Gene collection and functional classification
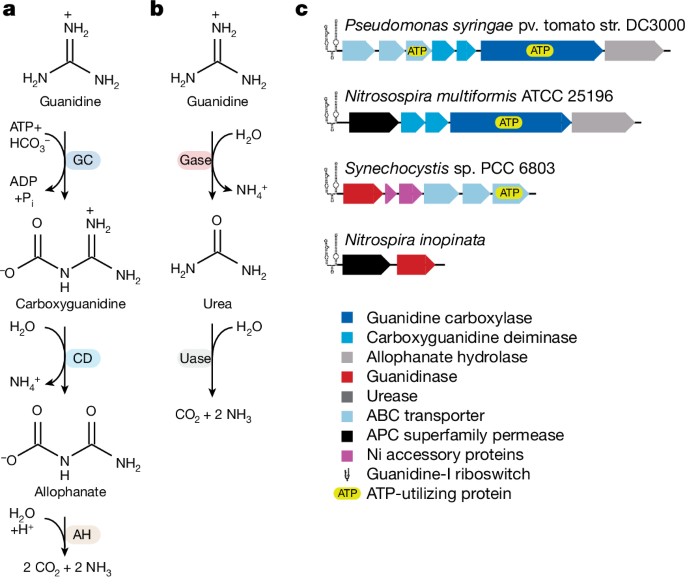
Supplementary Table 11 contains details for the collection and analyses parameters of each gene family and the following description is generalized. Predicted proteins from all publicly available genomes in GenBank as of 1 July 2022 (429,896 genomes) were screened with hmmsearch59 using âcollection HMMsâ for genes related to guanidine metabolism (Fig. 1). The resulting genes were used as query sequences against a combined HMM set of PFAMs, TIGRFAMS, NCBIFAMs and PANTHERFAMs for a set of acceptable âcross-check HMMsâ. Genes were further screened using specific e-value and coverage cut-offs. Cross-check HMM names were used to query UniProt and results were filtered for reviewed entries with evidence at the protein level to identify functionally characterized proteins and download them if not already present in the dataset. The portion of the protein sequence that was aligned to the âcollection HMMâ was extracted and clustered using usearch60 with specified -id and -query_cov values to identify centroids. HMM-based alignments of centroid sequences generated from the initial hmmsearch were used in FastTree61 to generate phylogenetic trees for each gene family of interest. For the APC superfamily permease and the allophanate hydrolase, all proteins that passed the e-value and coverage cut-offs were inferred to possess the expected function. Guanidinases were also required to possess threonine at N. inopinata position 105 (PF00491 HMM position 15), histidine at N. inopinata position 222 (PF00491 HMM position 134) and tryptophan at N. inopinata position 313 (PF00491 HMM position 223)18. Guanidine carboxylases were required to possess a conserved aspartic acid at K. lactis position 1,584 (TIGR02712 HMM position 956) and were further differentiated from urea carboxylase by having an aspartic acid at K. lactis position 1,330 (TIGR02712 HMM position 701)15. Carboxyguanidine deiminases were defined using the common ancestor of the two subunits (CgdA and CgdB) in the general tree for PF09347. This common ancestral node then gave rise to CgdA and CgdB as two monophyletic clades defined as such.
Riboswitch collection
Genomes from ammonia-oxidizing microorganisms were screened using infernal (v.1.1.3)62 using established RFAM covariance models for guanidine riboswitches I (RF00442), II (RF01068) and III (RF01763) and a model for the recently described guanidine IV riboswitch, which was constructed with infernal using âGGAM-1-curated.stoâ63. Scaffold IDs, coordinates and orientation were recorded and cross-referenced against gff files to identify downstream genes. A gene was considered to be under the control of a riboswitch if it was in the same orientation as the riboswitch and the 5â² end of the gene was within 1,000 nucleotides of the riboswitch. The inferred operon was then extended downstream until genes could be found with the opposite orientation.
Phylogenetic analyses
Ureohydrolase
HMM-based alignments of centroid sequences defined above were used in FastTree261 with the default parameters to generate a phylogenetic tree. The tree was midpoint rooted using the function midpoint() from the phangorn package, and functional clades were defined using the getMRCA() function within the ape package and visualized using the ggtree package in R.
Guanidinase
The most recent common ancestor of all guanidinases (as defined above) was identified in the ureohydrolase tree using the getMRCA() function and the descendant centroids were collected using the Descendants() function, both from the ape package64. All HMM alignments of ureohydrolases that were represented by centroids collected using the Descendants() function were additionally required to have covered PF00491 over 90% of its length reclustered using usearch (-id 0.9 -query_cov 0.9). The HMM-aligned portion of this sequence dataset was used to calculate phylogeny with IQ-TREE265, using the best model (LG+I+I+R5), and bipartition support was evaluated using ultrafast bootstraps. Logos for each resulting clade of guanidinases and close relatives were generated using the ggseqlogo package in R.
For co-phylogeny analyses of comammox guanidinases and ammonia monooxygenases, comammox genomes were screened for the presence of amoA and guanidinase genes. As most available genomes were MAGs, it was required that exactly one copy of each gene was identified per genome. This resulted in 54 genomes for analysis. The AlignTranslation and AlignSeqs functions from the Decipher package were used to align amoA nucleotide and guanidinase amino acid sequences, respectively. IQ-TREE2 was used to identify the best models (amoA, TPM3u+F+I+I+R3; guanidinase protein, LG+I+G4), calculate trees and evaluate bipartition support with ultrafast bootstraps. Trees were visualized in R using a combination of the cophylo function from the phytools package and the ggtree package.
