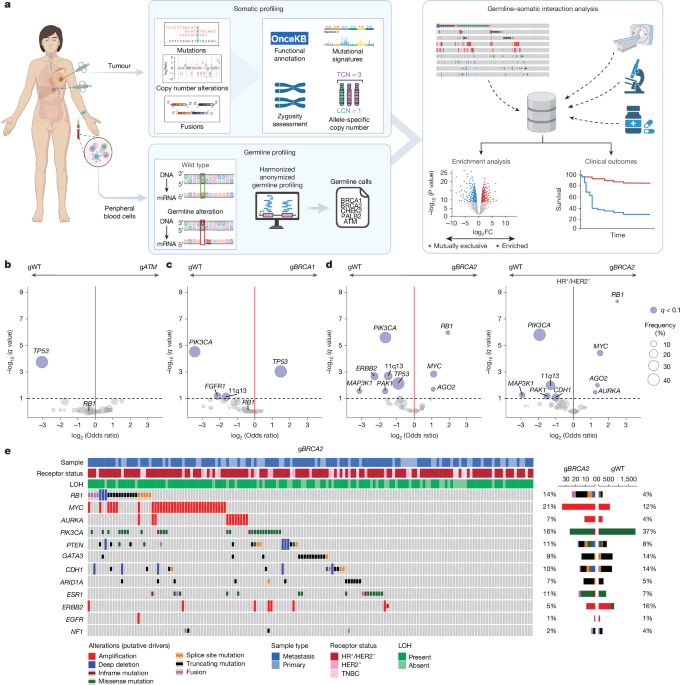
Study cohort and prospective sequencing
The study cohort comprised 6,927 tumour samples from 5,881 patients with breast cancer. All patients underwent prospective clinical tumour and normal DNA sequencing as part of their clinical care (February 2014 to September 2021). The present study was approved by the MSK Institutional Review Board (IRB) and all patients provided written informed consent for tumour and paired normal DNA sequencing and review of medical records for clinical annotations. Genomic sequencing was performed on tumour DNA extracted from formalin-fixed, paraffin-embedded tissue and normal DNA extracted from mononuclear cells from peripheral blood in all patients as previously described22. Patient samples were sequenced in a CLIA-compliant laboratory using one of several versions of the MSK-Integrated Mutation Profiling of Actionable Cancer Targets (IMPACT) targeted sequencing panel, which interrogates exonic and selected intronic regions of 341, 410, 468 or 506 genes depending on the assay version, with somatic mutation calling performed using an extensively validated pipeline followed by manual review, as previously described22,23. Tumours were obtained from the primary site in 50.1% (n = 3,470) of samples and from a metastatic site in 49.8% (n = 3,457) of samples.
Anonymized germline variant calling was performed using a sequence analysis pipeline validated for clinical use in a CLIA-compliant laboratory performing clinical sequencing of patient tumours and matched normal blood specimens as a part of routine clinical care21. Germline variants with a population allele frequency of less than 2% in gnomAD exomes (v2.0.1) and genomes (v2.1.1) were assessed for pathogenicity following ACMG guidelines. Variants predicted by Variant Effect Predictor to have high-impact consequences were considered putative LoF variants. Variants were designated as pathogenic if classified as pathogenic or likely pathogenic in-house or by ClinVar or predicted to be LoF in a TSG. We excluded variants flagged as potentially arising from clonal haematopoiesis or circulating tumour cells as previously described61.
We manually reviewed variants with discordant interpretations between ClinVar and in-house classifications, as well as variants with conflicting interpretations in ClinVar. Novel LoF variants were considered pathogenic unless located in the terminal exon, where such variants were considered pathogenic only if the same gene harboured previously designated pathogenic LoF variants located further downstream in the coding sequence. Low-risk variants in CHEK2 such as c.470T > C were excluded from the analysis.
Population frequencies were obtained from gnomAD exomes (v2.0.1) and gnomAD genomes (v2.1.1). Variants were annotated for pathogenicity interpretations using ClinVar (accessed December 2024) and in-house classifications (as of August 2022) by expert clinical geneticists. Ensembl Variant Effect Predictor was used to annotate variants for predicted functional consequences.
