and DPd that reveal their distinct brain-wide inputs (brain-wide ROIs to the DPs or DPd). To validate these results, Extended Data Fig. 11 shows anterograde tracer injections delivered to those brain-wide ROIs to validate their projections to either the DPs or DPd. Furthermore, we have previously demonstrated that PHAL and AAV produce similar brain-wide anterograde labelling patterns21 (Supplementary Fig. 6).
AAVretro-hSyn-Cre-WPRE (AAV retro Cre; 1.6 × 1013 genome copies (GC) per ml; Addgene, 105553), AAV1-Syn-Flex-GCaMP6f-WPRE-SV40, AAV1-hSyn-SIO-stGtACR2-FusionRed and AAV-Ef1a-mCherry-IRES-Cre (retrograde)53 were produced by Addgene.
Viral anterograde tracers AAV encoding enhanced green fluorescent protein (AAV-GFP; AAV2/1.hSynapsin.EGFP.WPRE.bGH) and tdTomato (AAV1.CAG.tdtomato.WPRE.SV40) and AAV1-hSyn-Cre-WPRE-hGh (anterograde trans-synaptic) were packaged by UPenn Vector Core. AAV5-Ef1a-DIO-hChR2(H134R)-eYFP, AAV9-CAG-FLEX-GFP (Cre-dependent channelrhodopsin expressing AAV) was produced by the University of North Carolina vector core facility.
For experiments investigating the synaptic contacts of DP axon terminals onto PVH CRH neurons, an AAV-synaptophysin-GFP injection was made into the DP of CRH-Cre Ai14 male mice (n = 4, aged 2 months). A 1:1 mixture of AAV Cre and AAV synaptophysin-GFP was generated and a 100 nl pressure injection was made into the desired ROI. The viruses used for the mixture were AAV1-hSyn-FLEX-tdTomato-T2A-SypGFP-WPRE (Addgene, 51509, 1.3 × 1013 GC per ml) and AAV1-hSyn-Cre-WPRE (Addgene, 105553, 1.8 × 1013 GC per ml).
The viral vectors used for the genetically engineered rabies tracing system were produced by Wickersham Lab at MIT and included AAV2/9-CAG-FlexmKate-T2A-TVA, AAV2/9-CAG-Flex-mKate-T2A-N2c-G and Rbv-CVS-N2c-ΔG-GFP (the modified rabies virus). All viral vectors were aliquoted and stored at 80 °C until use. Importantly, rabies virus has been widely used as a tool for reliably mapping presynaptic inputs to a specific brain region or starter cell population54,55,56. While its tropism appears broad, it remains possible that specific cell types in different pathways may be over- or under-represented relative to the performance of other retrograde tracers and viruses (for example, AAVretro, CAV2, retrobeads and so on)57,58. Moreover, the potential for cytotoxicity within the starter cell population may provide an opportunity for rabies to non-specifically spread to nearby synapses rather than through those specifically contacting starter cells. To control for these possibilities, retrograde tracing results must be validated using complementary anterograde approaches, such as by tracing axonal connections from the upstream region, or by optogenetically stimulating upstream axons and recording synaptic responses in the target cell population. As such, the data that we present were validated in several different ways.
Stereotaxic surgeries for neuroanatomical tract tracing
Surgical and microscopy imaging procedures for neuroanatomical tract tracing experiments have been described previously28,32,59. Stereotaxic coordinates of targeted injection centres were determined through the ARA11 and empirically adjusted when needed. On the day of the experiment, mice were deeply anaesthetized and mounted into a Kopf stereotaxic apparatus where they were maintained under isoflurane gas anaesthesia (Datex-Ohmeda vaporizer). Before the surgery, mice were given one subcutaneous injection of Ketoprofen (4 mg per kg) and a protective ophthalmic ointment was applied to their eyes. For single anterograde tracer injection experiments (PHAL or AAV), tracers were iontophoretically delivered through glass micropipettes (inner tip diameter, 24–32 μm) using alternating 7 s on–7 s off pulsed positive electrical current (Stoelting, current source) for 10 min, and AAVs were delivered through the same method for 2 min (inner tip diameter, 8–12 μm). For anterograde/retrograde co-injection experiments (PHAL/CTB-647 and AAV/FG), tracer cocktail iontophoretic injections were made through glass micropipettes (inner tip diameter, 28–32 μm) for 10 min (PHAL/CTB-647) or 5 min (AAV/FG).