Guanidine quantification
The following chemicals were purchased from Sigma-Aldrich: guanidine hydrochloride (â¥99%, G3272), benzoin (â¥99%, 8.01776), potassium hydroxide (â¥85%, 1.05033), ethanol (â¥99.8%, 02851), formic acid (â¥98%, 5.43804), β-mercaptoethanol (â¥99%, 8.05740), l-arginine (â¥99.5%, 11009) and sodium sulfite (â¥98%, 239321). Hydrogen chloride solution (32%, 20254.321) and acetonitrile (â¥99.9%, 20060.320) were purchased from VWR. 2-Methoxyethanol (â¥99.5%, 10582945) was purchased from Thermo Fisher Scientific. MilliQ water was obtained from a water purification system (0.071âµSâcmâ1; Elga Veolia, PURELAB Chorus). The derivatization protocol for guanidine was adapted from a previous study29. In brief, 150âµl of an aqueous solution potentially containing guanidine was cooled to 0â°C in a 0.5âml plastic tube (Eppendorf, Protein LoBind, 0030108434) and spiked with 75âµl of a benzoin solution in ethanol (4âmM), 75âµl of an aqueous solution containing both β-mercaptoethanol (0.1âM) and sodium sulfite (0.2âM), and 150âµl of an aqueous solution of potassium hydroxide (1.6âM). The resulting solution was mixed, heated in a bath of boiling water for 10âmin and cooled in an ice bath for 2âmin. Subsequently, 25âµl of an aqueous solution of hydrogen chloride (4.8âM) was added. The resulting solution was mixed and transferred to a 1.5âml plastic tube (Eppendorf, 0030120086) and centrifuged at 10,000g for 2âmin. Before analysis, the supernatant was diluted to obtain analyte concentrations in the optimal quantification range of the analytical instrument (that is, 0.05â5âμM). The predominant derivatization product (proposed structure in Supplementary Fig. 1c) was analysed using liquid chromatography (Agilent 1290 Infinity II) coupled to triple quadrupole mass spectrometry (Agilent, 6470) with a retention time of 3.73âmin. We used the InfinityLab Poroshell 120 Bonus-RP (Agilent, 2.7âµm, 2.1âÃâ150âmm) column for separation, an injection volume of 2âµl, a flow rate of 0.4âmlâminâ1, a column compartment temperature of 40â°C and the following eluents: aqueous (A): MilliQ water with 0.1% (v/v) formic acid; organic (B): acetonitrile with 0.1% (v/v) formic acid. The eluent gradient was as follows: 0â1.5âmin, 5% B; 1.5â4âmin, 5â61% B; 4â4.5âmin, 61â95% B; 4.5â7âmin, 95% B; 7â8âmin, 95-5% B; 8â10âmin, 5% B. The source parameters were set as follows: positive mode electrospray ionization; gas temperature, 250â°C; gas flow, 10âlâminâ1; nebulizer, 45âpsi; sheath gas temperature, 280â°C; sheath gas flow, 11âlâminâ1; capillary voltage, 3.5âkV; nozzle voltage, 0.5âkV. The following product ions of the derivatization product (m/z of parent ion: 252.2) were monitored: m/z: 182.1 (quantifier) and m/z: 104.1 (qualifier). The resulting chromatographs were integrated using MassHunter (Agilent, v.10.1). For absolute quantification, we used a series of guanidine solutions with a concentration range after dilution between 0.05 and 5âµM. For pure culture medium, activated sludge and soil extracts, calibration solutions were prepared in the respective matrix. Animal urine and faeces were quantified with calibration solutions in water. Accurate quantification for animal samples was confirmed by spiking 20âµM guanidine to an animal faeces sample with a recovery of >83%. Animal manure samples were freeze-dried, and subsamples were dispersed in 2âM KCl solution (1âml per 100âmg sample) and bead-beated for 15âmin in a Lysing matrix A tube (MPBiomedicals), then centrifuged at 20,000g for 15âmin. Wastewater treatment plant influent was quantified by standard addition. Calibration solutions were derivatized and analysed in the same way as and in parallel to the respective samples. For Orbitrap (high-resolution) MS analyses, we used liquid chromatography coupled to the Thermo QExactive mass spectrometer with the following parameters: positive electrospray ionization; capillary temperature, 275â°C; sheath gas, 15; aux gas, 10; sweep gas, 1; S-lens RF, 50.0; resolution, 140,000 (MS full-scan), 17,500 (MS/MS); NCE (stepped), 10,20,30. For growth experiment samples containing heavy-isotope-labelled guanidine, the total guanidine concentrations were inferred by assuming the measured isotopically unlabelled guanidine concentrations to correspond to 90% (we used 10% isotopically labelled guanidine).
Physiology experiments with N. inopinata and ammonia-oxidizing bacteria
The cells were grown in medium containing 54.4âmgâlâ1 KH2PO4, 74.4âmgâlâ1 KCl, 49.3âmgâlâ1 MgSO4·7 H2O, 584âmgâlâ1 NaCl, 147âmgâlâ1 CaCl2, 34.4âμgâlâ1 MnSO4·1H2O, 50âμgâlâ1 H3BO3, 70âμgâlâ1 ZnCl2, 72.6âμgâlâ1 Na2MoO4·2 H2O, 1âmgâlâ1 FeSO4·7 H2O, 20.0âμgâlâ1 CuCl2·2 H2O, 80âμgâlâ1 CoCl2·6 H2O, 3âμgâlâ1 Na2SeO3·5H2O, 4âμgâlâ1 Na2WO4·2H2O, 24âμgâlâ1 NiCl2·6 H2O and 0.5âmM pyruvate. The medium was buffered by addition of 4âmM HEPES, with the pH set to 8. For regular culture maintenance, cultures were kept in closed Schott bottles at 37â°C without shaking in the dark. When indicated, guanidine hydrochloride was added from a filter-sterilized 0.1âM stock solution to a final concentration of 50âμM.
For comparing guanidine utilization by pure cultures of N. inopinata and AOB, all strains were induced in 1âl batch cultures for 6âweeks with 0.5âmM ammonium and 1âµM guanidine fed weekly. Subsequently, the same amount of biomass per culture as determined using the Pierce BCA protein quantification kit (Thermo Fisher Scientific; calculated final concentration in the incubation, 10âµgâmlâ1) was collected, washed and resuspended in fresh medium in equal volumes and transferred to 96-well, flat bottom culture plates (Greiner Bio-One). In these plates the following incubations were done with either 50âµM guanidine only; 50âµM guanidine plus 150âµM ammonium; or 150âµM ammonium only for 14 days at 28â°C in the dark and without agitation (optimal growth conditions for the ammonia oxidizing organisms used, while 9â°C colder than the optimum for N. inopinata).