The external cohort consisted of tumour samples from gBRCA2 primary tumours, which underwent whole-exome sequencing (WES) from samples collected at the University of Pennsylvania and Mayo Clinic5. Patients gave written informed consent for research use of germline DNA and tumour specimens under IRB-approved protocol at the University of Pennsylvania and Mayo Clinic. As previously described62, somatic tumour DNA was extracted from formalin-fixed, paraffin-embedded primary breast cancer specimens using standard laboratory deparaffinization, whereas germline DNA was extracted from whole blood or saliva. Tumour DNA libraries were prepared using the NEBNext formalin-fixed, paraffin-embedded repair mix and NEBNext Ultra II DNA library prep kit (New England Biolabs), per the manufacturer’s instructions. Germline DNA libraries were prepared using the NEBNext Ultra DNA library prep kit (New England Biolabs), per the manufacturer’s instructions. DNA libraries were pooled and hybridized using SureSelect Target Enrichment System for Illumina Multiplex Sequencing (Agilent) and associated protocols. For WES, tumour and germline libraries were hybridized to SureSelect All Exon v5, SureSelect All Exon v6+COSMIC and SureSelect All Exon v7 captures (Agilent). WES was performed using an Illumina HiSeq 4000, and targeted sequencing was performed using an Illumina NovaSeq 6000. All sequencing was performed with 150 paired-end reads by the University of Pennsylvania Next Generation Sequencing Core.
FASTQ files from WES and targeted sequencing were aligned to the hg19 build of the human genome using the Burrows–Wheeler Aligner (BWA v0.7.17-r1188)63. Various bam file processing operations were performed using Samtools/htslib/bcftools (v1.11). The resulting bam files were processed according to Genome Analysis Toolkit (GATK v3.7) best practices (picardtools v2.20.7).
Zygosity inference
We inferred somatic zygosity for all germline pathogenic variants using locus-specific copy number and ASCN inference utilizing FACETS as previously described64. All ASCN solutions (FACETS outputs) from tumours with germline pathogenic variants were manually reviewed to ensure that the optimal solution was selected. Purity estimates were similarly inferred from the FACETS. We incorporated ASCN, purity and variant allele frequency (VAF) into a previously described framework65, allowing statistical inference of heterozygous, biallelic (loss of WT allele) or loss of mutant allele. The zygosity of the germline variant was considered indeterminate and excluded from zygosity analyses if the: (1) variant was homozygous in the germline; (2) read depth of coverage in the normal blood specimen was less than 50; (3) FACETS-derived total and minor copy numbers were not evaluable at the corresponding locus; or (4) an optimal solution could not be identified, most commonly due to low tumour purity. Using these criteria, out of the 472 total cases with germline pathogenic variant in the genes of interest, somatic zygosity status was evaluable for 447 (94.7%) of tumours. We excluded cases where copy number analysis and VAF were suggestive of loss of mutant allele, rather than WT (n = 22, 4.6%) from further analyses.
To initially determine whether a given germline variant was in allelic imbalance in the corresponding tumour specimen, we evaluated consistency between observed somatic VAF and expected VAF. The latter value was calculated as a function of ASCN and purity as previously defined65. Germline variants were considered heterozygous if their observed VAF was either (1) consistent with the expected VAF (within its 95% binomial CI) given balanced heterozygosity (total copy number (TCN) and lesser copy number (LCN) of either 2 and 1 or 4 and 2 in diploid and genome doubled tumours, respectively), or (2) less than the lower bound of the 95% CI of the expected VAF corresponding to a TCN and LCN of 3 and 1, respectively, which was either single copy gain of the mutant or WT allele. Germline variants in allelic imbalance of any kind were those with an observed VAF that was either within or greater than the 95% CI of the expected VAF corresponding to a copy number state other than balanced heterozygosity. For allelically imbalanced germline variants, loss of the WT was determined as those with an observed VAF within the 95% CI (or greater than the lower bound of the 95% CI) of the expected VAF corresponding to a LCN equal to 0 (observed VAF is concordant with the expected VAF when the lesser allele has a copy number of 0).