For multiple retrograde tracing experiments, at each injection site, 50 nl of the retrograde tracer was individually pressure-injected through glass micropipettes at a rate of 10 nl min−1 (Drummond Nanoject III). All injections were placed into the right hemisphere. After injections, incisions were sutured, and mice were returned to their home cages for recovery.
TRIO tracing (Cre-dependent TVA receptor mediated rabies tracing)
To reveal monosynaptic inputs to projection-defined neuronal populations in the DP (for example, DP neurons projecting to the ACA), we used a modified TRIO (tracing the relationship between input and output) strategy60. In brief, AAVretro-Cre was injected into a downstream projection target of DP (for example, ACA), and Cre-dependent TVA- and RG-expressing helper virus (AAV8-hSyn-FLEX-TVA-P2A-GFP-2A-oG) and mCherry-expressing G-deleted rabies virus (produced by the laboratory of I. Wickersham) were injected into the DP to label the DP projection neurons population (first order) and their brain-wide monosynaptic inputs (second order). The same strategy also was applied to trace brain structures that generate monosynaptic inputs to the DP-projecting neurons in the RE.
We used a modified rabies tracing strategy to identify monosynaptic inputs from the DPd to CRH-expressing neuroendocrine neurons in the PVH. In CRH-Cre transgenic mice, a Cre-dependent helper virus (AAV8-hSyn-FLEX-TVA-P2A-GFP-2A-oG) was injected into the PVH to drive expression of TVA and rabies glycoprotein (RG), followed by injection of mCherry-expressing G-deleted rabies virus (RbV-ΔG-mCherry). This approach selectively labels neurons providing direct monosynaptic input to CRH neurons. All viral vectors were produced by the Wickersham laboratory at MIT.
AAV1-Cre-based anterograde trans-synaptic tracing
This technique leverages the fact that when AAV1 is injected at sufficiently high concentration into a neuronal population, viral particles will travel down the axons and be released from the synaptic terminals where they can infect postsynaptic neurons. Detailed methodology was described previously61. In brief, anaesthetized mice were iontophoretically injected with Cre-dependent AAV-FLEX-RFP or GFP in the target structures (for example, the LHA), and pressure injected (20–80 nl) with AAV1-Cre in an upstream structure (for example, the DP). The AAV1-Cre is transported anterogradely down the axons and is released from the terminals, where it transfects postsynaptic cells that have been infected with high concentrations of Cre-dependent AAV-FLEX-RFP. The scant Cre expression is sufficient to unlock strong fluorophore expression in the downstream neurons, therefore revealing their axonal projections and terminals. After a 3-week post-operative recovery, the mice were anaesthetized with pentobarbital and perfused. The Cre injection site was verified by staining with mouse anti-Cre recombinase monoclonal primary antibody (see the ‘Tissue preparation and immunohistochemistry’ section for details). One caveat to using AAV1-Cre is that the virus can also travel retrogradely and should be used in situations in which the connection is known to be primarily unidirectional. In all of the experiments in which AAV1-Cre was used in the current paper, the connections were shown to be predominantly unidirectional (BLA/SUBv/AUDv to DP and DP to CEA/LHA/PAG; Supplementary Fig. 11). As an example, we injected AAV1-Cre into the DP and a Cre-dependent AAV into the LHA to show that LHA neurons that receive input from the DP project to the spinal cord (DP to LHA to spinal cord). If LHA projected back to the DP, we would possibly be showing that LHA neurons that project to DP also project to the spinal cord. However, we know that the DP to LHA connection is unidirectional (Supplementary Fig. 11b), which lends reasonable confidence to our conclusion of DP to LHA to spinal cord.
Details regarding how injection site locations were accurately determined are provided in the ‘Injection Site Analysis’ section of the Supplementary Information.
Data reproducibility
Reported connections underwent validation through at least one of the following methods. Injections targeting different regions were repeated to evaluate consistency of labels. The Cre-dependent anterograde and TVA receptor mediated rabies tracing also validated the connections. Furthermore, retrograde tracers were introduced into regions displaying anterograde terminal labelling to confirm anterograde connections, whereas anterograde tracers were administered into the sites of retrogradely-labelled projection cells to validate retrograde injection data. In some instances, channelrhodopsin assisted circuit mapping also validated connections.
