Ethics statement
Leukopaks from deidentified healthy donors with Institutional Review Board-approved consent forms and protocols were purchased from StemCell Technologies (200-0092). Residuals from leukoreduction chambers after Trima Apheresis from deidentified healthy donors with Institutional Review Board-approved consent forms and protocols were purchased from Vitalant.
All of the mice in this study were treated following a protocol (AN182757) approved by the UCSF Institutional Animal Care and Use Committee (IACUC).
Plasmids
For transient GFP expression, a scAAV plasmid with a CMV enhancer chicken B-actin intron (CAG) promoter was used (scAAV-CAG-GFP, Addgene, 83279). A similar ssAAV (ssAAV-CAG-GFP) has been used for biodistribution studies (Addgene, 28014).
A plasmid encoding the HIV protein gag fused to Cas9 expressing four Nuclear Localization Signal (4×NLS) was used to package Cas9 into EDVs59. For integrating genes at the TRAC locus, we used the same homology arms and sgRNA sequence as previously described1. An EGFRt sequence was cloned in following a 1928z-1XX sequence33. To potentially improve nuclease activity when combined with a Cas9-containing EDV, a U6 promoter expressing the sgRNA for targeting TRAC was cloned into the plasmids upstream of the left homology arm (pAAV-U6/TRAC-TRAC-1928-1XX-P2A-EGFRt). The extracellular domain of a BCMA-targeting CAR was used to replace the CD19-binding sequence to generate a pAAV-U6/TRAC-TRAC-BCMA-1XX-P2A-EGFRt.
Plasmids for the B7H3 experiment were designed by cloning the scFv targeting B7H3 into a 28z-1XX CAR. The construct was flanked by the same homology arms used previously targeting TRAC.
To generate a GFP fusion at the Clta N terminus, the GFP gene was cloned into an AAV plasmid containing homology arms targeting the CLTA exon 1 start codon (pAAV-CLTA-GFP). The design homology arm design and sgRNA sequences were taken from a previous study41.
To produce lentiviruses delivering an anti-CD19 CAR, a second-generation lentivirus comprising a transfer plasmid, a packaging plasmid and an envelope plasmid were used. The exact CAR sequence from pAAV-U6/TRAC-TRAC-1928-1XX-P2A-EGFRt cloned into a lentiviral transfer plasmid, under an EF1-α promoter, was used. The AAV-hT7 capsid variant generated in this study has been made available at Addgene (252215).
Cell lines
AAVs were packaged in HEK293T cells (ATCC, CRL-3216) or suspension-adapted HEK293 cells (Viral Production Cells 2.0, Gibco, A49784). Lentivirus were packaged in Lenti-X 293T cells (Takara, 632180). HEK293T and Lenti-X 293T cells were cultured in GlutaMAX DMEM (Gibco, 10566024) supplemented with FBS (10%, Corning, 35016CV), penicillin–streptomycin (100 U ml–1; Thermo Fisher Scientific, 15140122), sodium pyruvate (1 mM; Gibco, 11360070) and HEPES (10 mM; Corning, 25-060-CI). Suspension HEK293 cells were grown in viral production medium (Gibco, A4817901) supplemented with 20 mM GlutaMAX (Gibco, 35050061).
NALM6 parental lines and derivatives were cultured in RPMI 1640 (Gibco, 11875093) supplemented with FBS (10%), penicillin–streptomycin (100 U ml−1), sodium pyruvate (1 mM), HEPES (10 mM), β-mercaptoethanol (Gibco, 21985-023) and MEM non-essential amino acids (1×; Gibco, 11140050). OPM2-ffLuc-GFP cells were cultured in RPMI 1640 supplemented with penicillin–streptomycin (100 U ml−1) and FBS (10%). Raji (ATCC, CCL-86) cells were cultured in RPMI 1640 (Gibco, 11875093) supplemented with FBS (10%) and penicillin–streptomycin (100 U ml−1). JeKo1 (ATCC, CRL-3006) cells were cultured in RPMI 1640 (Gibco, 11875093) supplemented with FBS (20%) and penicillin–streptomycin (100 U ml−1). SupB15 (ATCC, CRL-1929) cells were cultured in Iscove’s modified Dulbecco’s medium with and supplemented with β-mercaptoethanol (Gibco, 21985-023), FBS (20%) and penicillin–streptomycin (100 U ml–1). MES-SA cells (ATCC, CRL-1976) were cultured in McCoy’s 5a modified medium (Gibco, 16600082) supplemented with 10% FBS and penicillin–streptomycin. Suspension HEK293 cells were maintained in an 8% CO2 incubator at 37 °C, on a 25 mm orbital shaker. All of the other cell lines were cultured under sterile conditions in a 5% CO2 incubator at 37 °C.
Primary cell isolation and culture
PBMCs
PBMCs were used either fresh or from frozen aliquots. Apart from fresh PBMCs used for in vivo experiments described in Figs. 1i, 3a and Extended Data Fig. 7a, all PBMCs were sourced from Leukopaks (StemCell Technologies, 70500.1). Fresh Leukopaks were either directly processed for further cell purification or were frozen in aliquots.
For the experiments shown in Figs. 1i and 3a and Extended Data Fig. 7a, PBMCs were isolated from TRIMA residuals (Vitalant) using Lymphoprep reagent (StemCell Technologies, 07801) and SepMate-50 tubes (StemCell Technologies, 85450) according to the manufacturer’s protocol.