Data anonymization
All patients (n = 5,881) consented to an IRB-approved protocol allowing analysis of somatic and clinical data (NCT01775072). A subset of 2,896 patients had additionally consented for identified analysis of germline variants under this protocol, whereas the remaining patients (n = 2,985) consented only to somatic analyses.
For analyses involving germline pathogenic variant status, we anonymized data into a unique anonymized ID (A-ID-#####), enabling germline calls to be conducted on matched normal samples for all patients regardless of germline consent status in accordance with MSK IRB guidelines for anonymized germline–somatic analyses, as previously described65,66. For all patients, data were binned to avoid unique clinical or somatic alteration values and thereby prevent re-identification65,66. In brief, for patients who did not provide consent for identified germline analysis, genomic data were anonymized with a deterministic one-way hash function. In these patients, germline variant calling was performed using the clinically validated pipeline described above67, and PFS and OS times were rounded to the nearest month. For analyses in which germline data were not required, a separate study-specific unique identifier (S-ID-#####) and fully anonymized clinical data were used for all the patients regardless of germline consent status. Continuous clinical (for example, time to progression) and somatic genomic data were therefore able to be used for analyses that did not incorporate germline data.
Clinical data annotation
For the MSK cohort, clinical data were obtained from the validated Breast Translational Program’s clinical database, which contains comprehensive, longitudinal information on patient demographics, pathology, treatment regimens and clinical outcomes4,39,68,69,70,71. Structured data were systematically curated using standardized clinical data annotation processes by a dedicated team of expert clinical data annotators and were maintained under rigorous quality-control procedures by experienced data managers. The data lock for the analysis was April 2025.
Each patient in the cohort was assigned a single receptor status. Recognizing potential intertumoural heterogeneity, we sought a unified definition as follows: (1) in cases in which any metastatic biopsy was sequenced, receptor status was defined by treating clinician interpretation at the time of assigned first-line treatment; (2) in cases in which only a primary tumour is sequenced, receptor status was defined by receptor status of the sequenced primary. We excluded cases in which sequencing was obtained for external consultation (and therefore, lacked clinical or pathological data, n = 310), cases where the diagnosis was ductal carcinoma in situ (DCIS), with no evidence of invasive breast cancer during the clinical course (n = 1) or cases in which multi-site sequencing demonstrated no intrapatient genomic overlap and multiple distinct receptor statuses consistent with multiple primary tumours (n = 4; Extended Data Fig. 1a). Using these definitions, the MSK cohort consisted of 3,703 patients (66.5%) with HR+/HER2– receptor status and 1,043 patients (18.7%) with triple-negative breast cancer, with the remainder (820, 14.7%) being HER2 amplified (HER2+).
Progression events were defined as (1) a radiographic or clinical disease progression prompting change in systemic therapy or recommendation for ablative local therapy directed at a site (or sites) of progressive disease; or (2) clinician assessment detailing radiographic and/or clinical progression, after which it was documented that the patient and physician decided to continue the same therapy post-progression. In such cases, the time of progression was defined as the date of documented progression rather than the date of therapy discontinuation.
Enrichment analysis
We compiled mutations, fusions and copy number alterations predicted to be functionally significant (oncogenic or likely oncogenic) by the OncoKB precision oncology variant database72. All putative RB1 homozygous deletions were manually re-reviewed. Cases in which the putative homozygous deletion spanned beyond the size of an amplicon (that is, an event that would be interpreted as incompatible with tumour survival) were not considered a functionally significant variant.
In cases in which a patient had multiple samples sequenced, we compiled the total somatic variants called from either the sequenced primary sample or all sequenced metastatic samples (omitting the primary), to avoid duplicate samples and to ensure that each set of variants was assigned either ‘primary’ or ‘metastasis’ as a covariate. For the purposes of this analysis, local recurrence samples were considered ‘primary’. Upon excluding samples with indeterminable receptor status (as described in the ‘clinical data annotation’ section), 5,566 patients were eligible for Firth-penalized logistic regression.