Tissue preparation and immunohistochemistry
Animals were euthanized with an overdose injection of sodium pentobarbital (6 mg per kg) 7 days (chemical tracers) or 14 days (viral tracers) after surgeries. Each animal was transcardially perfused with approximately 50 ml of 0.9% NaCl followed by 50 ml of 4% paraformaldehyde solution (PFA; pH 9.5). The brains were post-fixed in 4% PFA for 24–48 h at 4 °C, after which they were embedded in 3% type I-B agarose (Sigma-Aldrich) before sectioning. Four series of coronal sections were sliced at a thickness of 50 µm using a Compresstome (VF-700, Precisionary Instruments) and prepared for processing. One of the four-section series was immunostained for imaging. The series that contained coronal level 53 of the Allen Reference Atlas was selected to maintain similar section-level distributions across experiments.
The following primary antibodies were used across the studies: rabbit anti-PHAL (1:1,000; Vector Laboratories, AS-2300), rabbit anti-fluorogold antibody (1:5,000; Millipore-Sigma, AB153-I), mouse anti-Cre recombinase (1:4,000; EMD Millipore, MAB3120), rat monoclonal FOS antibody (1:1,000; Synaptic Systems, 226017), mouse anti-oxytocin antibody (4G11, 1:5,000; Millipore-Sigma, MAB5296), rabbit anti-vasopressin antibody (1:5,000, Millipore-Sigma, AB1565), rabbit-anti somatostatin antibody (1:500, Millipore-Sigma, SAB4502861) and streptavidin for biocytin-labelled recorded neurons (1:1.000; Thermo Fisher Scientific, S21374).
In brief, sections were transferred to a blocking solution containing normal donkey serum (Vector Laboratories) and Triton X-100 (VWR) for 1 h. After three 5-min rinses, the sections were incubated in a KPBS solution comprising donkey serum, Triton X-100 and a primary antibody at the appropriate dilution (the concentration for each primary antibody is described above) for 48–72 h at 4 °C (Vector Laboratories, AS-2300). The sections were rinsed three times in KPBS and then soaked for 3 h in the secondary antibody solution, which contained donkey serum, Triton X-100 and the corresponding secondary antibody solution at the appropriate dilution (1:500 for anti-rabbit IgG conjugated with Alexa Fluor 488, 555 or 647; Invitrogen, A-21206 (488), A-31573 (647); anti-mouse IgG conjugated with Alexa Fluor 488 or 647: Life Technology: A-21202 (488) and A-31571 (647); anti-mouse IgG conjugated with Alexa Fluor 647: Jackson ImmunoResearch, 715-605-150). The sections were again rinsed with KPBS three times. All selected section series were counterstained with a fluorescent Nissl stain, NeuroTrace 435/455 (NT, 1:500, Invitrogen, N21479). The sections were then mounted and coverslipped using 65% glycerol.
RNAscope experiments
Deeply anaesthetized animals were perfused and brains were post-fixed in 4% PFA overnight followed by sequential immersion in 15% and 30% sucrose for cryoprotection. The brains were then sectioned (20 µm) and the RNAscope Multiplex Fluorescent v2 assay was performed according to the manual provided by Advanced Cell Diagnostics. In brief, slides containing tissue sections were post-fixed in the prechilled 4% PFA for 15 min at 4 °C, and then dehydrated in a series of ethanol solutions 50%, 70% and 100% for 5 min at room temperature. Hydrogen peroxide was applied to each section for 10 min at room temperature, and target retrieval was then performed at 99 °C for 5 min. Protease III was then added to the samples and incubated in a HybEZ oven for 30 min at 40 °C. The subsequent RNAscope assay was composed by multiple steps of hybridization (probe and AMP binding) and signal development (HRP channel binding), each followed by washing the slides in 1× wash buffer as described in the manufacturer’s protocol (Multiplex fluorescent v2 kit). Opal fluorophores (Akoya Biosciences) were diluted at 1:1,500 to visualize RNA signals. After the RNAscope assay, the sections were counterstained with DAPI (1:500, Thermo Fisher Scientific, D1306) and mounted. Images were acquired using a Dragonfly High Speed Confocal Microscope System (Andor). The following probes were used in this study: Cux2, Etv1, Tle4, SS, CRH, and TRH.
Image acquisition and 2D processing
All 50-µm sections were scanned as high-resolution virtual slide image (VSI) files using the Olympus VS120 high-throughput microscope. Post-acquisition processing methods, including image registration, tracer segmentation and quantification, are described in detail below. In brief, all images were flipped to ensure that the injection site and its corresponding ipsilateral labelling were in the correct right hemisphere. This was performed using the VS-Desktop software that accompanies the microscope. Images were then imported into Outspector—our custom software for image processing (see the ‘Outspector: our proprietary 2D image processing pipeline’ section of the Supplementary Methods) and processed on a high-performance computing cluster. A first-pass fully automatic registration was executed and subsequently its results were inspected and fine-tuned when needed. Tracer segmentation was performed on registered images with the default parameters. Segmentation results were also inspected, followed by interactive parameter adjustments if needed. Segmented pixel counts and cell counts in each brain region at all section levels were generated and used for analysis.