T cells
Human T cells were isolated from frozen aliquots of PBMCs. After thawing, T lymphocytes were purified using the EasySep Human T cell isolation kit (StemCell Technologies, 17951) and activated with Dynabeads Human T Expander CD3/CD28 at a 1:1 bead-to-cell ratio (Gibco, 11141D) in X-VIVO 15 medium (Lonza, BP04-744Q) supplemented with human serum (5%, Gemini Bioproducts, 100-512), IL-7 (5 ng ml−1, Miltenyi Biotec, 130-095-367) and IL-15 (5 ng ml−1, Miltenyi Biotec, 130-095-760) at a density of 1 × 106 cells per ml. Unless otherwise specified, beads were removed after 48 h, and T cells were maintained at 1 × 106 cells per ml every 2 to 3 days by addition of fresh medium.
NK cells
NK cells were isolated from fresh PBMCs (TRIMA residual, Vitalant) by negative selection using the EasySep Human NK Cell Enrichment Kit (StemCell Technologies). Isolated NK cells were cultured at an initial density of 106 cells per ml in NK MACS medium (Miltenyi) supplemented with human platelet lysate (5%, Elite Cell), penicillin–streptomycin (0.5%) and IL-2 (1,000 U ml−1, Peprotech), as previously described60. After an overnight rest period, cells were activated for 7 days with anti-CD2- and anti-NKp46-coated (Miltenyi) beads at a 1:2 bead-to-cell ratio.
Macrophages
Fresh PBMCs were obtained from StemCell Technologies (70500.1) and subjected to the EasySep Human CD14 positive selection kit II (StemCell Technologies, 17858) according to the manufacturer’s protocol. Then, 0.5 × 106 isolated CD14 monocytes were seeded into 24-well plates for differentiation into human monocyte-derived macrophages in RPMI with 10% FBS, 1% penicillin–streptomycin, 1× GlutaMAX, 1× HEPES and 20 ng ml−1 recombinant human GM-CSF (PeproTech, 300-03) for 9 days.
HSCs
HSCs were isolated from G-CSF-mobilized peripheral blood of de-identified healthy donors and CD34 enriched and then cryopreserved (Fred Hutchinson Cancer Center, Hematopoietic Cell Procurement and Processing Core). Cryopreserved cells were thawed and incubated for 1 h at 37 °C in RPMI 1640 medium (Thermo Fisher Scientific) supplemented with 30% FBS, 1% penicillin–streptomycin, 10 μg ml−1 DNase I and 20 U ml−1 heparin. As previously described61, HSPCs were then cultured at 1 × 105 cells per ml in StemSpan Serum-Free Expansion Medium II (StemCell Technologies) or Good Manufacturing Practice Stem Cell Growth Medium (SCGM, CellGenix) supplemented with a human cytokine cocktail (PeproTech) that included 100 ng ml−1 stem cell factor, 100 ng ml−1 thrombopoietin, 100 ng ml−1 FMS-like tyrosine kinase 3 ligand, 100 ng ml−1 interleukin-6, 20 mg ml−1 streptomycin and 20 U ml−1 penicillin.
EDV and LVV production
Transfection
Lenti293T cells were seeded into packaging medium consisting of OptiMEM-GlutaMAX (Gibco, 51985034) supplemented 5% FBS and 1 mM sodium pyruvate. Cells were transfected the next day with the appropriate plasmids to produce EDVs or replicative-deficient LVV. A total of 60 μg of plasmids complexed with 181 μl of Lipofectamine 3000 and 165 μl p3000 were used per T225 flasks. EDVs were produced by transfecting producer cells with an envelope plasmid (env) and two packaging plasmids, one encoding a Gag–Cas9 fusion (Gag–Cas9), and one rescue packaging plasmid (PsPax2). A U6 promoter and sgRNA sequence was incorporated in both packaging vectors to generate EDV-carrying RNPs (Supplementary table 1). Env-to-gagCas9-psPax2 mass ratios were respectively 1:2:1 (v/v) were produced by transfecting producer cells with an env plasmid, a PsPax2 plasmid and a transfer plasmid encoding a 1928z-1XX sequence. The mass ratios of Env to psPax2 to transfer were 1:3:0.75 (ref. 27).
Particle pseudotyping
WT EDVs and LVVs were generated using the pMD2.G plasmid as the env plasmid. For T cell-targeted EDVs and LVVs, the pMD2.G plasmid was substituted with a construct encoding a mutated VSV-G glycoprotein (K47Q, R354A) fused through a P2A linker to an scFv specific for CD3.
Collection and isolation
The medium was replaced after 6 h with fresh packaging medium supplemented with 0.2% (v/v) ViralBoost (Alstem, VB100). LVVs and EDVs were collected 48 h after transfection. Cell debris was pelleted by centrifugation at 2,000g for 10 min. The supernatant was clarified through a 0.45 μm filter before purification by ultracentrifugation on a sucrose cushion (50 mM Tris-HCl, 100 mM NaCl, 0.5 mM EDTA, 20 w/v% sucrose, pH 7.4). The supernatants were spun down at 91,000g for 90 min. Pellets were resuspended in ice-cold PBS and immediately stored at −80 °C.