Receptor status was also defined on a ‘per-patient’ basis as described above. Genes with alterations in less than 2% of the cohort were omitted from the analysis. For each remaining combination of germline gene X and somatic gene Y, we performed a Firth-penalized logistic regression to account for the sparsity of the dataset. Receptor status and sample type (metastatic versus primary) were included as covariates. The analysis was also repeated for each receptor status subtype, as well as repeated for biallelic versus mono-allelic germline variants. The latter step was only performed for samples in which zygosity was evaluable (n = 5,516). Putative P values were adjusted for multiple hypothesis testing using the Benjamini–Hochberg method; q < 0.10 was deemed to be statistically significant.
ASCN definitions
In the MSK cohort, somatic LOH events of RB1 were defined based on manual FACETS review of pre-CDK4/6i treatment samples. Pre-CDK4/6i ASCN analysis was performed on samples collected before first-line CDK4/6i treatment or as part of a matched pre-treatment and post-treatment CDK4/6i pair. Of 922 patients meeting this criteria, 196 samples were excluded due to low purity or other technical limitations (such as ‘waterfalling’ artefact), or paucity of heterozygous single-nucleotide polymorphisms allowing for confident lesser copy number inference. RB1 LOH was defined as LCN = 0, irrespective of TCN, whereas heterozygous state was any LCN > 0.
In the PALOMA-3 cohort, baseline ctDNA samples from the palbociclib combination arm were sequenced using a 1,729 amplicon custom AmpliSeq panel, which included 119 single-nucleotide polymorphisms located within the RB1 gene. ctDNA-based LOH analysis was conducted using a bespoke pipeline48; RB1 LOH was defined as previously described73.
In the MSK cohort, we further separated the RB1 LOH (LCN = 0) group into (1) het loss, defined as a state with TCN = 1 and LCN = 0; and (2) other LOH, defined as a state with TCN > 1 and LCN = 0. Fraction genome altered was also calculated for each pre-treatment sample and defined as the fraction of log2 copy number variation (gain or loss) of more than 0.2 divided by the total size of the copy number profiled region.
HRD inference analysis
To study the implications of HRD on clinical outcomes and mechanism of resistance, we inferred HRD from two orthogonal methods, which have been validated for use with targeted next-generation sequencing data. IMPACT-HRD quantifies genomic scars associated with HRD by analysing ASCN alterations determined with the FACETS algorithm (v0.5.14) and computing those genomic scars with the impact-hrd package (https://github.com/mskcc/facets-suite/blob/master/R/copy-number-scores.R). All IMPACT-HRD assessments were completed using R (v4.1.2). In particular, three metrics were evaluated: number of telomeric allelic imbalances, large-scale transitions and losses of heterozygosity. The overall HRD phenotype is defined as the unweighted sum of these three metrics.
Clinical outcome analysis
We determined the association between genomic alterations and PFS with disease progression on therapy with CDK4/6i or patient death. Disease progression was defined as the date of the radiological study or clinical assessment that established progression of disease and prompted a change in systemic treatment, intervention with locally directed therapy (for example, radiation therapy), or otherwise an annotation in the chart documenting progression of disease. We categorized CDK4/6i regimens based on their ET partner (aromatase inhibitor versus selective oestrogen receptor degrader). Patients with ablation of only known sites of disease with radiotherapy or surgical resection before initiation of CDK4/6i therapy were excluded, as were patients who discontinued therapy due to toxicity within 2 weeks. When ET or CDK4/6i changed to another ET and CDK4/6i, respectively, for reasons other than disease progression (for example, toxicity, patient preference or insurance coverage), the time on successive regimens was combined to more accurately capture real-world PFS on the CDK4/6i + ET combination.
We used both univariate and multivariate Cox proportional hazard models (adjusted for ET partner (fulvestrant versus aromatase inhibitors) and treatment line, where applicable). For patients with multiple lines of therapy from the same class of treatment, only the first treatment line from that class that was started after the MSK-IMPACT biopsy was included in the analysis. For analyses pertaining to ASCN, fraction genome altered and whole-genome duplication were used as additional covariates. These are recognized poor prognostic factors and may be a confounding factor given increased likelihood for tumours with measures of copy number instability to harbour LOH of any specific region.