Outspector
Outspector is our informatics workflow specifically designed to reliably warp, reconstruct, annotate and analyse the labelled pathways in a high-throughput manner. Tissue sections from each analysed case were assigned and registered to standard templates of the corresponding ARA levels. All images shown in this Article are from the raw data of unwarped, unregistered VSI images. Threshold parameters were individually adjusted for each case and tracer. Adobe Photoshop was used to correct conspicuous artifacts in the threshold output files. Each colour channel was brightness/contrast adjusted to maximize labelling visibility (Nissl Neurotrace 435/455 is converted to bright field), and TIFF images are then converted to JPEG file format. The detailed calculations used for all Outspector processing steps and analysis are provided in the ‘Outspector: our proprietary 2D image processing pipeline’ section of the Supplementary Methods.
Quantifying colocalization
Using spots function on Imaris, thresholds were set to select most positive cells for each channel (405, 488, 561 and 642). After automatic detection, manual quality control was necessary. The spatial relationship between DAPI in 405 and RNA puncta for Fezf2 was used as a visual aid to correct false negatives and remove false-positive spots from the automatic detection. Each channel was quality controlled multiple times to ensure accuracy.
Tissue processing and imaging for 3D single-neuron labelling
To elucidate the finely detailed morphology of DP neurons, we injected with either AAV-retro Cre or AAV1-Cre into MORF3 mice. Then, 3–4 weeks after the injections, the mice were perfused. Immunostaining with the V5 antibody allows the visualization of the finely detailed neuronal morphology of these MORF3 neurons. This approach enabled us to image, reconstruct and analyse the cell-type-specific morphological features of DP cortical projection neurons. Following the same strategy, we also revealed sparsely labelled neurons with finely detailed dendritic morphology in the LHA.
The SHIELD clearing protocol was used for the 3D tissue processing62. Mice were transcardially perfused with cold saline and SHIELD perfusion solution. Brains were dissected, subjected to a SHIELD-based brain clearing method and immunostained for V5 (to reveal MORF3 with secondary antibody conjugated with Alexa Fluor 647) using eFLASH25,62,63. Brains were processed using the SmartClear and SmartLabel machines (LifeCavas SmartBatch). Whole brains were imaged using the Life Canvas SmartSPIM lightsheet microscope at ×15 (0.42 μm × 0.42μm × 0.42 μm). Three lasers (488, 555, 647 nm) were used, respectively, to image Syp–eGFP axonal terminals, tdTomato-positive cortical neurons and their axons and MORF3-positive postsynaptic neurons.
For acquiring images on thick sections, the whole brain was cut into 500 µm sections and were cleared in the SDS buffer at 37 °C for 72 h. The sections were then washed three times with KPBS and incubated in KPBS at 4 °C for 24 h. The sections were mounted and cover-slipped onto 25 × 75 × 1 mm glass slides with an index matching solution (100%, EasyIndex, LifeCanvas Technologies, EI-Z1001). The sections were imaged with a high-speed spinning-disk confocal microscope (Andor Dragonfly 202 Imaging System, Andor, Oxford Instruments Company, CR-DFLY-202-2540). ×10 magnification (NA 0.40, Olympus, UPLXAPO10X) was used to acquire an overview, after which ×30 magnification (Olympus, UPLSAPO30xSIR, NA 1.05, W.D.: 0.80 mm) at 1 µm z steps or ×60 magnification (Olympus, UPLASAPO60XS2, NA 1.30, W.D.: 0.30 mm) were used to acquire high-resolution images for fine-detailed dendritic morphology and putative synaptic connections. The sections were imaged with four excitation wavelengths: 405 nm (blue Nissl background), 488 nm (for AAV-GFP) and 647 nm (far red for MORF3) with respective emission detection wavelengths of 450 nm, 525 nm and 600 nm.