For growth experiments, N. inopinata pure culture cells pregrown on 10âµM guanidine and 0.5âmM ammonium (with weekly refeedings) for 1âmonth were collected by centrifugation (4,500g, 30âmin), washed with N-free medium three times and resuspended in fresh medium. Aliquots of 200âml were distributed into 250âml serum bottles. Aliquots used as dead controls were autoclaved (120â°C, 20âmin) before substrate additions. The following N substrates were added (always 150âµM N) to five replicate bottles each: (A) 15N-guanidine (10% 15N-guanidine hydrobromide, 90% guanidine hydrochloride); (B) guanidine (as guanidine hydrochloride); (C) 15N-guanidine (10% 15N-guanidine hydrobromide, 90% guanidine hydrochloride) and ammonium (each 150 µM N); (D) ammonium only; (E) no N addition (starved control); (F) dead (autoclaved) control with 15N-guanidine (10% 15N-guanidine hydrobromide, 90% guanidine hydrochloride). Moreover, all bottles received 13C-bicarbonate additions (13C-NaHCO3; 1âmM final concentration, 99% 13C) to detect chemolithoautotrophic growth and 0.5âmM sodium pyruvate as a reactive-oxygen-species scavenger66. Serum bottles were closed with sterile, HCl-cleaned blue butyl rubber stoppers (Chemglass) and incubated at 37â°C in the dark without agitation. Samples of 2âml for the determination of cell numbers (using qPCR) and of N-compound concentrations were taken with sterile syringes and needles and replaced with air every 1 to 14 days (frequent sampling in the beginning of the experiment, more spaced-out sampling after incubations containing ammonium were ended) over a time course of 126âdays (12âdays for treatments containing ammonium). Substrates were replenished after depletion. After 107âdays of incubation, 10âml samples were removed from treatments A, B, E and F, fixed with 3% formaldehyde (final concentration) for 30âmin at room temperature, filtered onto polycarbonate filters (0.2âµm pore size, GTTP, 40ânm gold sputtered), washed with sterile 1à PBS, dried and stored frozen until further use. Cells were visualized after staining with 4â²,6-diamidino-2-phenylindole (DAPI, 10âµgâmlâ1, 5âmin at room temperature) using a confocal laser-scanning microscope (inverted Leica TCS SP8X CLSM equipped with a 405ânm UV diode). At the end of the growth experiments, the absence of heterotrophic contaminants was confirmed by inoculation into heterotrophic growth medium (LB and TSY).
Ammonium, urea, nitrite and nitrate concentrations were measured by colorimetric protocols published previously67. In brief, combined ammonia and ammonium concentrations were determined using the indophenol blue method. Nitrite concentrations were measured spectrophotometrically using the Griess method after reacting with sulfanilamide and N-1-naphthyl-ethylenediamine dihydrochloride. Nitrate was measured by the same method after reduction to nitrite with vanadium chloride. Urea concentrations were measured using the thiosemicarbazide-diacetylmonoxime method68, according to a previous study18.
For quantification of N. inopinata cell numbers, qPCR was performed using the primers 515F/806R, targeting the V4 region of the 16S rRNA gene as described previously69,70. Standards were generated from purified PCR products generated from N. inopinata genomic DNA as template. The standards were quantified according to the Qubit dsDNA HS Assay Kit instructions. Standards containing 109 gene copies per µl were aliquoted and stored frozen at â20â°C until further use. Each standard aliquot was used and defrosted only once to freshly prepare tenfold serial dilutions (108â102 gene copies per µl). The qPCR assays were performed as follows: the frozen culture aliquots were four times freeze-thawed for cell disruption. A total of 0.25âµM of each primer was used in a mixture of 10âµl SYBR Green Supermix (Bio-Rad), 2âµl cell lysate or standard, and water in a final volume of 22âµl per reaction. The qPCR cycler (C1000-CFX96, Bio-Rad) settings were as follows: 95â°C for 15âmin; 40 cycles of 95â°C for 30âs, 50â°C for 1âmin and 72â°C for 45âs (plate read); and finishing with 72â°C for 2âmin and a melting curve performance from 40â°C to 95â°C with an increase of 0.5â°C every 5âs. The efficiencies of the standard curves had an average of 86% and an R2 of 0.999. Growth rates (division rates) were calculated as follows:
$$v({{\rm{d}}}^{-1})={\log }_{2}({N}_{i+1}/{N}_{i})/t$$
(1)
where v is the rate of division (dâ1), N is the qPCR determined cell number at timepoint iâ+â1 and i, and t is the time interval between time point iâ+â1 and i in days.
For visualization of stable N and C isotope assimilation into N. inopinata cells from the supplied 15N-guanidine and 13C-bicarbonate, gold-sputtered filters containing cells from two replicate bottles (Treatment A, replicate A1 and A2) and a natural abundance (NA) control were glued onto antimony-doped silicon wafers (7.1âÃâ7.1âÃâ0.11âmm, Active Business Company) using superglue (Loctide). NanoSIMS measurements were performed on the NanoSIMS 50L instrument (Cameca) at the Large-Instrument Facility for Environmental and Isotope Mass Spectrometry at the University of Vienna. Before image acquisition, each analysis area was preconditioned by sequence of high and extreme low ion impact energy (EXLIE) Cs+ depositions as follows: high energy (16âkeV) at 50âpA beam current to a fluence of 5âÃâ1014âionsâcmâ2; EXLIE (50âeV) at 400âpA beam current to a fluence of 5âÃâ1016âionsâcmâ2; high energy to an additional fluence of 2.5âÃâ1014âionsâcmâ2. Data were acquired as multilayer image stacks by sequential scanning with a finely focused primary Cs+ ion beam (approximately 80ânm probe size at a 2 pA beam current) over 45âÃââ45 μm2 areas with 512âÃâ512 pixel image resolution. The primary ion beam dwell time varied between 1âms (A1, 74 planes; NA, 50 planes) and 5âms (A2, 21 planes) per pixel per cycle. The detectors of the multicollection assembly were positioned to enable parallel detection of 12C2â, 12C13Câ, 12C14Nâ, 12C15Nâ, 31Pâ and 32Sâ secondary ions. Image data analysis was performed using the OpenMIMS ImageJ plugin (OpenMIMS v.3.0.5, ImageJ v.1.54f), where the acquired datasets were aligned, deadtime and QSA corrected, processed (for example, accumulation, stable isotope ratio calculation) and exported for visualization of 13C and 15N enrichment (as 13C and 15N atom%).