For OS analysis, we implemented a left-truncated model to account for the immortal time from diagnosis of metastatic disease (time zero) to enrolment on sequencing protocol. Similarly to the univariate analyses, we used univariate and multivariate Cox proportional hazard models. In addition to ET partner, age at metastatic diagnosis was also included as a covariate. We rejected the null hypotheses with a two-sided α = 0.05.
For the matched-pairs analysis, we included all patients with available paired pre-CDK4/6i and post-CDK4/6i sequencing data. Pre-CDK4/6i samples consisted exclusively of tumour specimens sequenced using MSK-IMPACT with available ASCN and zygosity assessment for RB1. Post-treatment samples included post-progression CDK4/6i tumour specimens sequenced using MSK-IMPACT as well as ctDNA sequenced using either MSK-ACCESS74 or Guardant360 (ref. 75). For analyses comparing RB1 heterozygous loss with other allelic configurations, we focused specifically on drivers of resistance rather than subclonal events. We therefore excluded post-treatment alterations in which the VAF was less than 0.30 of the maximum allele frequency of high-confidence variants present in the particular sample of interest.
For assessment of acquired tumour suppressor loss in the HRD versus BRCA2 versus non-HRD group, we considered tumour suppressor genes that have been implicated in CDK4/6i resistance: RB1 (refs. 35,39,76), PTEN77, LATS2 (ref. 39), FAT1 (ref. 39), TP53 (ref. 71), ARID1A78, LATS1 (ref. 39) and NF1 (ref. 79). We excluded samples in which the ASCN was not evaluable in all these genes of interest, or in which there was already a biallelic LoF of one of the genes predicted to confer immediate resistance (RB1, PTEN and NF1). We included patients with baseline TP53 loss, as it has been shown to facilitate cell cycle re-entry and is therefore associated with acquired resistance on an intermediate timescale71. We defined pre-treatment samples as either (1) gBRCA2, (2) non-BRCA2 HRD (either harbouring a germline variant in BRCA1, PALB2 or classified as HRD-positive by the HRD-IMPACT assay) versus (3) non-HRD. All clinical outcome analyses were conducted with R software (v4.5.1) and the survival and exact2x2 packages.
In vivo PDX models
Targeted sequencing of post-mortem and PDX studies
Post-mortem tissue samples were selected for DNA extraction, library preparation and targeted sequencing. Up to 30 mg frozen tissue was digested with 40 µl of proteinase K (600 mAU ml−1) in 360 µl Buffer ATL at 56 °C. Genomic DNA was isolated using the DNAeasy Blood & Tissue Kit (69504, Qiagen) according to the manufacturer’s protocol, including treatment with RNase A. DNA was eluted in 60 µl 0.5X Buffer AE heated to 55 °C.
After PicoGreen quantification and quality control using an Agilent BioAnalyzer, 100 ng of genomic DNA was used to prepare libraries using the KAPA Hyper Prep Kit (07961901001, Roche) with eight cycles of PCR. Of each barcoded library, 100–135 ng was captured by hybridization in a pool of 9 samples using the IMPACT assay (IDT), designed to capture all protein-coding exons and select introns of 505 commonly implicated oncogenes, tumour suppressor genes and members of pathways deemed actionable by targeted therapies. Captured pools were sequenced using an Illumina NovaSeq 6000 in PE100 run mode using the NovaSeq 6000 S4 Reagent Kit (200 cycles). All experiments were carried out at MSK’s Integrated Genomics Organization.
The demultiplexed FASTQ files from the post-mortem samples were aligned to the human genome reference GRCh37/hg19 using bwa mem (v0.7.17-r1188)63 and deduplicated using Picard MarkDuplicates (v2.21.8). Quality-control metrics of the alignments included (1) unique passing filter (PF)-aligned read pairs, (2) mean target coverage, (3) mean insert size, and (4) major or minor contamination.