3D reconstructions, visualizations and analysis of neuronal morphology
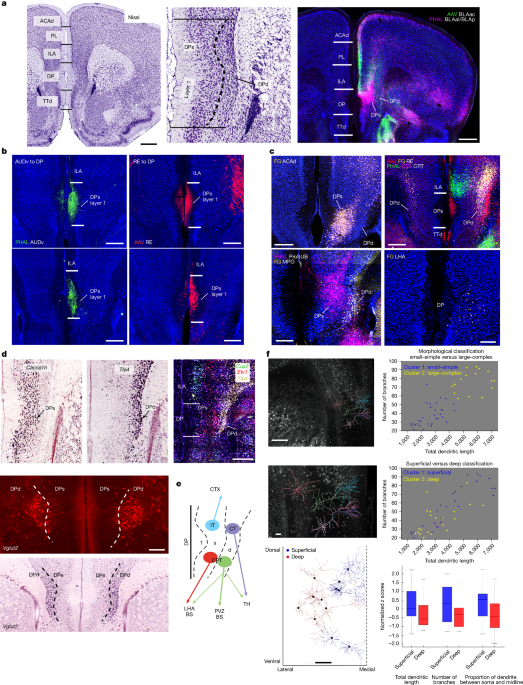
Manual reconstruction of neurons was performed using Vaa3D64,65 or Fast Neurite Tracer66, followed by morphological repairing, sorting and typing of the dendritic arbours. Reconstructions were finalized by loading them in Imaris and neuTube67 and finalized into the widely used swc file format68. In total, 47 reconstructed DP neurons were morphologically analysed in MATLAB, leveraging functions from the Trees Toolbox environment69. The 47 DP neurons were pooled from four MORF3 mice. Specifically, analysis scripts measured all neurons across a series of parameters including total dendritic wiring length, number of branches, average branch angle, average root angle, branch asymmetry, average branch order, average Strahler order and the distance from the midline. The coefficients of variances across the parameters were then measured, followed by classification using k-means clustering. The size and complexity of dendritic arbours, defined by total dendritic length and the total number of branches, emerged as the most varied. We then classified all of the neurons, first based on dendritic arbour size and complexity, followed by superficial versus deep-layer classification based on the distance of all cell bodies from the midline. Both classifications created two neuronal clusters each. The superficial versus deep-layer neurons were then compared for size, complexity and dendritic orientation. Similarly, the two morphological clusters (small-simple and large-complex) were compared for their distance from midline.
Statistical analyses
These comparisons were made using one-tailed nonparametric Wilcoxon signed rank tests. The P values from the tests were corrected using FDR correction to account for multiple comparisons.
In vitro patch-clamp slice recordings
In wild-type mice, AAV1-Cre (anterograde trans-synaptic Cre) was injected into the AUDv, while AAV-hSyn-ChR2-YFP (UPenn Vector Core) was injected into either the ENTI (n = 5; Fig. 5a) or PIR (n = 6 Fig. 5c) of different animals. In separate experiments, the AAV-hSyn-ChR2-YFP was injected into the DP of CRH-Cre Ai14 mice (n = 7; Fig. 4g). Sample sizes were chosen on the basis of previous experience. Randomization and blinding were not performed as the mice were not assigned to different groups.
Two weeks after channelrhodopsin and tracer injections, acute brain slices were prepared for electrophysiological recordings. After anaesthesia with isoflurane, the mouse was decapitated and the brain was rapidly removed and immersed in ice-cold high sucrose slice solution: 208 mM sucrose, 2.5 mM KCl, 1.25 mM NaH2PO4, 26 mM NaHCO3, 1.3 mM MgCl2, 8 mM MgSO4 and 10 mM glucose; saturated with 95% O2 and 5% CO2; pH 7.4. Brain slices (thickness, 350 μm) were cut in the coronal plane using a vibrating microtome (Leica, VT1000S). Slices were allowed to recover for at least 30 min in a submersion chamber filled with warmed (32–34 °C) ACSF containing 130 mM NaCl, 3 mM KCl, 1.25 mM NaH2PO4, 26 mM NaHCO3, 2 mM CaCl2, 2 mM MgCl2 and 10 mM glucose, oxygenated with 95% O2 and 5% CO2, pH 7.2–7.4, 290–310 mOsm, and then cooled gradually to room temperature until recording. The presence of RFP and ChR2 (YFP) labelling was determined with green and blue fluorescence excitation, respectively, in the slices before recording. Patch pipettes with about 3–4 MΩ impedance were used for whole-cell patch clamp recordings. Glass pipettes contained a Cs-methanesulfonate-based internal solution with the following salt concentrations: 130 mM Cs-methanesulfonate, 10 mM CsCl, 4 mM NaCl, 1 mM MgCl2, 5 mM mM 150 mM MgATP, 5 mM EGTA, 10 mM HEPES, 5 mM GTP, 10 mM phosphocreatine and 0.1 mM leupeptin (pH 7.2 with 151 CsOH, 270 mOsm) and biocytin (0.2%). For isolation and recording of monosynaptic responses to blue-light stimulation, tetrodotoxin (Na+ channel blocker, 1 μM) and 4-aminopyridine (K+ channel blocker, 100 μM) were added to the external solution. Signals were recorded from red-labelled neurons with a MultiClamp 700B amplifier (Molecular Devices) running pClamp software (v.10.5) under voltage clamp mode at a holding voltage of −70 mV for