N. inopinata shotgun proteomics
For protein analysis, biomass was dissolved in lysis buffer (8âM urea, 2âM thiourea, 1âmM PMSF). Protein extraction was done by incubation at 95â°C, while shaking at 1,400ârpm for 5âmin. Subsequently, the samples were treated for 3âmin in an ultrasonication water bath (Elmasonic S30 H). To the cell suspension, 6.75âµl 25âmM 1,4 dithiothreitol (in 20âmM ammonium bicarbonate) was added and incubated for 1âh at 60â°C and 1,400ârpm shaking. Next, 150âµl 10âmM iodoacetamide (in 20âmM ammonium bicarbonate) was added and incubated for 30âmin at 37â°C with 1,400ârpm shaking in the dark. Finally, 200âµl of 20âmM ammonium bicarbonate was added and the protein lysates were proteolytically cleaved overnight at 37â°C with trypsin (2.5âµl of 0.1âµgâµlâ1 trypsin, Promega). The cleavage was stopped by adding 50âµl 10% formic acid. The peptide lysates were desalted using ZipTip μC18 tips (Merck Millipore). The peptide lysates were resuspended in 15âµl 0.1% formic acid and analysed using nanoliquid chromatographyâMS (UltiMate 3000 RSLCnano, Dionex, Thermo Fisher Scientific). MS analyses of eluted peptide lysates were performed on the Q Exactive HF mass spectrometer (Thermo Fisher Scientific) coupled with a TriVersa NanoMate (Advion). Peptide lysates were injected onto a trapping column (Acclaim PepMap 100 C18, 3âμm, nanoViper, 75âμmâÃâ2âcm, Thermo Fisher Scientific) with 5âμlâminâ1 by using 98% water/2% acetonitrile with 0.5% trifluoroacetic acid, and separated on an analytical column (Acclaim PepMap 100 C18, 3âμm, nanoViper, 75 μmâÃâ25âcm, Thermo Fisher Scientific) at a flow rate of 300 nlâminâ1. Mobile phase was 0.1% formic acid in water (A) and 80% acetonitrile/0.08% formic acid in water (B). Full MS spectra (350â1,550âm/z) were acquired in the Orbitrap at a resolution of 120,000 with automatic gain control target value of 3âÃâ106âions.
Acquired LCâMS data were analysed with the Proteome Discoverer (v.2.5, Thermo Fischer Scientific) using SEQUEST HT and INFERYS Rescoring. Protein identification was performed using a database constructed from predicted proteins of N. inopinata downloaded from MicroScope71 and common contaminating proteins. Searches were conducted with the following parameters: trypsin as enzyme specificity and two missed cleavages allowed. A peptide ion tolerance of 10âppm and an MS/MS tolerance of 0.02âDa were used. As modifications, oxidation (methionine) and carbamidomethylation (cysteine) were selected. Peptides that scored qâ>â1% based on a decoy database and with a peptide rank of 1 were considered identified. Differential expression of proteins was evaluated using the DEqMS72. Normalized spectral abundance factors were also calculated for visualization purposes only.
Heterologous expression and purification of N. inopinata guanidinase
The guanidinase gene of N. inopinata was amplified with self-designed, specific PCR primers which already contained the vector-specific linker overhangs for Gibson cloning (5â²-CTGGAAGTTCTGTTCCAGGGGCCCATGGCGAAAAAGAGAACGTACC-3â² and 5â²-CCCCAGAACATCAGGTTAATGGCGTCAGCGTTTCTTTCGATTGCC-3â²), using high-fidelity Phusion Plus PCR Master Mix (Thermo Fisher Scientific). The purified product was cloned into the pCoofy4 (pETM44; His6-MBP) expression vector by using the GeneArt Gibson Assembly EX Cloning Kit (Invitrogen) according to the manufacturerâs protocol. The sequence of the insert was verified by Sanger sequencing.
Cultures were grown at 37â°C in auto-induction ZYP-5052 medium73 supplemented with 0.5âμM, 20âμM, or 1âmM NiSO4 for 5âh before cooling down at 4â°C for 15âmin, followed by overnight expression at 20â°C. Cells were lysed in the presence of a protease inhibitor cocktail in 50âmM HEPES, 200âmM NaCl, 5% glycerol, pHâ7.4 using a cell disruptor (Constant Systems) and centrifuged at 4â°C and 45,000g for 30âmin. Guanidinase fused N-terminally to a His-MBP-tag was purified by affinity chromatography using MBPTrap HP columns (Cytiva). Subsequently, the His-MBP-tag was cleaved overnight with HRV-3C protease added at a mass ratio of protease to protein of 1:50. Guanidinase was further purified by MBPTrap HP columns (Cytiva), followed by size-exclusion chromatography on the HiLoad Superdex 200 26/600pg column (Cytiva) equilibrated with 20âmM HEPES, 200âmM NaCl, 5% glycerol, pHâ7.4. For the 20âμM nickel in expression batch, this nickel concentration was maintained in all buffers during purification.
The sample was concentrated to around 10âmgâmlâ1 by ultrafiltration by using Vivaspin centrifugal concentrators (Sartorius) and flash-frozen and stored at â80â°C. Protein identity and purity were analysed using SDSâPAGE.
Size-exclusion chromatography combined with multiangle light scattering
SEC-MALS was performed using a Superdex 200 increase 10/300 GL (Cytiva) operated at 20â°C on the 1260 Infinity HPLC system (Agilent Technologies) coupled to a miniDawn Treos MALS detector (Wyatt Technology). The samples were injected (80âμl at 1âmgâmlâ1) onto a column extensively equilibrated with 20âmM HEPES, 150âmM NaCl, pHâ7.4. Measurement was performed using BSA as a control. The protein concentration was measured with a RI-101 refractive index detector (Shodex) and the average molecular mass was calculated using the program Astra (Wyatt Technology). The first-order fit Zimm formalism was used for analysis of light-scattering data as a data process procedure in Astra, and a generic protein dn/dc value of 0.185âmlâgâ1 was used for guanidinase and BSA.