Particle titration
EDVs were titred by measuring the sgRNA concentration in the particles, as previous studies showed that the number of sgRNA equates to the number of functional Cas9 RNPs inside the EDVs27. The sgRNA concentration was determined using quantitative PCR with reverse transcription (RT–qPCR). EDVs were diluted tenfold in DirectDetect buffer (Zymo Research, R1400) according to the manufacturer’s instructions. For RT–qPCR, custom TaqMan small RNA assays were designed to detect the sgRNA sequence (Thermo Fisher Scientific, CTZTEYN (for CLTA sgRNA) and CTCE4RX (for TRAC sgRNA)). Synthetic sgRNA sequences with the appropriate spacers were used as standards. We added 4 μl of samples or standards to 6 μl of RT–qPCR mastermix (1× Luna Luna Universal One-Step RT–qPCR mix with 0.25× RT primer and 1× small RNA assay probes from TaqMan small RNA assay kit in nuclease-free water) in 384-well plates. RT–qPCR was performed on the QuantStudio 5 Real-Time PCR System (Thermo Fisher Scientific) using the following parameters: carryover prevention (25 °C, 30 s), reverse transcription (55 °C, 15 min), initial denaturation (95 °C, 1 min) and 45 cycles of denaturation (95 °C, 10 s), extension (60 °C, 60 s) with a plate read.
For the comparative study between the combined EDV and AAV treatment and the LVV treatments, two normalization methods were used. The first normalization was based on the number of anti CD3-pseudotyped particles (LVV and EDV) injected. Physical titres of LVV and EDV preparations were measured through ELISA-based p24 quantification (Lenti-X p24 Rapid Titer Kit, Takara, 632200) according to the manufacturer instructions. The total p24 amount per EDV dose was calculated based on the sgRNA titres (here, 5 × 1011 sgRNA per mouse) and a matching p24 amount of LVV was injected. The mice injected with these samples are referred to as the LVVhigh dose group.
Functional normalization was also performed. Transducing units (TU) were measured based on the ability to generate CAR T cells in vitro, according to the formula below.
$$\mathrm{TU}\,\mathrm{per}\,\mathrm{ml}=\frac{\mathrm{number}\,\mathrm{of}\,\mathrm{cells}\,\mathrm{transduced}\times \mathrm{percentage}\,\mathrm{of}\,{\mathrm{CAR}}^{+}\,\mathrm{cells}}{\mathrm{volume}\,\mathrm{added}\times \mathrm{dilution}\,\mathrm{factor}}$$
As AAVs have been found to be non-limiting in in vitro settings, a fixed amount of AAV was used with decreasing doses of EDVs to calculate the total TUEDV per preparation. LVVs were also used in decreasing doses to calculate total TULVV. The sgRNA dose of EDV was converted into TUEDV and a similar TULVV dose was injected. The mice injected with these samples are referred to as the LVVlow dose group.
According to manufacturer guidance and previous work27, 5 × 1011 sgRNA molecules are the equivalent to approximately 1.1 × 1011 EDV particles or 8.8 µg of p24. Approximatively three times less LVV was injected for the functional normalization.
AAV production and quantification
AAV2-ITR-containing plasmids were used to package vector genomes into various AAV capsids by transfection of HEK293T cells with adenovirus helper and AAV Rep-Cap plasmids using polyethyleneimine (Polysciences, 23966). For small-scale production, HEK293T cells were seeded into 150 mm dishes, and a total of 20 μg of plasmid complexed with 200 μl PEI was used per dish. The mass ratio of cargo:Rep-Cap:helper was 1:1.3:1.8.
Cell pellets were collected 72 h after transfection and resuspended in AAV lysis buffer (50 mM Tris, 150 mM NaCl) before being lysed by three rounds of rapid freeze–thawing. Polyethylene glycol (PEG) was used for precipitation of supernatants. Crude lysates and pelleted supernatants were combined before downstream purification.
For large-scale production, suspension-adapted HEK293 cells were transfected using the VirusGEN AAV Transfection Kit (Mirus, MIR 6750) according to the manufacturer’s recommendations. A total of 2 μg of plasmid were used per millilitre of cell culture, while keeping the same mass ratios. Producer cells were lysed 72 h after transduction by chemical disruption (AAV Lysis Buffer, Gibco, A50520) and crude lysates were concentrated by PEG precipitation.
AAV-containing solutions were incubated for 1 h at 37 °C with 25 U ml–1 benzonase (Millipore Sigma, 70-664-3). Solutions were clarified through centrifugation and AAV vectors were further purified using iodixanol (OptiPrep, StemCell Technologies, 07820) gradient ultracentrifugation.
AAV vector titres were determined by qPCR on DNase-I-treated (NEB, B0303S), proteinase-K-digested (Qiagen, 1114886) AAV samples after purification, using primers targeting the viral genome. qPCR was performed with PowerUp SYBR Green Master Mix (Applied Biosystems, A25918) on a StepOnePlus Real-Time PCR System (Applied Biosystems, 4376600). Relative quantity was estimated by comparison to a serial dilution of a vector plasmid standard of known concentration.