excitatory currents, filtered at 1 kHz and sampled at 10 kHz. Blue light (470 nm) stimulation was delivered using a 0.5 ms pulse at ~1 mW power for 5 trials, delivered through a LED (CoolLED). Signals were analysed using Clampfit (v.10.5). For responding neurons, peak responses were averaged across trials (technical replicate). The averaged value of each cell was then combined with the other cells for each experiment to calculate the mean, median, s.d. and so on (biological replicate). To demonstrate that light-evoked synaptic responses were indeed excitatory, the selective glutamate receptor antagonists CNQX (20 µM) for AMPA receptors and APV (50 µM) for NMDA receptors were added to the bathing solution.
After recording, the slices were fixed with PFA and then transferred to PBS for biocytin staining to reveal and validate the location of the recorded neurons. The biocytin staining also revealed the morphology of the recorded DP or PVH neurons (see the ‘Tissue preparation and immunohistochemistry’ section for details).
Statistical analyses
To compare the median peak amplitude of currents evoked by optogenetic stimulation of AUDv to DP neurons receiving either ENTl or PIR input, nonparametric Mann–Whitney U-tests were used and P value adjustments were not necessary.
Optogenetic stimulation and measurements of plasma CORT hormone
A total of 15 mice (male C57BL/6, aged 2 months) were randomly assigned to either the experimental or control group. Sample sizes for each of the experiments were chosen based on our previous experience and, based on the results, sufficient power was achieved to detect significant differences. Each of the animals received bilateral injections of AAV1-Cre into the SUBv and the BLAa, allowing Cre to anterogradely transport to and trans-synaptically label DP neurons. The animals also received a bilateral injection of Cre-dependent AAV-FLEX-ChR2 (30 nl) into the DP. Consequently, only DP neurons postsynaptic to the SUBv and BLAa express ChR2 and generate projections to the hypothalamic neuroendocrine zone, including the PVH. Optic fibres (200 μm core diameter, 0.37 numerical aperture; Inper) were implanted above the injection site at a depth of −1.6 mm ventral to the bregma skull surface and secured to the skull using super glue and dental cement.
The optogenetic stimulation was performed 7–8 weeks after the surgery. Before testing, the mice were individually habituated to the testing cage (new mouse cage) and optic fibres over 4 consecutive days, for 30 min daily. On the test day, the mice were habituated to the testing cage alone for 2 h and attached to optic fibres for 15 min before optogenetic stimulation. A 473 nm laser, pulsing at 20 Hz with 15 ms pulses, was applied for 30 s on and 30 s off cycles, for a total of 20 min. The light irradiance was around 4–5 mW mm−2 in the target region. After laser stimulation, the optic fibres were removed, and the mice remained undisturbed in the testing cage for 25 min before euthanasia. To minimize circadian rhythm effects on CORT levels, animals were tested between 1.5 to 5 h after lights turned off in the dark phase, with control and ChR2 groups counterbalanced during the test.
Serum collection and CORT assaying
After rapid cervical dislocation, terminal trunk blood was collected with heparinized microcapillary tubes (Thermo Fisher Scientific) and transferred into microfuge tubes, which were left on ice for at least 15–20 min. The blood samples were centrifuged at 2,500 rpm for 15 min at 4 °C. The supernatant (serum) was collected and transferred into new tubes and the serum samples were frozen at −80 °C. The samples were packed and shipped on dry ice to the Center for Research in Reproduction Ligand Assay and Analysis Core at the University of Virginia School of Medicine, where they were processed (in duplicate) using a radioimmunoassay kit for CORT measurements. The brains were sectioned at 50 μm thickness and Nissl stained to assess the location of the implanted optic fibres. The researcher assessing the accuracy of the bilateral implant location was blinded to the treatment condition of the animal. Only animals with accurate bilateral implants in the DP were included in the analysis.