Variant calling was performed using a previously described pipeline80. In brief, structural nucleotide variants were detected in the tumour–normal pairs using Mutect (v1.1.6)81, whereas indels were detected using a consensus of Varscan 2 (v2.4.6)82, Strelka (v2.9.10)83, Scalpel (v0.5.4)84 and Platypus (v0.8.1.2)85. Variants found with more than 0% global allele frequency in the 1000 Genomes database (phase III) or more than 0.01% across any population in the ExAC database (release 0.3.1) or that were covered by 10 reads in the tumour or 5 reads in the germline were filtered out. Variants for which the tumour variant allele fraction was more than five times than that of the normal variant allele fraction were filtered out. The aggregated set of variants identified in the tissues and xenografts were re-genotyped in all samples using SAMtools mpileup (v1.19.2)86. Copy number alterations were detected using Facets (v0.6.2)64. In addition, off-target reads were used to estimate log2 ratios using CNVkit (v0.9.8)87. Structural variants were detected using the consensus of Manta (v1.6.0)88, SvABA (v1.1.0)89 and GRIDSS (v2.13.2)90. The aggregated set of structural variants identified were re-genotyped in all samples using Paragraph (v2.3)91 and annotation of the structural variants was done using vcf2maf (v1.6.22; https://github.com/mskcc/vcf2maf) and AnnotSV (v3.5.3)92. Reversion mutations and structural variants affecting BRCA2 were further classified using aardvark (v0.35)93,94.
MSK PDXs (PDX-L and PDX-R)
Animal studies
Mouse studies were conducted through the MSK antitumour core facility in compliance with institutional guidelines under an Institutional Animal Care and Use Committee-approved protocol (MSK IRB 12-10-016). PDXs were established by implanting freshly collected autopsy samples from a patient in MSK’s Last Wish Program (MSK IRB 15-021), which enables patients to donate their bodies post-mortem for research. The samples were collected at MSK under approved IRB biospecimen protocols (MSK IRB 12-245 and 06-107).
Animals were maintained in accordance with the Guide for the Care and Use of Laboratory Animals in an AAALAC-accredited facility. All procedures outlined in the study were approved by the MSK Cancer Center Institutional Animal Care and Use Committee. Animals were housed in individually ventilated caging systems (Thoren Caging Systems), on autoclaved aspen chip bedding (PJ Murphy Forest Products) and were provided a γ-irradiated commercial diet (PicoLab Rodent Diet 20, 5053 LabDiet, PMI Nutrition International), and acidified water (pH 2.5–2.8) ad libitum. Mice were housed at a population density that ranged from 1 to 5 mice per cage in an environment providing a temperature of 21.1–22.2 °C (70–72 °F), 30–70% humidity, 10–15 fresh air exchanges hourly and a 12–12-h light–dark cycle (lights on, 06:00–18:00). For PDX studies, the tumours were expanded by serial subcutaneous transplantation.
In vivo studies
The 0.18 mg/90-day-release oestrogen pellets were implanted into 6-week-old female NSG mice 5 days before tumour implantation. When xenografts reached 100 mm3, mice were randomized to treatment arms of vehicle (saline), fulvestrant (200 mg kg−1 subcutaneous, twice weekly), ribociclib (75 mg kg−1 PO, 5 days per week) or combination therapy. Tumour size was measured twice a week. The animals were euthanized at the end of the experiment and tumours were collected for histological and biochemical analyses. The maximum allowed tumour size was 2,000 mm3. The sample size for PDX experiments was calculated based on previous experience with this model and drug response. No statistical method was used to predetermine sample size. The investigators were not blinded to allocation during experiments and outcome assessment.
For analysis, a linear mixed-effect model was used for comparing the growth curves between the treatment conditions. In detail, the model included the tumour volume as the dependent variable, individual mouse ID as a random intercept, and day, treatment and the interaction term between day and treatment as fixed effects. In PDX-R, the model comparing ribociclib to vehicle failed to converge. This was attributable to one extreme outlier mouse in each group at multiple timepoints identified using the Tukey method (tumour volume exceeding quantile 3 + 1.5 × interquartile range). Once excluding these outliers, the mixed effect model successfully converged and satisfied the model assumptions. A sensitivity analysis confirmed that the exclusion of these two mice did not alter the biological conclusion regarding the significance. Complete tumour raw data are included in the supplement.