AAV capsid library generation and evolution
The AAV6 capsid library was generated by performing saturation mutagenesis of seven residues in the VR-IV region as reported previously62. In brief, to generate the library, an overlap extension PCR was performed using two amplicons amplified from a modified AAV6 backbone containing seven tandem stop codons replacing the randomized region (amino acids 454–460 VP1 numbering) to prevent potential amplification of the WT sequence. The first amplicon consisted of 34 bp of the AAV6 cap immediately 5′ of the randomized region, the randomized region and the AAV6 cap up to the Sbf1 site 3′ of the randomized region. The second amplicon consisted of the AAV6 cap starting from the BsiWi site 5′ of the randomized region up to the 34 bp of the AAV6 cap immediately 5′ of the randomized region. The two resulting amplicons were combined in an equimolar ratio in a second PCR for overlap extension. The final assembled amplicon was digested using BsiWI-HF (NEB, R3553S) and SbfI-HF (NEB, R3642S) and ligated into the pITR2-Rep2-dead(GFP)Cap6 backbone using T4 DNA ligase (NEB, M0202S). The pITR2-Rep2-dead(GFP)Cap6contains AAV2 ITRs and Rep along with the AAV6 Cap gene interrupted by a filler sequence derived from GFP inserted out of frame into the cognate BsiWI and SbfI site to eliminate any potential WT AAV6 from the library ligation. The ligation products were concentrated and purified by ethanol precipitation. Purified products were electroporated into DH10B ElectroMax cells (Invitrogen, 18290015) and directly plated onto multiple 5,245-mm2 bioassay dishes (Corning, 431111) with LB/ampicillin agar to maintain library diversity. Plasmid DNA from AAV6 capsid libraries was purified from pooled colonies grown on LB agar plates with ampicillin using the ZymoPURE II plasmid maxiprep kit (Zymo Research, D4203).
AAV6 capsid libraries were produced by co-transfection of adherent HEK293T cells with adenovirus helper plasmid (pXX680,49 19.25 μg per plate) and the Rep Cap plasmid library (8.75 μg per plate) mixed in a 3:1 ratio with PEI MAX (Polysciences, 24765-1). Viral medium was collected at day 4 and 6, and the cell pellet at day 6. The medium was PEG precipitated, and the cell pellet was lysed through chemical disruption. AAVs were purified as described above.
Library evolution was performed by transducing 48-h-activated T cells with the capsid library at a MOI of 1 × 104. To evolve the library, primary human T cells were isolated and activated with CD3/CD28 beads and recombinant IL-7/IL-15 and then co-cultured with the capsid library at a low MOI of 1 × 104. Cells were then washed twice with PBS to remove any unbound AAV, and cellular and viral DNA was extracted from cells using the IBI genomic DNA extraction kit (IBI Scientific, IB47280). The Cap region was amplified by PCR before being digested and ligated back into the pITR2-Rep2-dead(GFP)Cap6 backbone to generate the next-round library.
A total of three rounds of evolution was performed to generate an evolved library. Each round of library was performed on two different donors, for a total of six different donors used.
AAV6 capsid library sequencing and analysis
Parental and evolved libraries were processed for Illumina NovaSeq sequencing. Parental and evolved libraries were each treated with DNase I and purified by iodixanol gradient centrifugation. To dissociate the capsid, virus was heated in a PCR tube (95 °C, 15 min) with Tween-20, which prohibits capsid reassembly that would interfere with amplification.
Round-1 PCR was performed with defined primer sets (Supplementary Table 1) for 18 cycles using Q5 polymerase (NEB, M0492S), and amplicons were PCR-purified (IBI Scientific, IB47010). In round 2, indices for demultiplexing and the P5 and P7 flow cell adaptor sequences were added in a 15-cycle PCR, and amplicons were run on and purified from a 1% agarose gel. The amplicon band was gel-purified, amplicon quality was verified using a Bioanalyzer (Agilent) and concentrations were quantified by Qubit (Invitrogen). Libraries were prepared using the Illumina NovaSeq 6000 S-Prime reagent kit (300 cycles, Illumina, 20028312) according to the manufacturer-provided instructions and sequenced by Illumina NovaSeq.
Demultiplexed reads were analysed using an in-house Perl script as done in another context previously63. Reads were probed for the nucleotide sequences corresponding to the library region, and the occurrence of each nucleotide sequence was counted and ranked. These sequences were converted to amino acid sequences and pooled by like-sequence, counted and ordered by percentage rank. A second Perl script was used to calculate enrichment between the evolved library and parental library as done previously63. Data in Supplementary Table 2. To generate the amino acid position-specific scoring matrix, sequences that were above 0.1% of the reads of the evolved library and enriched more than 100-fold from the parental library (34 sequences) were selected and run through PSSMSearch (http://slim.icr.ac.uk/pssmsearch/).
Combined EDV + AAV transduction
Unless otherwise specified, combined EDV + AAV transduction of T cells was performed as follows. 48-h-activated T cells were seeded at 1 × 106 cells per ml in T cell medium, and particles were added at a specified MOI. The volume of EDV/AAV added did not exceed 20% of the culture volume. After incubating the culture overnight, the AAV-containing medium was exchanged for fresh medium, and T cells were subsequently cultured in standard conditions.
Genome-wide CRISPR screens identify AAVs host factors
Genome-wide CRISPR–Cas9 screening
A genome-wide sgRNA KO library targeting 19,114 genes (comprising 76,441 sgRNAs) was sourced from Addgene (Brunello Library, 73178) and amplified according to the guidelines for maintaining library representation64. Viral packaging of the library in lentivirus was conducted as previously described. Screens were performed with cells isolated from two healthy donors. T cells were obtained from thawed PBMC aliquots and activated using Dynabeads. Then, 36 h after activation, T cells were transduced with the lentiviral library at an MOI of approximately 0.3. The lentiviral titre was predetermined using T cells from the same donors. For this experiment, integration rates were measured at 40–45% after puromycin selection. After a further 36 h, beads were removed and Cas9 protein was introduced via nucleofection. Nucleofection was performed using the P3 Primary Cell 96-well Nucleofector Kit (Lonza V4SP-3096) on the Amaxa 4D Nucleofector unit (Lonza, AAF-1002B). Cells were resuspended at 1 × 108 per ml in P3 buffer and Cas9 protein (40 μM stock) was added to the cells at a 1:20 ratio and then electroporated using pulse code EH-115.