Statistical analyses
To compare the CORT levels between the experimental and control groups, Student’s t-test and nonparametric Mann–Whitney U-tests were used. Owing to the two-group comparisons, P value adjustment was not necessary.
Social interaction experiments
Surgery
Six VGLUT2-Cre (for DP) and six VGLUT1-Cre (for ILA) male mice (aged 6–8 weeks) were anaesthetized using a 1–2% isoflurane–oxygen mixture and injected unilaterally with 350 nl (1 nl s−1) pAAV.Syn.Flex.GCaMP6f.WPRE.SV40 (Addgene, 100833-AAV1) into the DP. Then, 30 min after virus injection, a 6.1-mm-long (0.5 mm diameter) relay lens (Inscopix) was implanted over the DP: anteroposterior, +2.1 mm, mediolateral, 0.6 mm; dorsoventral, −3.2 mm. The lens was securely affixed to the skull using cyanoacrylate glue and dental cement and then covered with Kwik-Sil (WPI). Sample sizes for each of the experiments were chosen based on previous experience. Randomization was not necessary as VGLUT2-Cre animals received injection into the DP and the VGLUT1-Cre mice received injection in the ILA. Also, each of the animals underwent both social and non-social interaction behaviour tests (see below).
Three weeks later, the mice were anaesthetized as described previously and a UCLA V4 miniscope with an attached baseplate was positioned over the lens to visualize GCaMP6f expression. The focal plane was adjusted to optimize cell visibility, and the baseplate was secured in place with dental cement. A plastic cap was used to cover and protect the lens over the baseplate.
Behavioural tests
Mice with lens implantation in the DP or ILA were habituated to handling and the miniscope for 4–5 days. On the day of the open-field test, the miniscope was attached to the baseplate and the mice freely ran in an open arena (45 cm × 45 cm × 30 cm) for 20 min. On the day of same-sex social interaction, mice performed social interaction counterbalanced with object exploration in the same arena as the open-field test for 8 min per session. The next day, mice performed social interaction with opposite sex social targets for 8 min. Each social trial included an 8-min recording session, beginning with a 1-min period of the subject mouse alone, followed by a 7-min interaction with the social partner. On the day of same-sex social interaction counterbalanced with object exploration, after the first trial, the subject mouse rested with the LED off for approximately 5 min before starting the second trial. Calcium imaging and behaviour (miniCam) were recorded simultaneously at 30 Hz.
Pre-processing and extraction of calcium signals
Calcium imaging videos were motion corrected (NoRMCorre: https://github.com/flatironinstitute/NoRMCorre; MiniAn: https://github.com/denisecailab/minian) and analysed with extended constrained non-negative matrix factorization (CNMF-E)-algorithm-based packages (MiniscopeAnalysis: https://github.com/etterguillaume/MiniscopeAnalysis; MiniAn: https://github.com/denisecailab/minian) to identify cells and extract cell activity. Denoised and demixed trace data were used for further analysis.
Identification of behaviour responsive cell
To determine whether an individual neuron’s activity was significantly modulated by social interaction, we used ROC analysis to calculate an area under the curve (AUC) value for denoised calcium trace correspondence to binarized manually scored interaction bouts. The interaction bouts were defined when the imaged mouse actively investigated the social target/object within a distance of one-quarter to one-third of mouse body length. To determine whether the AUC values indicated that the neuron was significantly excited or inhibited by the interaction, we compared observed AUC values to a null distribution derived from circularly shuffled calcium signals (1,000 shuffles). Significance was determined using percentile thresholds, classifying neurons as significantly responsive if their auROC values surpassed the 97.5th percentile for excited neurons or the 2.5th percentile for inhibited neurons. Neurons that were neither excited nor inhibited were deemed non-responsive.
Calcium event calculation
Calcium transients were identified using automatic detection algorithms in MATLAB (peakfinder) (https://github.com/GradinaruLab/striatum2P).
Statistical analyses
To examine whether there were differences in DP-miniscoped or ILA-miniscoped animals in the time spent interacting during non-social (object) versus social (same- or opposite-sex partner) testing, an omnibus one-way ANOVA was performed followed by post hoc Tukey’s multiple-comparison test, which controls for the family-wise error rate. To assess the percentage of behaviourally responsive cells and frequency of calcium events during non-social (open-field) and social (same- or opposite-sex partner) testing, an omnibus two-way ANOVA was performed followed by post hoc analyses using either Tukey’s tests (for within DP or ILA comparisons) or Šidák’s tests with adjusted P values (for across DP and ILA comparisons) for multiple comparisons.
