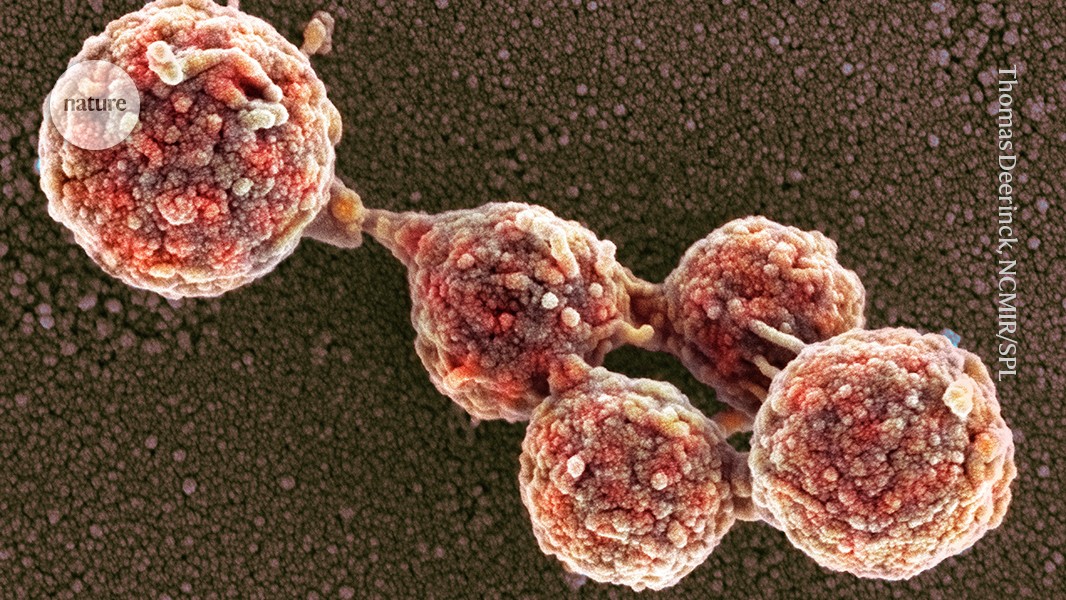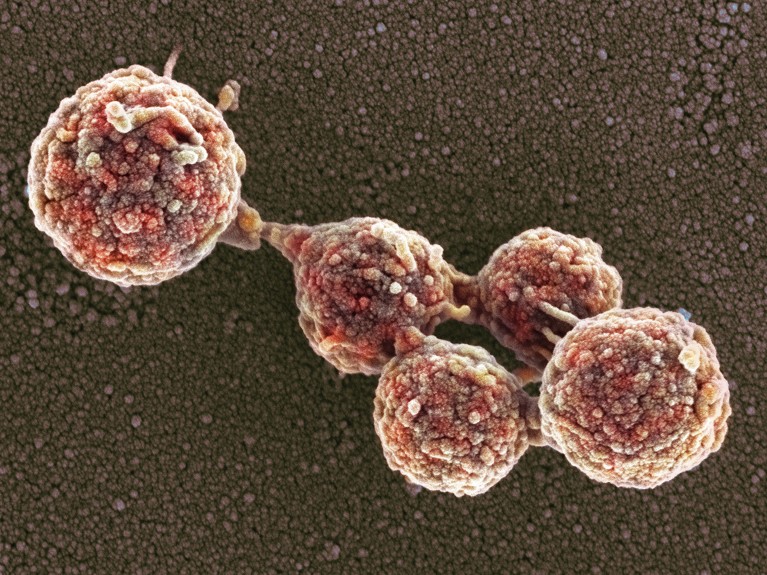
These Mycoplasma capricolum bacterial cells have absorbed a genome engineered from a closely related bacterium, Mycoplasma mycoides.Credit: Thomas Deerinck, NCMIR/Science Photo Library
Researchers have resurrected ‘dead’ bacterial cells by replacing their defunct DNA with the working genome of another species.
AI can write genomes — how long until it creates synthetic life?
The feat — reported on the preprint server bioRxiv this month1 — could boost efforts to re-engineer microbial life by moving entire genomes into bacteria to imbue them with useful properties, such as making drugs or biofuels.
Such genome transfers, including the one that gave rise to these ‘zombie cells’, have so far been accomplished only between species within a single bacterial class. But if researchers can routinely make zombies from other bacteria, the approach could be used to test engineered genomes from more commonly studied species, such as the laboratory staple Escherichia coli.