Cells recovered for 2 days, then underwent puromycin selection at 2 µg ml−1 (Gibco A1113803) for another 2 days. After selection, cells were washed twice, cultured for two more days, and restimulated with Dynabeads for 48 h at a 1:10 beads-to-cell ratio to enhance transcriptional activity while limiting activation-induced cell death.
Cells were transduced at an MOI of 5 × 103 in serum-free conditions. After overnight transduction, cells were washed, and serum was reintroduced by replenishing cells with fresh medium. Then, 48 h after AAV transduction, cells were prepared for sorting by staining with 7-AAD live/dead reagent (eBioscience, 00-6993-50), followed by fixation in 4% formaldehyde in PBS (15 min, 4 °C) at a concentration of 107 cells per ml. Fixed cells in FACS buffer were sorted into four bins based on GFP expression. More than 4.5 × 107 cells per replicate were sorted at the UCSF Parnassus Flow Cytometry Core (PFCC).
After cell sorting, genomic DNA extraction was performed according to established protocols65. sgRNA barcodes underwent PCR amplification utilizing Ex Taq DNA Polymerase (Takara Bio) through 28 thermal cycles, followed by amplicon purification with SPRIselect Beads (Beckman Coulter). Quality assessment was conducted using D1000 ScreenTape assay on a TapeStation system (Agilent). For sequencing preparation, P7 and P5 primers for lentiGuide compatibility (generating 354-nucleotide products). PCR amplification of sgRNA sequences was performed using standard protocols (https://portals.broadinstitute.org/gpp/public/) with P5/P7 primers obtained from IDT.
Library pooling preceded sequencing on a NovaSeq X platform (300 × 151 × 12 × 24 × 151 configuration) across two lanes at UCSF’s Center for Advanced Technology. Post-sequencing FASTQ files underwent processing and analysis using MAGeCK v.0.5.9.5. Data visualization was performed using volcano plots with the log-transformed fold change on the x axis (negative values indicating GFP-low bin enrichment) and −log10[P] on the y axis, representing the inverse error probability where higher values correspond to increased statistical confidence. Data are provided in Supplementary Table 3.
Individual validation
To validate the function of genes identified in the genome-wide screens, individual gene knockouts were performed by delivering Cas9–RNP complexes. T cells activated for 48 h were resuspended at a concentration of 1 × 108 cells per ml in P3 buffer. RNP complexes were generated by incubating Cas9 protein (40 μM stock) with targeting sgRNAs (80 μM stock) at a molar ratio of 2:1 for 15 min at 37 °C. For each experiment, 3 μl of RNP complexes were added to 20 μl of resuspended T cells. Alternatively, CD7-KO T cells were generated using base-editing by adding 1 µg mRNA encoding an adenine base editor and 100 pmol CD7-targeted sgRNA to 20 µl resuspended T cells. Electroporations were conducted using pulse code EH-115.
Edited T cells were cultured for 6–8 days, reactivated with Dynabeads at a 1:10 beads-to-cell ratio for 48 h, then transduced with various AAVs carrying sc-CAG-GFP at specified MOIs. The medium was changed the next day, and GFP expression was assessed using flow cytometry 48 h after transduction.
Animal work
All mice in this study were treated following a protocol approved by the UCSF Institutional Animal Care and Use Committee (IACUC) and were housed under a 12 h–12 h light–dark cycle with food and water available ad libitum. Unless specifically mentioned, all procedures were performed in a BSL-1 room. NSG (005557) and NSG-MHCI/II double-KO (025216) mice were acquired from the Jackson Laboratory. Only female mice were used in this study.
Humanization
Unless specifically mentioned, all humanization was performed using frozen aliquots of PBMCs. PBMCs were thawed on the day of injection, washed, counted and kept in ice-cold RPMI medium before injection. NSG-MHCI/II double-KO mice (aged 7–12 weeks) were injected i.v. with 1 × 107 of thawed PBMCs.
Organ collection for phenotyping
Spleens were collected, crushed in FACS buffer, strained and then treated with 1 ml ACK lysing buffer (Quality Biological, 118-156-101) for 2 min. The lysing process was quenched by adding 20 ml FACS buffer, the cells were then strained and resuspended in FACS buffer and stained for flow cytometry.
Bone marrow was collected by crushing the hind legs (femur and tibia) using a mortar and pestle in FACS buffer. The crushed bones marrows were then strained and treated with 1 ml ACK lysing buffer (Quality Biological, 118-156-101) for 2 min. The lysing process was quenched by adding 20 ml FACS buffer, the cells were then strained and resuspended in FACS buffer and stained for flow cytometry.
Circulating blood was collected through submandibular cheek bleeding. Blood samples were collected in EDTA-coated tubes. Red blood cells were lysed with RBC lysis buffer (BioLegend, 420302) for 15 min at room temperature.