DP neuronal activities in response to social and environmental stimuli
Male mice in response to social stimuli
Twenty-four male mice, aged between 12 and 14 weeks, were used for the study. To reduce potential stress by the testing environment, each mouse was habituated to a clean cage for four consecutive days, spending 2 to 4 h in the cage and with approximately 30 min in the recording position daily. On the testing day, the mice were habituated in the clean cage for about 2 h before the experiment. Unfamiliar male mice, female mice and postnatal day 7 pups were used as the social stimuli. Each mouse was exposed to three instances of the same type of social stimulus, lasting about 6 min each, for a total duration of 20 min, with six mice in each group. In the control group, the cages were opened three times as well, but without introducing any intruders. The mice remained undisturbed in their individual cages for 80 min before perfusion.
Acute-restraint stress
Animals (n = 4, male) subjected to acute-restraint stress were removed from the colony room, transferred to a behavioural testing room for 1 h and then placed into a Tailveiner restrainer for mice (Braintree Scientific). This device consists of a clear tube, within which a plug was adjusted to limit voluntary movement. Acute restraint stress was applied for 30 min and terminated 30 min before the animals were euthanized. Throughout all studies, control mice (n = 4, male) underwent numerous sensory manipulations that were carried out in parallel with those for the mice in the restraint stress group.
Acute looming sound stimuli in comparison with white noise and foot shock
Subjects
TRAP2-CreER70 and Ai14 mice were acquired from the Jackson Laboratory (www.jax.org/strain/030323; www.jax.org/strain/007908). Homozygous TRAP2-CreER and Ai14 mice were bred to generate TRAP2-CreER/Ai14 offspring.
Drug preparation
4-Hydroxytamoxifen (4-OHT; Sigma-Aldrich, H6278) was dissolved at 20 mg ml−1 in ethanol by shaking at 37 °C for 15 min, then corn oil was added to give a final concentration of 10 mg ml−1, and the ethanol was evaporated by vacuum under centrifugation. 4-OHT (intraperitoneally (i.p.), 50 mg per kg) was injected in all TRAP2-CreER experiments70,71.
Stimulation for TRAP
Four TRAP-Cre mice were randomly assigned into two groups: control (n = 2) and foot-shock-based fear conditioning (n = 2). For auditory stimulation (noise group), on day 1 and 2, the TRAP2/Ai14 mice were adapted to custom-made boxes in a sound-proofed room for 1 h per day. On day 3, the mice were placed into their home cage for 12 h in a sound-proofed room before white-noise exposure. The mice were then placed in the custom-made boxes and exposed to 1 h broadband white-noise stimulation (70 dB SPL).
For the foot-shock group, on day 1, the mice were acclimatized in custom-made boxes in a sound-proofed room for 1 h. On day 2 (conditioning), the mice were put into custom-built boxes with a metal grid floor. After 10 min habituation to the conditioning chamber, the animals were exposed to the 20 s noise (70 dB SPL) co-terminated with a 0.75 mA foot shock (5 Hz for 1 s with the duration of each pulse = 100 ms) five times72. On day 3, the mice were subjected only to 1 h broadband white-noise exposure. Thereafter, mice received an injection of 4-OHT (i.p., 50 mg per kg) and were returned to their home cage. The cage was moved back to our animal facility 12 h after injection of 4-OHT. Sound stimulation and injection of 4-OHT were conducted in a sound-proofed room. On day 12, the mice were further presented with looming sound stimuli. A train of ten consecutive crescendo sound stimuli (with intensity increased linearly from 20 to 70 dB SPL in 0.4 s and followed by 0.6 s interval with intensity at 20 dB SPL) was repeated for 10 min with an interval of 10-s between the train stimuli. The mouse was euthanized after a 1.5 h rest.
Control mice were treated the same procedures as the auditory stimulation group but without any sound stimuli.
Images used from open-access resources
Images from the mouse ARA used in this paper are available online (https://mouse.brain-map.org/)11. Images of the flatmaps are freely downloadable from the cited article73. Bright-field in situ hybridization images from the ABA are available online (https://mouse.brain-map.org/)74. The Common Coordinate Framework (CCF) was used to generate the background 3D images of the brain, and are available online (https://alleninstitute.github.io/abc_atlas_access/descriptions/Allen-CCF-2020.html)75.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.