Protein T
m determination
The Prometheus NT.48 instrument (NanoTemper Technologies) was used to determine the melting temperatures (Tm). Before measurements, samples of the guanidinase expressed with 0.5âµM Ni2+ were centrifuged for 10âmin at 16,000g at 4â°C to remove any large aggregates. To identify the buffer/pH, at which the Tm of the protein was the highest, the protein was diluted using a DSF-buffer/pH screen containing different buffers and pH values74. The capillaries were filled with 10âμl of sample and placed onto the sample holder. A temperature gradient of 1â°Câminâ1 from 20 to 95â°C was applied and the intrinsic protein fluorescence at 330 and 350ânm was recorded. Data were processed using MoltenProt75, where the melting temperatures from the curves were estimated using the two-state reversible unfolding model.
MS for heterologous expression experiments
Protein identity and purity were verified by intact protein mass spectrometry. A total of 40âng of the sample was injected into a column on the LCâMS system: Dionex nano HPLC, Waters XBridge C4, flow rate 250âμlâminâ1 step gradient 12â40â80% ACN Synapt G2Si, resolution mode. Reconstruction of average mass was done with MaxEnt1software76.
Metal determination by ICP-MS
To quantify Ni2+ and Mn2+ concentrations of the purified guanidinase, the samples were acid-digested and measured using ICP-MS. For acid digestion, HCl 30% (Supelco Suprapur, 100318, Merck), HNO3 65% (3-fold subboiled, provided in analytically pure quality; 1.00441.1000, Merck) were used. H2O2 (31%, ROTIPURAN Ultra, HN69.1) was purchased from Carl Roth. Deionized water was produced with 0.075âµSâcmâ1 using an Elga Veolia, PURELAB Chorus 3 RO. 180âµl of the sample was pipetted into 7âml PFA vials (Savillex), corresponding to a total sample amount of between 2 and 2.5âmg. Subsequently, 0.5âml HCl and 1.5âml HNO3 were added. After closing the vials gas tight, they were heated to 120â°C on a hot plate (Savillex). The temperature was kept constant for 12âh. After the samples had cooled down to room temperature, a total of 500âµl of H2O2 was added in 50âµl steps. Vials were closed again and heated at 120â°C for 12âh. Subsequently, vials were opened and the samples were brought to dryness at 120â°C. After cooling, the digestions were dissolved in 2âml HNO3 and brought again to dryness at 140â°C. Finally, the digestions were dissolved in 1âml HNO3 and 2âml deionized water. Vials were closed and heated again at 120â°C for 12âh to ensure complete dissolution. The digestions were then quantitatively transferred to 15âml centrifuge tubes (polypropylene, metal free) and filled up to 10âml with deionized water. Twofold dilutions of the digestions were measured with an Agilent 7900 Single Quad ICP-MS instrument (Agilent Technologies) in no-gas mode. The operation parameters for the plasma were set to the following values: RF power: 1,550âW; RF matching, 1.80âV; sample depth, 10âmm; nebulizer gas flow, 0.8âlâminâ1; makeup/dilution gas, 0.4âlâminâ1. The parameters for data acquisition were as follows: acquisition mode, spectrum; sweeps/ replicate, 80; replicates, 3; integration time/mass, 0.1âsec. External calibration standards with an element concentration of 0.025 to 25âµgâlâ1 were used for quantification. The limit of quantification values achieved for Mn2+ and Ni2+ were â¤0.17âµgâlâ1 and â¤0.61âµgâlâ1, respectively. The limit of detection (LOD) was â¤0.05âµgâlâ1 for Mn2+ and â¤0.18âµgâlâ1 for Ni2+. The measured concentrations of the diluted digestions ranged from 1.9 to 21.6âµgâlâ1 for Mn2+ and from <LOD to 22.8âµgâlâ1 for Ni2+. To exclude any contamination by the buffer used, this was also digested and measured. Here the concentrations ranged from <LOD to 0.07âµgâlâ1 for Mn2+ and from <LOD to 0.29âµgâlâ1 for Ni2+. Given that the concentrations of metals in all of the analysed samples were either significantly above the limit of quantification or below the LOD, the presence of these metals in the buffer solution was deemed not to have a relevant impact on the overall results.
Protein crystallization
For initial screening, guanidinase expressed with 0.5âμM Ni2+ was concentrated to 12.3âmgâmlâ1 using the Amicon ultra centrifugal filter unit with 30âkDa MWCO and crystallized in MRC two-well crystallization plates with 50âµl of mother liquor set up using the TTPLabtech Mosquito pipetting robot system using the drop ratios 150ânl:200ânl and 200ânl:200ânl (protein:reservoir). Initial screens were performed using JCSGâ+âHT, Index Screen, Morpheus Screen, PACT Premier screen and Crystal Screen at room temperature. Several hits were obtained from Crystal Screen and the condition F3, containing 0.5âM (NH4)2SO4, 0.1âM Na3 citrate pHâ5.6 and 1âM Li2SO4 was used as a template for optimization screening by varying the (NH4)2SO4 and Li2SO4 concentrations. The best crystals were obtained at 1âM (NH4)2SO4 and 0.5âM to 0.7âM Li2SO4.