B cell depletion models
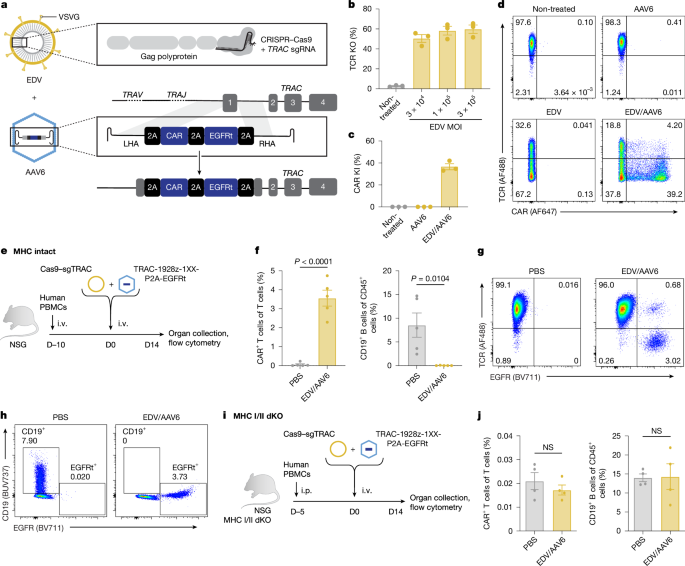
For experiments with MHC-intact mice, 8-week-old NSG mice were injected i.v. with 1 × 107 frozen PBMCs. EDVs and AAVs were injected 10 days later. For experiments with MHC-KO mice, NSG-MHCI/II double-KO mice were engrafted i.p. with 2 × 107 fresh human PBMCs isolated from Trima residuals, followed by EDV and AAV injection 5 days later.
For all experiments, mice were injected i.v. with a mixture of AAV (1 × 1012 vg per mouse) and EDV (2.5−5 × 1011 sgRNAs per mouse) in PBS and euthanized for organ collection 14 days after EDV/AAV injection.
Tumour models
Experimental scheduling
Across all models and to support tumour engraftment, mice were first injected with a given dose of tumour cells before humanization. Humanization was performed either i.p. or i.v., at the indicated time. Mice were injected at the indicated time with a mixture of AAV (1 × 1012 vg per mouse) and EDV (2.5−5 × 1011 sgRNA per mouse) in ice-cold PBS. Only MHC-I/II double-KO mice were used for tumour challenges.
Tumour measurement and end points
Mice were humanely euthanized at an IACUC-approved humane end point, including respiratory distress, hunched posture, body condition score of 2 or less, 15% weight loss, impaired or decreased mobility or neurological signs that interfere with normal function.
For haematological malignancies (NALM6 and OPM2), tumour burden was monitored at regular intervals by BLI using the Xenogen IVIS Imaging System (Xenogen) with Living Image software (Xenogen) for acquisition of imaging datasets. Both dorsal and ventral images were obtained for each animal. The dorsal and ventral signals were separately quantified through region of interest (ROI). The resulting signal summations (in units of photons per s) were normalized to the ROI area so that all measurements are given in photons per s per cm2.
For solid tumour challenge, the tumour volume was measured using callipers by first determining the length as the longest dimension and the width as the shortest perpendicular dimension of the tumour. Tumour volume was calculated according to the standard formula below.
$$\frac{1}{2}\times \mathrm{length}\times {\mathrm{width}}^{2}$$
B-ALL experiments
NSG MHC-I/II double-KO (aged 7–9 weeks) mice were purchased from Jackson Laboratories. Apart from the experiment described in Fig. 1i–j, mice received 5 × 105 NALM6 ffLuc-GFP by i.v. injection and were humanized 3 days later by i.v. injection of 1 × 107 of PBMCs from frozen stocks. For the experiment in Fig. 1i–j, NSG-MHCI/II double-KO mice were i.v. injected with 2.5 × 105 NALM6-FFLuc-GFP, followed 3 days later by i.p. injection with 2 × 107 fresh PBMCs.
In all cases, EDV and AAV were injected the day after PBMC injection. Mice were injected either i.p. or i.v. with a mixture of AAV (1 × 1012 vg per mouse) and EDV (2.5–5 × 1011 sgRNAs per mouse).
For the experiment in Fig. 4c–j, LVV injection was performed at the same time as EDV and AAVs. Injected doses of LVV were normalized to EDV/AAV treatment based on the titres (the quantification methods are described above). Two normalization methods were used. An equivalent number of EDV and LVVs was injected, based on the quantification of particles in the two viral preparations (LVVhigh). Volumes were also normalized based on the functionality, so that an equivalent TU was injected between the EDV/AAV and the second LVV condition (LVVlow). For this experiment, the EDV-to-LVV volume ratio was 1:0.7 for the first method, and 1:0.2 for the latter.
Ex vivo manufactured cells were generated from T cells isolated from the same PBMCs aliquots. T cells were isolated and activated for 2 days. Cells were left untreated, treated with a WT pseudotyped LVV delivering a CD19-28z-1XX cargo, or electroporated with TRAC targeted sgRNAs and Cas9 protein. Electroporated cells were either left for recovery or transduced with an AAV delivering an HDRT to precisely knock-in the same CAR CD19-29z-1xx at the TRAC locus.
CAR expression was measured by flow cytometry 5 days after treatment, and then again before freezing at day 10 of expansion. Cells were counted and frozen on the CryoStor CS10 (StemCell Technologies, 100-1061). T cells were thawed on the day of injection, washed in RPMI and injected i.v. in non-humanized age-matched mice.