Immunoblotting
Frozen PDX tumours were thawed on ice, cut into small pieces and placed in Lysing Matrix tubes (6910100, MP Biomedicals). SDS lysis buffer was added, and the sample was homogenized for 40 s and then boiled at 100 °C for 5 min. The supernatant was transferred and subjected to sonication at 40–45 amp for 30 s, repeated twice. After sonication, lysates were boiled again and then centrifuged. Protein concentration was quantified with BCA protein assay (23225, Thermo Scientific). Of protein, 25 µg was loaded for 3–8% Tris-acetate gel (NuPage) electrophoresis and transferred to nitrocellulose membranes. Blots were blocked with 5% non-fat milk in TBST for 1 h at room temperature and then incubated with primary antibody at 4 °C overnight. The following primary antibodies were used at 1:1,000 dilution: BRCA2 (123491, Abcam), pRB S780 (8180S, Cell Signaling Technology) and Rb (9313S, Cell Signaling Technology). Secondary antibodies conjugated with fluorescence (#926-32211 and #926-68070, LI-COR Bioscience) were incubated for 1 h at room temperature. Blots were imaged by Odyssey Clx Imaging System (LI-COR Biosciences); raw images are included in Supplementary Fig. 1.
Whole-genome sequencing of PDX samples
Post-CDK4/6i PDX samples were selected for DNA extraction, library preparation and whole-genome sequencing. The tissue samples were homogenized in 500 µl MagMAX DNA Cell and Tissue Extraction Buffer (A45469, Thermo Fisher) for up to 40 s and DNA from lysate was extracted using the MagMAX DNA Multi-Sample Ultra 2.0 Kit (A36570, Thermo Fisher) on the KingFisher Apex System (Thermo Fisher) according to the manufacturer’s protocol. The samples were eluted in 80 µl elution solution.
After PicoGreen quantification and quality control using an Agilent TapeStation, 500 ng of genomic DNA were sheared using a LE220-plus Focused-ultrasonicator (500569, Covaris) and sequencing libraries were prepared using the KAPA EvoPrep Kit (10212250702, Roche) with modifications. The libraries were subjected to a 0.5× size selection using aMPure XP beads (A63882, Beckman Coulter) after post-ligation cleanup. The libraries were not amplified by PCR and were pooled at equal volume. The samples were sequenced using an Illumina NovaSeq X in PE150 run mode using the NovaSeq X 25B Reagent Kit. All experiments were carried out at MSK’s Integrated Genomics Organization.
The demultiplexed FASTQ files were aligned to a chimeric genome reference comprising the human reference GRCh37/hg19 and the mouse GRCm38/mm10 using bwa mem (v0.7.17-r1188)63. Read pairs where at least one end (R1 and/or R2) had a primary alignment the mouse genome were filtered out and the remaining read pairs were re-aligned to the human reference GRCh37/hg19 as described above. Quality control of the alignments was done as previously described above in which the percent of mouse content was quantified as the number of unique PF read pairs aligned to the mouse genome relative to the total unique PF aligned read pairs. Copy number alterations were detected using Facets (v0.6.2)64. Structural variants were detected using the consensus of Manta (v1.6.0)88, SvABA (v1.1.0)89 and GRIDSS (v2.13.2)90. The aggregated set of structural variants identified in the xenografts were re-genotyped in all samples using Paragraph (v2.3)91, and annotation of the structural variants was done using vcf2maf (v1.6.22; https://github.com/mskcc/vcf2maf) and AnnotSV (v3.5.3)92.
Institut Curie PDXs
PDX-C (HBCx-118) models of ER+ MBC were obtained by engrafting biopsies from spinal bone metastases of patients with ER+ breast cancer progressing under ET. Specifically, this model was derived from a patient with ER+ MBC previously exposed to combination fluorouracil, epirubicin and cyclophosphamide (FEC) chemotherapy and tamoxifen in the adjuvant setting, and aromatase inhibitor and paclitaxel in the metastatic setting. The protocol was approved by the Institut Curie Hospital committee (Comité de Revue Institutionnel). Bone metastasis biopsies were engrafted with informed consent from the patient into the interscapular fat pad of female Swiss nude mice (Charles River Laboratories), which were maintained under specific pathogen-free conditions. Their care and housing were in accordance with institutional guidelines and the rules of the French Ethics Committee: CEEA-IC (Comité d’Ethique en matière d’expérimentation animale de l’Institut Curie, National registration number: #118). The project authorisation no. is 02163.02. The housing facility was kept at 22 °C (± 2 °C) with a relative humidity of 30–70%. The light–dark cycle was 12 h light–12 h dark.