“For me, this paper represents a significant step forward for genome engineering in synthetic biology,” says Olivier Borkowski, a synthetic biologist at the French National Research Institute for Agriculture, Food and Environment (INRAE) and Paris-Saclay University.
Dawn of the dead
More than 15 years ago, researchers chemically synthesized the 1.1-million base-pair genome of the bacterium Mycoplasma mycoides and transplanted it into living cells of the closely related species Mycoplasma capricolum2, creating what they called the first synthetic cell.
‘Minimal’ cell raises stakes in race to harness synthetic life
The team, which included some researchers from the latest study, added a gene to the synthetic M. mycoides genome that imparted resistance to the antibiotic tetracycline. This meant that if the researchers transplanted the genome into M. capricolum and then grew those recipient cells in the presence of tetracycline, the cells would survive only if they had successfully absorbed the synthetic genome — a sign that the test had worked.
A 2016 study successfully transplanted genomes between species in the same class of bacteria as Mycoplasma, called Mollicutes3. But efforts to transplant genomes more widely have failed, says John Glass, a synthetic biologist at the J. Craig Venter Institute (JCVI) in La Jolla, California, and an author of both the new preprint and the synthetic-cell study. Apparent successes with other bacteria turned out to be false positives because the genomes of the recipient cells were able to incorporate antibiotic-resistance genes through a process called homologous recombination. That meant that the recipient cells would survive even if they didn’t absorb the entire donor genome. (M. capricolum didn’t give false positives in the experiments 15 years ago because it doesn’t have this recombination ability.)
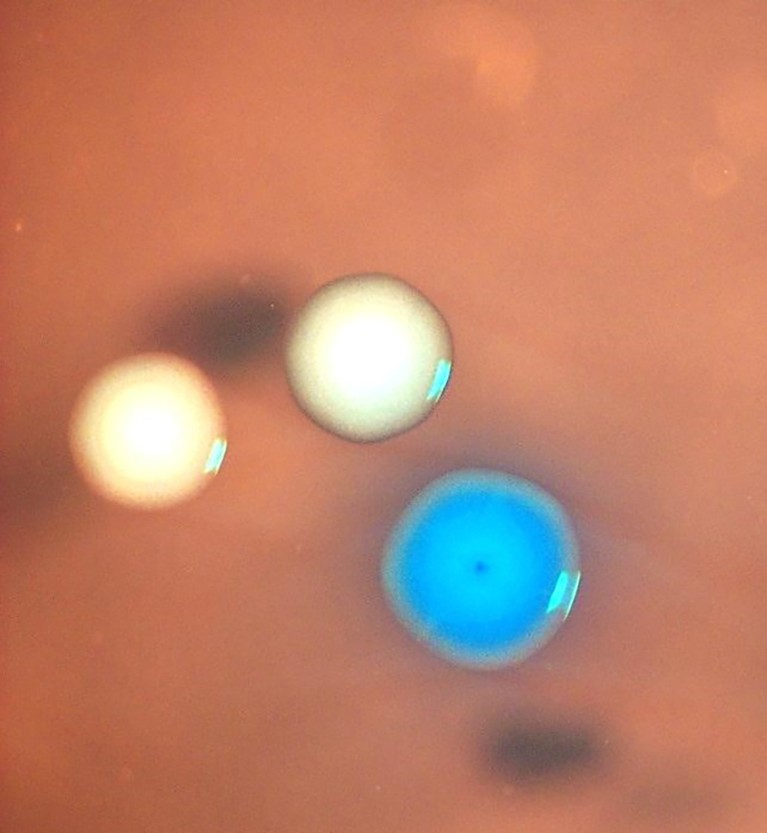
In their latest experiment, JCVI researchers killed M. capricolum cells and then resurrected them by inserting a synthetic genome from another bacterial species. The blue colony here is one that’s expressing that foreign genome.Credit: Nacyra Assad-Garcia
In search of a way to transplant genomes without having to worry generally about false positives, Glass and his colleagues reported in their preprint this month inactivating the genomes of recipient cells, leaving the cells unable to replicate, or functionally dead. This also prevents the recipient genomes from incorporating foreign DNA such as resistance genes through recombination. The team created these ‘dead’ M. capricolum cells by treating them with a DNA-damaging chemotherapy drug called mitomycin C.
When engineered M. mycoides genomes were transferred into M. capricolum cells treated with mitomycin C, a small fraction of the recipient cells survived. “The cell is destined to die, but we give it life,” says co-author Zumra Peksaglam Seidel, a synthetic biologist at the JCVI. She and her colleagues call these survivors ‘zombie cells’ in their preprint, which has not been peer-reviewed.