Multiple myeloma model
MHC-I/II double-KO mice (aged 8 weeks) were purchased from Jackson Laboratories, mice were injected i.v. with 1 × 106 OPM2 ffLuc-GFP, then humanized i.v. 5 days later with 1 × 107 PBMCs from frozen vials. EDV/AAV treatment was performed 5 days after humanization through i.v. injection.
Sarcoma model
MHC-I/II double-KO mice (aged 8 weeks) were obtained from Jackson Laboratories. MES-SA cells (4 × 106) were suspended 1:1 in PBS and Matrigel and injected subcutaneously into the flank of immunocompromised mice. Matrigel supported cell viability and tumour establishment. Tumour growth was monitored regularly.
TCR-seq analysis
To isolate in vivo engineered CAR T cells, splenocytes were labelled with anti-EGFR-biotin (BioLegend, 352934) and isolated using anti-biotin MicroBeads (Miltenyi Biotec, 130-090-485) on LS columns (Miltenyi Biotec, 130-042-401). T cell clonality was analysed on RNA from isolated CAR T cells using SMART Human TCR a/b Profiling Kit (Takara Bio, 634780). Samples were pooled and sequenced on the Illumina MiSeq system using 2 × 300 bp paired-end chemistry and the MiSeq Reagent Kit v.3 (MS-102-3003, Illumina). FASTQ files were analysed using MiXCR to identify TCR clonotypes. Clonotypes were defined by unique CDR3 amino acid sequences combined with V and J gene assignments, and clonotypes with fewer than two reads were excluded from analysis. Clonality metrics and visualization were performed using custom Python scripts.
Biodistribution
For biodistribution studies, NSG-MHCI/II double-KO mice (aged 7 weeks) were first humanized with frozen PBMCs (1 × 107 per mouse). Then, 10 days later, mice were injected i.v. with AAV6 or 312 delivering a ssAAV-CAG-GFP. After 1 week, mice were euthanized and transcardial perfusion was performed, then organs were collected and snap-frozen in liquid nitrogen. DNA extractions were performed using the PureLink Genomic DNA Mini kit (Invitrogen, K182001). Eluted DNAs were further cleaned and concentrated using NA Clean & Concentrator-5 Kit (Zymo, 1159U31).
Viral genomes were quantified by qPCR targeting the GFP gene, and the total copies were normalized to the DNA input.
Cytokine quantification in vivo
NSG-MHCI/II double-KO mice (aged 9 weeks) were humanized with frozen PBMCs (1 × 107 per mouse). Ten days later, mice were i.v. injected with different AAVs (1 × 1012 per mouse), EDVs (5 × 1011 per mice) or a combination of both. Mice were euthanized at the indicated times and blood was immediately collected through cardiac puncture in non-treated tubes. Blood samples were allowed to clot, and the resulting serum was diluted 1:2 in PBS, stored at −80 °C, and shipped for analysis. Serum contents were analysed for a panel of 41 human and mouse cytokines (Eve Technologies, HUMU41)
Flow cytometry
Cells were stained to distinguish dead cells in 100 μl PBS for 30 min at room temperature using either Zombie Violet (BioLegend, 423114), Zombie NIR (BioLegend, 423105) or Ghost Dye Red 780 (Tonbo, 13-0865-T100).
Cells were stained in 100 μl FACS buffer (2% FBS and 1 mM EDTA in PBS) for 30 min at room temperature using the following reagents: anti-TCRa/b AF488 (BioLegend, 306712), anti-TCRa/b BV421 (BioLegend, 344646), anti-G4S AF647 (Cell Signaling Technologies, 69782), anti-CD45 PE (BioLegend, 368510), anti-CD25 BV711 (BioLegend, 356138), anti-CD69 PE (BioLegend, 310906), anti-CD4 BUV395 (BioLegend, 583550), anti-CD8 BV421 (BD, 568217), anti-CD8 BV711 (BioLegend, 344734), anti-CD8 PE-Cy7 (Invitrogen 25-0087-42), anti-EGFR AF488 (BioLegend, 352908), anti-EGFR BV711 (BioLegend, 352290), anti-CD19 BUV737 (BD, 612756), anti-CD3 BUV395 (BD, 563546) and anti-CD56 BV711 (BioLegend, 318336).
For quantification of cells during flow cytometry, 50 μl of CountBright Absolute Counting Beads (Invitrogen, C36950) was added to each sample according to the manufacturer’s protocol.
Spectral flow cytometry
Preparation of reference controls for spectral flow cytometry
For determining optimal controls for unmixing, reference controls for each marker were prepared on Ultra Comp eBeads Plus (Invitrogen 01-3333-42), Slingshot HyParComp (Slingshot SSB-20-A) and cells (1 million human PBMCs), except for LIVE/DEAD blue. Beads were stained with individual antibodies at a dilution of 1:100. Cells were stained with individual antibodies using the same concentration used in the surface and intracellular stain master mixes. Surface markers were diluted in FACS buffer and intracellular markers were diluted in permeabilization buffer. All of the reference controls underwent the same protocol as the fully stained samples, including incubation/staining times, washes and fixation/permeabilization steps. The same antibody lot was used for both reference controls and full-stained samples. For LIVE/DEAD blue staining, 1 million cells were heat killed at 65 °C for 15 min. Half of the cells were stained with LIVE/DEAD blue under the same conditions as full-stain samples (50 µl staining volume in PBS, 25 min at room temperature, protected from light), washed, then mixed with the remaining unstained dead cells. An unstained human PBMC sample was used in the reference group as a universal negative for the CD45-PE single-cell reference control. Reference controls were acquired on a five-laser Aurora spectral flow cytometer (Cytek Biosciences).
