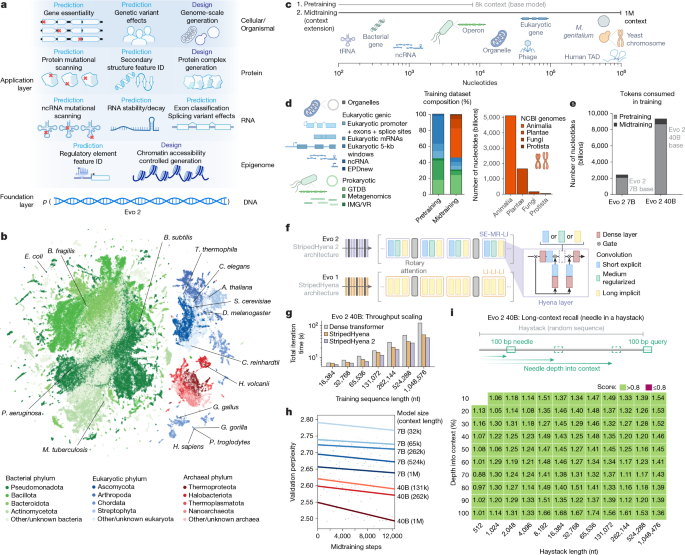
Nguyen, E. et al. Sequence modeling and design from molecular to genome scale with Evo. Science 386, eado9336 (2024).
Merchant, A. T., King, S. H., Nguyen, E. & Hie, B. L. Semantic design of functional de novo genes from a genomic language model. Nature 649, 749–758 (2026).
Avsec, Ž et al. Effective gene expression prediction from sequence by integrating long-range interactions. Nat. Methods 18, 1196–1203 (2021).
Linder, J., Srivastava, D., Yuan, H., Agarwal, V. & Kelley, D. R. Predicting RNA-seq coverage from DNA sequence as a unifying model of gene regulation. Nat. Genet. 57, 949–961 (2025).
Ku, J. et al. Systems and algorithms for convolutional multi-hybrid language models at scale. Preprint at https://doi.org/10.48550/arXiv.2503.01868 (2025).
Vaswani, A. et al. Attention is all you need. In Adv. Neural Information Processing Systems Vol. 30 (eds Guyon, I. et al.) (NIPS, 2017).
Kaplan, J. et al. Scaling laws for neural language models. Preprint at https://doi.org/10.48550/arXiv.2001.08361 (2020).
Radford, A. et al. Language models are unsupervised multitask learners. OpenAI Blog 1, 9 (2019).
Gao, T., Wettig, A., Yen, H. & Chen, D. How to train long-context language models (effectively). In Proc. 63rd Annual Meeting of the Association for Computational Linguistics 1, 7376–7399 (ACL, 2025).
Dubey, A. et al. The Llama 3 herd of models. Preprint at https://doi.org/10.48550/arXiv.2407.21783 (2024).
Liu, S. J. et al. In vivo perturb-seq of cancer and microenvironment cells dissects oncologic drivers and radiotherapy responses in glioblastoma. Genome Biol. 25, 256 (2024).
Poli, M. et al. Hyena hierarchy: towards larger convolutional language models. In Proc. 40th International Conference on Machine Learning (eds Karuse, A. et al.) 28043–28078 (2023).
Poli, M. et al. Mechanistic design and scaling of hybrid architectures. In Proc. 41st International Conference on Machine Learning 235, 40908–40950 (2024); https://proceedings.mlr.press/v235/poli24a.html.
Meier, J. et al. Language models enable zero-shot prediction of the effects of mutations on protein function. Preprint at bioRxiv https://doi.org/10.1101/2021.07.09.450648 (2021).
Notin, P. et al. ProteinGym: large-scale benchmarks for protein design and fitness prediction. Adv. Neural Inf. Process. Syst. 36, 64331–64379 (2023).
Benegas, G., Albors, C., Aw, A. J., Ye, C. & Song, Y. S. A DNA language model based on multispecies alignment predicts the effects of genome-wide variants. Nat. Biotechnol. 43, 1960–1965 (2025).
Shine, J. & Dalgarno, L. The 3′-terminal sequence of Escherichia coli 16S ribosomal RNA: complementarity to nonsense triplets and ribosome binding sites. Proc. Natl Acad. Sci. USA 71, 1342–1346 (1974).
Kozak, M. The scanning model for translation: an update. J. Cell Biol. 108, 229–241 (1989).
Nijkamp, E., Ruffolo, J. A., Weinstein, E. N., Naik, N. & Madani, A. ProGen2: exploring the boundaries of protein language models. Cell Syst. 14, 968–978 (2022).
Li, F.-Z., Amini, A. P., Yue, Y., Yang, K. K. & Lu, A. X. Feature reuse and scaling: understanding transfer learning with protein language models. In Proc. 41st International Conference on Machine Learning 235, 27351–27375 (2024).
Weinstein, E. N., Amin, A. N., Frazer, J. & Marks, D. Non-identifiability and the blessings of misspecification in models of molecular fitness. In Adv. Neural Information Processing Systems https://proceedings.neurips.cc/paper_files/paper/2022/file/247e592848391fe01f153f179c595090-Paper-Conference.pdf (2022).
Dalla-torre, H. et al. Nucleotide Transformer: building and evaluating robust foundation models for human genomics. Nat. Methods 22, 287–297 (2024).
Stanke, M., Steinkamp, R., Waack, S. & Morgenstern, B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 32, W309–W312 (2004).
de Almeida, B. P. et al. SegmentNT: annotating the genome at single-nucleotide resolution with DNA foundation models. Nat. Methods 22, 2301–2315 (2025).