For efficacy studies, tumour fragments were transplanted into female 8-week-old Swiss nude mice. When tumours reached a volume comprised between 100 and 200 mm3, xenografts were randomly assigned to the different treatment groups of vehicle and palbociclib 75 mg kg−1 PO 5 days per week. Tumour size was measured with a manual caliper twice per week. Tumour volumes were calculated as V = a × b2/2, a being the largest diameter, by the smallest. Tumour volumes were then reported to the initial volume as relative tumour volume. Means (and s.d.) of relative tumour volume in the same treatment group were calculated, and growth curves were established as a function of time. For each tumour, the percent change in volume was calculated as (Vf − V0/V0)/100, V0 being the initial volume (at the beginning of treatment) and Vf the final volume (at the end of treatment).
PDX-P1 and PDX-P2
All animal work was conducted according to AstraZeneca’s Global Bioethics Policy (https://www.astrazeneca.com/content/dam/az/Sustainability/Bioethics_Policy.pdf), in accordance with the PREPARE guidelines and reported in line with the ARRIVE guidelines.
Studies with PDX-P1 (ST4316B) were performed under contract with XenoStart at AAALAC-accredited facilities and performed in accordance with protocols approved by the START ‘Institutional Animal Care and Use Committee’ and AstraZeneca’s ‘Platform for Animal Research Tracking and External Relationships’ (PARTNER) group. Mice were acclimated for a minimum of 24 h and housed on irradiated corncob bedding (Teklad) in individual HEPA-ventilated cages (Sealsafe Plus, Techniplast USA) on a 12-h light–dark cycle at 21–23 °C and 40–60% humidity. Animals were fed water ad libitum (reverse osmosis, 2 ppm Cl2) and an irradiated standard rodent diet (Teklad). Xenografts were established by subcutaneous surgical implantation of an approximately 70 mg tumour fragment into the right flanks of 6–12-week-old female athymic nude animals. Tumours reached 0.15–0.3 cm3 before the animals were randomized into groups. Tumour volume (mm3) was calculated as width2 × length × 0.52.
Studies with PDX-P2 (HBCx-22) were performed under contract with Xentech under authorization by the ‘Direction Départementale de la Protection des Populations, Ministère de l’Agriculture et de l’Alimentation’, France and in accordance with protocols approved by Xentech along with AstraZeneca’s PARTNER group. Mice were delivered to the facility at least 7 days before the experiment for acclimatizing to environmental conditions. Mice were housed in polysulfone plastic (PSU) individually ventilated cages (213 mm width × 362 mm diameter × 185 mm height) bedded with sterilized and dust-free bedding cobs. Animals had controlled light–dark cycle (14-h circadian cycle of artificial light) at 20–24 °C and 40–75% humidity. Each mouse was offered a complete pellet diet (150-SP-25, SAFE) and filtered, sterilized tap water ad libitum throughout the study.
Xenografts were established by subcutaneous surgical implantation of approximately 20 mm3 into the flank of female athymic nude -Foxn1nu mice. Tumours reached 0.1–0.3 cm3 before animals were randomly assigned into treatment groups. Tumour volume (mm3) was calculated as [length × width2]/2.
Animals were randomized into treatment groups, saruparib 1 mg kg−1, camizestrant 10 mg kg−1 and palbociclib 50 mg kg−1 dosed PO daily, according to the tumour size criteria outlined above to obtain treatment arms with homogeneous geomean volumes. For both studies, tumours were measured twice weekly. Changes in tumour volume and growth inhibition were determined by bilateral Vernier caliper measurement (length × width). Length was the longest diameter across the tumour, and width was the corresponding perpendicular. Conscious animals were euthanized by cervical dislocation with secondary confirmation at the end of the study or for welfare condition. For analysis, two-way ANOVA followed by Sidak’s multiple comparisons test was used to compare tumour volumes up to day 35 (last day of evaluable tumour volume among all groups) between the treatment conditions saruparib and palbociclib.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.