Data were unmixed using SpectroFlo v.3.3 software (Cytek Biosciences). Unmixing quality was assessed using bead versus cell controls and optimal reference controls were chosen as described in a previous study66. Moreover, the unstained reference control was used for unmixing with autofluorescence extraction. The resulting unmixed .fcs files were analysed using manual gating in FlowJo v.10.10 software (BD Biosciences). Panel performance was further assessed by generating NxN plots and manually inspecting marker expression in CD4+ and CD8+ T cells. Supplementary Table 2 reports the single stain reference control that was used for each marker in the panel.
Spectral flow staining and data acquisition
In total, 2 million isolated splenocytes were stained for each sample. Moreover, 2 million splenocytes were taken from one sample to serve as a group-specific unstained control for unmixing. To block Fc receptor binding, cells were incubated with FcR blocking reagent, mouse (Miltenyi 130-092-575) for 30 min at 4 °C. LIVE/DEAD fixable blue (Invitrogen, L34961) viability stain was reconstituted in 50 µl of DMSO and serially diluted 1:1,000 in PBS. Viability stain was added to all full-stain sample cells in a 50 µl staining volume and incubated at room temperature for 25 min protected from light. Cells were washed once with PBS and once with FACS buffer before proceeding to surface staining. To make the surface antibody master mix, antibodies conjugated to fluorophores excited by ultraviolet (UV), violet and blue lasers were added first into BD brilliant buffer (BD 00-4409-42). The remaining surface antibodies excited by yellow-green and red lasers were then added, followed by FACS buffer. Then, 50 µl of surface master mix was added to each well except for the unstained controls, to which 50 µl of FACS buffer was added. Cells were incubated with surface master mix at room temperature for 25 min protected from light, then washed twice with FACS buffer. Cells were fixed and permeabilized using the FOXP3/transcription factor staining kit (eBioscience 00-5523-00) at room temperature for 1 h protected from light (200 µl per well). The samples were washed once in a permeabilization buffer (full-stain samples) or FACS buffer (unstained sample). Unstained cells were resuspended in FACS buffer and full-stain samples were resuspended in permeabilization buffer and kept overnight at 4 °C protected from light. The next day, the intracellular antibody master mix was prepared by first adding ultraviolet, violet and blue excited fluorophores to BD brilliant buffer (BD 00-4409-42). The remaining intracellular antibodies excited by yellow–green and red lasers were then added, followed by a permeabilization buffer. Then, 50 µl of intracellular master mix was added to each well and cells were incubated for 1 h at 4 °C protected from light. After staining, cells were washed with permeabilization buffer then FACS buffer and resuspended in FACS buffer. Antibody information is provided in Supplementary Table 4.
Samples were acquired on a five-laser Aurora spectral flow cytometer (Cytek Biosciences). The reference control data were used within 1 week of acquiring by choosing ‘duplicate experiment with reference controls’ in the SpectroFlo v.3.3 software (Cytek Biosciences). In addition to acquiring the full-stain samples, a group-specific unstained control was recorded for unmixing with autofluorescence extraction of the full-stain samples. The samples were unmixed using the same single-stain reference controls and the same gating of positive and negative populations that was used when generating the single-stain controls. Full-stain samples were checked for unmixing errors using NxN plots after unmixing. The resulting unmixed .fcs files were analysed using manual gating in FlowJo v.10.10 software (BD Biosciences).
Editing frequency analysis by dPCR
Between 2 and 4 days after editing, cells were collected and QuickExtract DNA extraction solution (Epicentre) was used to collect genomic DNA. In some cases, extracted genomic DNA was concentrated using SPRIselect (Beckman Coulter). The genomic DNA was then digested using HindIII-HF according to the manufacturer’s instructions (New England Biolabs). The percentage of targeted alleles within a cell population was measured using Qiagen’s QIAcuity One 5-Plex Digital PCR System and QIAcuity Software Suit (v.2.5.0.1; Qiagen) using the following reaction mixture: 1–4 μl of digested genomic DNA input, 3 μl of QIAcuity Probe PCR Master Mix (Qiagen), primer/probes (0.8 µM; Integrated DNA Technologies) and volume was brought up to 12 μl with H2O. The sample reaction mixture was then added to QIAcuity Nanoplate 8.5 K 24-well (Qiagen). QIAcuity dPCR (Qiagen) settings were as follows: 95 °C (10 min), 95 °C (15 s), 59.6 °C (30 s), and return to step-two ×40–50 cycles. Analysis of droplet samples was performed using the QIAcuity Software Suit (Qiagen). To determine percentages of alleles targeted, the numbers of Poisson-corrected integrant copies per ml was divided by the numbers of reference DNA copies per ml. The following primers and 6-FAM/ZEN/IBFQ-labelled hydrolysis probes were purchased as custom-designed PrimeTime qPCR assays from Integrated DNA Technologies (Supplementary Table 1).
Statistics and reproducibility
The statistical methods and the number of mice or donors are indicated in figure legends for each experiment. In vivo experiments were performed with multiple donors or preceded by pilot experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.