Ji, Y., Zhou, Z., Liu, H. & Davuluri, R. V. DNABERT: pre-trained Bidirectional Encoder Representations from Transformers model for DNA-language in genome. Bioinformatics 37, 2112–2120 (2021).
Findlay, G. M. et al. Accurate classification of BRCA1 variants with saturation genome editing. Nature 562, 217–222 (2018).
Huang, H. et al. Functional evaluation and clinical classification of BRCA2 variants. Nature 638, 528–537 (2025).
Patel, A. et al. DART-Eval: a comprehensive DNA language model evaluation benchmark on regulatory DNA. Neural Inf. Process. Syst. 37, 62024–62061 (2024).
Cunningham, H., Ewart, A., Smith, L. R., Huben, R. & Sharkey, L. Sparse autoencoders find highly interpretable features in language models. Preprint at https://doi.org/10.48550/arXiv.2309.08600 (2023).
Bricken, T. et al. Towards monosemanticity: decomposing language models with dictionary learning. Transformer Circuits Thread https://transformer-circuits.pub/2023/monosemantic-features (2023).
Templeton, A. et al. Scaling monosemanticity: extracting interpretable features from Claude 3 Sonnet. Transformer Circuits Thread https://transformer-circuits.pub/2024/scaling-monosemanticity/ (2024).
Bussmann, B., Leask, P. & Nanda, N. BatchTopK Sparse Autoencoders. Preprint at https://doi.org/10.48550/arXiv.2412.06410 (2024).
Camargo, A. et al. Identification of mobile genetic elements with geNomad. Nat. Biotechnol. 42, 1303–1312 (2023).
Vorontsov, I. E. et al. HOCOMOCO in 2024: a rebuild of the curated collection of binding models for human and mouse transcription factors. Nucleic Acids Res. 52, D154–D163 (2024).
Gupta, S., Stamatoyannopoulos, J. A., Bailey, T. L. & Noble, W. S. Quantifying similarity between motifs. Genome Biol. 8, R24 (2007).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cells 38, 576–589 (2010).
Sandoval-Velasco, M. et al. Three-dimensional genome architecture persists in a 52,000-year-old woolly mammoth skin sample. Cell 187, 3541–3562.e51 (2023).
Meng, G., Li, Y., Yang, C. & Liu, S. MitoZ: a toolkit for animal mitochondrial genome assembly, annotation and visualization. Nucleic Acids Res. 47, e63 (2019).
Gibson, D. G. et al. Complete chemical synthesis, assembly, and cloning of a Mycoplasma genitalium genome. Science 319, 1215–1220 (2008).
Karr, J. R. et al. A whole-cell computational model predicts phenotype from genotype. Cell 150, 389–401 (2012).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Fredens, J. et al. Total synthesis of Escherichia coli with a recoded genome. Nature 569, 514–518 (2019).
Li, Y. et al. Competition-level code generation with AlphaCode. Science 378, 1092–1097 (2022).
Brown, B. et al. Large language monkeys: scaling inference compute with repeated sampling. Preprint at https://doi.org/10.48550/arXiv.2407.21787 (2024).
Allis, C. D. & Jenuwein, T. The molecular hallmarks of epigenetic control. Nat. Rev. Genet. 17, 487–500 (2016).
Schreiber, J., Lu, Y. Y. & Noble, W. S. Ledidi: Designing genomic edits that induce functional activity. Preprint at bioRxiv https://doi.org/10.1101/2020.05.21.109686 (2020).
Linder, J. & Seelig, G. Fast activation maximization for molecular sequence design. BMC Bioinformatics 22, 510 (2020).
Zrimec, J. et al. Controlling gene expression with deep generative design of regulatory DNA. Nat. Commun. 13, 5099 (2022).
de Almeida, B. P. et al. Targeted design of synthetic enhancers for selected tissues in the Drosophila embryo. Nature 626, 207–211 (2023).
DaSilva, L. F. et al. DNA-diffusion: leveraging generative models for controlling chromatin accessibility and gene expression via synthetic regulatory elements. Nat. Genet. 58, 180–194 (2026).
Sarkar, A. et al. Designing DNA with tunable regulatory activity using score-entropy discrete diffusion. Preprint at bioRxiv https://doi.org/10.1101/2024.05.23.595630 (2024).
Dauparas, J. et al. Robust deep learning-based protein sequence design using ProteinMPNN. Science 378, 49–56 (2022).
Verkuil, R. et al. Language models generalize beyond natural proteins. Preprint at bioRxiv https://doi.org/10.1101/2022.12.21.521521 (2022).
Bloomfield, D. et al. AI and biosecurity: The need for governance. Science 385, 831–833 (2024).
Pathak, A. K. et al. Pervasive ancestry bias in variant effect predictors. Preprint at bioRxiv https://doi.org/10.1101/2024.05.20.594987 (2025).
Schubach, M., Maass, T., Nazaretyan, L., Röner, S. & Kircher, M. CADD v1.7: using protein language models, regulatory CNNs and other nucleotide-level scores to improve genome-wide variant predictions. Nucleic Acids Res. 52, D1143–D1154 (2024).
Cheng, J. et al. Accurate proteome-wide missense variant effect prediction with AlphaMissense. Science 381, eadg7492 (2023).
Pampari, A. et al. ChromBPNet: bias factorized, base-resolution deep learning models of chromatin accessibility reveal cis-regulatory sequence syntax, transcription factor footprints and regulatory variants. Preprint at bioRxiv https://doi.org/10.1101/2024.12.25.630221 (2025).
Durrant, M. G. et al. Bridge RNAs direct programmable recombination of target and donor DNA. Nature 630, 984–993 (2024).