X-ray data collection, model building and refinement
Crystals were cryo-protected using 20% glycerol, flash-frozen in liquid nitrogen and diffraction datasets collected at beamline ID30B at the European Synchrotron Radiation Facility (ESRF, France) under cryogenic conditions. The collected datasets were processed with XDS and converted to the mtz file format using XDSCONV77. The phase problem was solved with Phaser-MR78, using its AlphaFold79 prediction as a search model. The structure was further refined in iterative cycles of the manual model building using COOT80 and maximum-likelihood refinement using the PHENIX software suite81. The final stages of refinement used the automated addition of hydrogens, and TLS refinement with one TLS group per chain. The models were validated with MolProbity82 and PDBREDO83. Figures were created using PyMOL (The PyMOL Molecular Graphics System, v.2.0, Schrödinger) (Supplementary Table 12). Anomalous datasets were collected at ID30B at a wavelength of 1.8929âà , close to the manganese anomalous scattering absorption edge, and at a wavelength of 1.4825âà , close to the nickel anomalous scattering absorption edge. The anomalous datasets were processed as described above and the obtained mtz files were refined using the finalized model obtained from the native dataset. The anomalous maps obtained from refinement were averaged using phenix.ncs_average supplying the refined pdb structure file and the corresponding anomalous map in ccp4 file format. Averaged anomalous maps were visualized using PyMol.
Substrate-dependent oxygen uptake measurements
Whole-cell substrate oxidation kinetics were determined from oxygen-uptake measurements as previously described34,48,84. Here oxygen-uptake measurements were performed using a microrespirometry (MR) system equipped with a four-channel MicroOptode meter (Opto-F4 UniAmp) and O2 MicroOptodes. Real-time O2 concentration monitoring was supported through SensorTrace Rate software (Unisense).
N. inopinata biomass was cultivated in batch cultures in the same growth medium as described above and ammonium (1âmM) or urea (0.5âmM) as sole substrates. Ammonium and guanidine were also used as co-substrates and here ammonium grown cultures (1âmM) were supplemented with guanidine (10â20âµM) around 12âh before MR experiments to induce the expression of the guanidine transporter and the guanidinase. In all cases, active N. inopinata biomass was collected (3,000g, 6âmin, 20â°C) from substrate replete cultures, washed and resuspended in identical but substrate-free medium, and incubated in a recirculating water bath (>30âmin, 37â°C). Samples were taken for chemical analysis to ensure the absence of detectable ammonium, nitrite, nitrate and urea before MR experiments. All chemical species were determined photometrically as described above.
MR experiments were conducted in a glass MR chamber (~2âml) containing a glass-coated magnetic stir bar, on an MR2 stirring rack (350ârpm), in a recirculating water bath (37â°C). MR chambers were overfilled with concentrated biomass to ensure the absence of a gaseous headspace, closed with an MR injection lid and submerged in the water bath. An O2 MicroOptode was inserted into each MR chamber and left to equilibrate (~1âh), before a stable background signal was determined (15â30âmin). The background rate of oxygen depletion was subtracted from all subsequent rate determinations in each MR chamber. A Hamilton syringe (10âμl; Hamilton) was used for all substrate (ammonium, urea, guanidine) injections. Both single- and multiple-trace oxygen uptake measurements were performed.
For single-trace measurements, a single-substrate injection was performed, and the oxygen uptake was recorded until complete substrate depletion in the presence of excess O2 (>30âμM O2). The single-injection scheme was used to determine the molar ratio of urea and guanidine consumed per O2. The whole-cell kinetics of N. inopinata with urea and guanidine as substrates, respectively, were performed with single-injection traces. Here a single injection of urea (~20âμM) or guanidine (~20âμM) into the MR chamber was performed. Moreover, the whole-cell kinetics of total ammonium oxidation in N. inopinata precultivated with urea (0.5âmM) or ammonium plus guanidine (1âmM and ~20âμM) was determined with single-trace measurements. Here a single injection of NH4Cl (~25âμM) was performed. In all cases, the experiments were halted after complete substrate depletion in the presence of excess O2 (>30âμM O2). Nitrate was the only detectable end product in all MR chambers used for whole-cell urea and guanidine kinetic calculations.
Multiple-trace measurements were used to determine the inhibitory effect of guanidine on the rate of maximum ammonium oxidation in N. inopinata. The maximum rate of ammonium oxidation was achieved with an initial injection of NH4Cl (250â500âμM). Subsequently, several injections of varying guanidine concentrations were performed, and discrete slopes of oxygen depletion were calculated after each injection (~2â5âmin).
In all cases, for both single and multiple injections, MR chamber contents were immediately centrifuged (19,000g, 15âmin, 20â°C) after the measurements and the cell pellets and supernatant were stored separately for protein and chemical analysis, respectively (â20â°C). For protein analysis, the total protein content was determined photometrically using the Pierce BCA Protein Assay Kit (Thermo Fisher Scientific). The chemical analyses (ammonium, nitrite, nitrate and urea) were performed as described above.
In vitro enzyme activity assay of guanidinase
Guanidine degradation by the heterologously expressed and purified guanidinase (in the presence of different Ni2+ concentrations; see above) was measured at 37â°C, pHâ7.5, in a buffer containing 20âmM Tris-HCl and 50âmM NaCl by measuring urea production over 25âmin of incubation. The measurements of the enzyme expressed in the presence of 1âmM Ni2+ were done in the presence of 1âmM Ni2+. Kinetics were calculated from measurements at 50, 100, 250, 500, 1,000, 2,500, 5,000, 10,000, 25,000, 50,000 and 100,000âμM guanidine starting concentrations. For screening alternative substrate use, the guanidinase expressed in the presence of 1âmM Ni2+ was used. Then, 10âmM of methylguanidine, agmatine, arginine, creatine, guanidinobutyrate and guanidinopropionate each were incubated with the purified guanidinase enzyme or BSA at 37â°C for 30âmin in three or six replicates. Guanidinase pH dependence was screened at 37â°C with incubations at pHâ5.5, 6, 6.5, 7, 7.5, 8, 8.5, 9, 9.5, 10, 10.5 and 11 (set by addition of HCl or NaOH). Temperature dependence was screened at pHâ7.5 with incubations at 14, 20, 28, 37, 46, 50, 55, 60, 65, 70, 80 and 90â°C. These incubations were done in triplicates.
Calculation of cellular substrate oxidation kinetic properties
The cellular kinetic properties of total ammonium, urea and guanidine oxidation were calculated from single-trace substrate-dependent oxygen uptake measurements. The substrate oxidation rates were calculated from oxygen uptake measurements using a substrate-to-oxygen consumption ratio. For total ammonium oxidation, a substrate-to-oxygen ratio of 1:2 was used. Single-trace experiments were used to confirm the substrate-to-oxygen ratio for urea (3.9â±â0.31, nâ=â3) and guanidine (6.17â±â0.24, nâ=â4) oxidation. Thus, for total urea oxidation and total guanidine oxidation, substrate-to-oxygen ratios of 1:4 and 1:6 were used, respectively. All substrate oxidation rates were normalized to total cellular protein in each MR chamber. In the case of total ammonium oxidation, the Km(app) for unprotonated NH3 was calculated based on the Km(app) for total ammonium, incubation temperature, pH and salinity85.
The cellular kinetic properties of total ammonium, urea and guanidine oxidation were determined with a MichaelisâMenten model fit to the data using equation (2) where V is the reaction rate (μM per mg protein per h), Vmax is the maximum reaction rate (μM per mg protein per h), S is the total substrate concentration (μM), and Km(app) is the reaction half saturation concentration (μM). An unconstrained nonlinear least-squares regression analysis was used to estimate the Km(app) and Vmax values86,87.
$$V=\left({V}_{\max }\times \left[S\right]\right)\times {\left({K}_{{\rm{m}}\left({\rm{app}}\right)}+\left[S\right]\right)}^{-1}$$
(2)
The reaction half-inhibition concentration for total ammonium oxidation (Ki, μM), inhibition by guanidine, was also determined. The Ki was determined graphically with a Dixon plot analysis88. Inverse total ammonium oxidation rates were plotted against total guanidine concentration. Total ammonium oxidation rates resulting in a linear trend were used for these analyses. Linear best fit trendlines from each biological replicate were used to determine intersection focal points and estimate Ki values. Furthermore, a linear regression of the percentage of the total ammonium oxidation rate at varying guanidine concentrations was used to determine the Ki.
The specific substrate affinity (ao; litres per g wet cells per h) of ammonium, urea and guanidine oxidation was calculated using equation (3). The factor of 5.7âg wet cell weight per g of protein was used32,48,89.
$${a}^{^\circ }=({V}_{\max }\times {5.7}^{-1})\times {{K}_{{\rm{m}}({\rm{app}})}}^{-1}$$
(3)
WWTP community structure analyses
The Ribe WWTP (GPS coordinates: 55.33, 8.74) has biological N and P removal (enhanced biological phosphorus removal) and treats municipal wastewater with 20% industrial contribution (organic loading) corresponding to a total of 25,000 person equivalents. It is designed with recirculation and has return sludge sidestream hydrolysis. It does not have primary settling. Suspended solids around the time of sampling were ~3.1âgâlâ1. The Haderslev WWTP (GPS coordinates: 55.25, 9.51) has biological N and P removal and treats municipal wastewater with 5% industrial contribution corresponding to a total of 100,000 person equivalents. It is designed with alternating conditions and includes side stream hydrolysis. It does not have primary settling. Suspended solids around the time of sampling were ~3.2âgâlâ1. The Klosterneuburg WWTP (GPS coordinates: 48.29, 16.34) treats municipal wastewater corresponding to a total of 50,000 person equivalents with a two-stage, biological hybrid process. Suspended solids around the time of sampling were ~4.4âgâlâ1.
For characterizing the community structure, amplicon sequencing of the V1 to V3 regions of bacterial 16S rRNA genes was performed on samples from the Ribe and Haderslev WWTPs from the MiDAS BioBank collection. Applied PCR primers were 27F (5â²-AGAGTTTGATCCTGGCTCAG-3â²) and 534R (5â²-ATTACCGCGGCTGCTGG-3â²) with barcodes and Illumina adapters (IDT). PCR reactions (25âμl) were run in duplicate for each sample, using 1à PCRBIO Ultra Mix (PCR Biosystems), 400ânM of both the forward and reverse primer, and 10âng template DNA. The PCR conditions were 95â°C for 2âmin; followed by 20 cycles of 95â°C for 20âs, 56â°C for 30âs and 72â°C for 60âs; and a final elongation step at 72â°C for 5âmin. The PCR products were purified using 0.8à CleanNGS beads and eluted in 25âµl nuclease-free water. The amplicon libraries were pooled separately in equimolar concentrations, diluted to 4ânM and paired-end sequenced (2âÃâ300âbp) on the Illumina MiSeq sequencer using v3 chemistry (Illumina). A 20% phage PhiX control library was added to mitigate low-diversity library effects. The forward and reverse sequence reads were merged using the software usearch60 with the -fastq_mergepairs command, filtered to remove phiX sequences using usearch -filter_phix and quality filtered using usearch -fastq_filter with parameter -fastq_maxee set to 1.0. Dereplication was performed by usearch -fastx_uniques with the option -sizeout, and amplicon sequence variants (ASVs) were resolved using the usearch -unoise3 command. An ASV table was created by mapping the quality-filtered reads to the ASVs using the usearch -otutab command with the -zotus and -strand plus options. Taxonomy was assigned to ASVs using the usearch -sintax command with the parameters -strand both and -sintax_cutoff 0.8. The absence of comammox organisms in the sample from Klosterneuburg used for guanidine degradation measurements was confirmed by PCR using comammox clade A and clade B specific primer sets38. Ribe and Haderslev sample DNA in the same concentration were used as positive controls.
Substrate incubation experiments with biomass from WWTPs
Activated sludge samples were collected from the aerated tanks of the Ribe and Haderslev WWTPs on 22 October 2021. Four litres of sludge from each WWTP were scooped into large sterile plastic bottles. The samples were transported to the laboratory on the same day, and were stored in the dark at ambient temperature, that is, ranging from 4 to 10â°C, until the incubations were started. The incubations with sludge from Ribe were started on the same day as collection, and the incubation with samples from Haderslev were started on the day after collection. Before each incubation, the sludge was diluted approximately 1:4 as follows: the sludge was allowed to completely settle (1âh), then 1.5âl of the clear supernatant was gently collected to a new sterile flask without disturbing the flocs and, finally, 0.5âl of the remaining sludge was fully resuspended and added to the 1.5âl of supernatant. Well-mixed aliquots of 100âml of the diluted sludge were then distributed to 200âml sterile glass microcosms and covered with aluminium foil to enable gas exchange with the atmosphere. Substrates were added to the following final concentrations: guanidine, 50âµM; ammonia, 150âµM; and urea, 75âµM. These different concentrations were chosen to account for the number of amino groups among the molecules. No substrate controls were also included. The samples were incubated at 23â°C with shaking at 100ârpm. All substrate and control treatments were performed in triplicate. Microcosms were subsampled immediately before and after initial substrate additions at T0. Additional subsamples from the Ribe incubation series were taken at 3, 6, 12 and 24âh. Additional subsamples from the Haderslev incubation series were taken at 2.5, 4, 8, 16, 24 and 48âh. Subsamples for metatranscriptomics were immediately flash-frozen with liquid-N2 and stored at â80â°C until processing. Parallel samples (1âml) for chemical analyses were centrifuged at 12,000g for 5âmin, and the supernatant was taken and frozen immediately at â80â°C.
RNA extraction and purification
Total nucleic acids were extracted from activated sludge samples (500âµl), which were thawed on ice and centrifuged (5âmin, maximum speed, 4â°C), using the RNeasy PowerMicrobiome Kit (Qiagen) according to the manufacturerâs instructions with the addition of phenol:chloroform:isoamyl alcohol (25:25:1) and β-mercaptoethanol (10âμlâmlâ1 final concentration). Bead beating (40âs at 6âmâsâ1, four times with 2âmin interval on ice) on the Fastprep FP120 (MP Biomedicals) system was performed for cell lysis instead of vortexing to improve lysis of bacteria with rigid cell walls. The total nucleic acid extracts were subjected to DNase treatment to remove DNA contaminants using the TURBO DNA-free kit (Invitrogen), and further cleaned up and concentrated with RNAclean XP beads (Beckman Coulter) before rRNA depletion. The integrity and quality of the purified total RNA were assessed on a Tapestation 2200 (Agilent) with the Agilent RNA ScreenTape (Agilent) system, and the concentration was measured using the Qubit RNA BR Assay Kit (Thermo Fisher Scientific). The average RNA integrity number was above 7.0 for all of the samples.
rRNA depletion, library preparation and sequencing
Total RNA was rRNA-depleted using the NEBNext rRNA Depletion Kit for Bacteria (New England Biolabs) with 100â300âng total RNA as input. The NEBNext Ultra II Directional RNA Library Prep Kit (New England Biolabs) was used to prepare cDNA sequencing libraries according to the manufacturerâs instructions. The libraries were pooled in equimolar concentration and 2.0ânM was sequenced on an S4 flow cell on the NovaSeq 6000 platform (Illumina) using the v1.5 300 cycle kit (Illumina, 20012863).
Identification of differentially transcribed genes
rRNA-depleted reads were adapter-screened, quality-filtered and mapped to published MAGs using bbmap v.38.92. Adapter removal and quality filtering was conducted using bbduk (ktrim=r k=21 mink=11 hdist=2 minlen=119 qtrim=r trimq=15). Metatranscriptomic reads from WWTP Ribe were mapped to genome accession GCA_016722055.1 and reads from WWTP Haderslev were mapped to genome accession GCA_016712165.1, which were the dominant comammox MAGs in the respective WWTP37. Both mappings were carried out using bbmap (minid=0.98 idfilter=0.98 ambiguous=toss pairedonly=t killbadpairs=t mappedonly=t bamscript=bs.sh) to produce bamfiles. Counts for each gene were calculated using bedtools coverage (-counts) using BAM files from bbmap and GFF files downloaded from GenBank for each genome. Counts for each coding gene were examined for potential outliers, which identified MBK8278324.1 and MBK9947797.1 as potentially misannotated small RNAs and were removed from subsequent calculations. Differential transcription was evaluated by treating different timepoints as replicates and comparing treatments as factors using DESeq290. TPM was calculated and used for visualization purposes only.
Soil incubations
Soil was collected from a long-term fertilization experiment managed by the Austrian Agency for Health and Food Safety located at the Ritzlhof field experiment (48°â11â²â17.9â²â²âN 14°â15â²â16.5â²â²âE) in May 2023. The soil is classified as a Cambisol and has been fertilized since 1991 with solid cattle manure at an application rate of 525âkg N per ha per year91. Soil incubations were conducted in 125âml Wheaton bottles capped with grey butyl stoppers. In brief, 30âg soil was added to each replicate (nâ=â3) bottle, and amended with 820âµl water, ammonium, guanidine or ammonium + guanidine for a final concentration of 30âµg N per g dry-weight soil. Soils were incubated at 23â°C and sampled at 0, 1, 2, 3, 5, 7, 12 and 27 days. Acetylene (0.02%, v/v) was used to inhibit all lithotrophic ammonia oxidation. Acetylene was supplied by adding 0.3âml of 10% acetylene gas to sealed bottles. Bottles were opened every 1â3 days, and acetylene was resupplied. For chemical analyses, around 2âg soil was extracted in water and 2âM KCl and extracts were frozen at â20â°C until analysis. Nitrate and nitrite were quantified in water extracts and ammonium, urea, and guanidine were quantified in KCl extracts as described above. Approximately 1âg soil was sampled for molecular analysis and was frozen at â80â°C until analysis. DNA extracts were performed using the ZymoBIOMICS DNA/RNA Miniprep Kit according to the manufacturerâs instructions. AOB, AOA and comammox clade A and B amoA qPCRs were carried out as previously described38,92,93.
Statistics
Statistical analysis on chemical, protein and qPCR data from physiological experiments and WWTP sample incubations were performed using two-tailed t-tests in SigmaPlot v.14.5 and R. No statistical methods were used to predetermine sample size, and blinding and randomization of samples were not used.
Inclusion and ethics statement
All collaborators on this study fulfil the criteria for authorship required by Nature journals, they have been included as authors as their work was essential in designing and performing the study. The roles and responsibilities were agreed among collaborators ahead of the research. No living animals or animal-derived material were used in this study, except dropped animal manure and urine. Animals were not forced to excrete.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.