Overview of study recruitment
Participants in this study were recruited from the customer base of 23andMe. Participants provided informed consent and volunteered to participate in the research online, under a protocol approved by the external Association for the Accreditation of Human Research Protection Programs-accredited Salus Institutional Review Board (https://www.versiticlinicaltrials.org/salusirb). Participants were included in the analysis on the basis of consent status verified at the time data analyses were initiated.
The 23andMe GLP1 survey was launched to research participants in August 2024. The survey aimed to capture participants’ experiences with GLP1 receptor agonist medication, and was targeted to 23andMe participants who had previously responded in the affirmative to the question, ‘Have you ever taken prescription medications to help you lose weight?’. The survey included questions regarding drug brand, dosing regimen, time on treatment, efficacy (including pre-treatment weight and weight on treatment), and side effects, as well as reasons for pursuing or stopping GLP1 treatment. We focused the survey and subsequent analysis on primarily six drug varieties; Ozempic, Wegovy, compounded semaglutide, Mounjaro, Zepbound and compounded tirzepatide, the first three of which represent variations of semaglutide, and the last three represent variations in tirzepatide. A full list of survey questions can be found in Supplementary Table 24.
Phenotype definitions
Using the information derived from the surveys, we defined phenotypes that aimed to capture aspects of drug efficacy and side effects. We defined our efficacy phenotype as the contrast between pre-treatment BMI to post-treatment BMI (or current BMI, if treatment is on-going). In general, for study participants who reported taking more than one GLP1 medication, we selected the GLP1 medication that they reported taking for the longest period of time. Specifically, we defined a percentage BMI change phenotype as:
$${{\rm{\Delta BMI}}}_{ \% }=100({\mathrm{BMI}}_{2}-{\mathrm{BMI}}_{1})/{\mathrm{BMI}}_{1}$$
where BMI1 and BMI2 represent pre-treatment and post-treatment BMI, respectively, measured in weight in kilograms per height in metres squared. We applied quality control filters to people with weight less than 36 kg or greater than 181 kg, height less than 1.39 m or greater than 2.06 m, BMI less than 14 kg m−2 or greater than 70 kg m−2, or age less than 18 years. In aggregate, these initial filters removed 80 people (0.29%). Inspection of the ΔBMI% phenotype revealed a heavy tailed distribution, so we further quality controlled the ΔBMI% phenotype to remove outlier participants with BMI changes above 20% or below −45% (Extended Data Fig. 10). The ΔBMI% estimates were set to missing for participants who did not pass quality control.
To enable genetic associations to be interpreted in units of weight rather than ΔBMI%, we also defined a corresponding Δweight phenotype, defined as the change in weight from baseline in kilograms. We note that, because adult height is treated as constant during the treatment window, the percentage change in BMI (ΔBMI%) is mathematically identical to the percentage change in weight (Δweight%).
For the side effect phenotypes, we defined separate case–control phenotypes for each side effect recorded in the survey, contrasting those who self-rated their side effects as moderate or severe (cases) to those who self-rated their side effects as mild or non-existent (controls). As before, for study participants who reported taking more than one GLP1 medication, we selected the GLP1 medication that they reported taking for the longest period of time.
We further defined phenotypes to represent covariates, specifically for drug type (semaglutide = 1 versus tirzepatide = 0), dosage and days on treatment. For the dosage phenotype, we used the reported most recent weekly dosage in milligrams; this was either the final dose or the current dose for people still taking medication.
Comparison of self-report and EHR data
As part of the 23andMe experience, research participants are offered the opportunity to share EHR information collected on their Apple iPhone devices. Specifically, the Apple Health application enables connection to healthcare providers for the purposes of sharing EHR information with third parties through Apple HealthKit (https://developer.apple.com/documentation/healthkit). 23andMe research participants can elect to share their EHR information for research purposes. We used these data to perform comparisons with the self-report survey data. Full details of comparison analyses are provided in Supplementary Information.
Non-genetic predictors of BMI loss
To analyse the dependence of achieved BMI loss on non-genetic factors such as drug type, dosage and time on treatment, we fit the following model:
$$\begin{array}{c}{{\rm{\Delta BMI}}}_{ \% }\sim \mathrm{age}+\mathrm{sex}+{\mathrm{BMI}}_{1}+\mathrm{drugType}+\mathrm{dose}+{\mathrm{days}}_{\mathrm{treat}}\\ \,+\,\mathrm{drugType}:\mathrm{dose}+\mathrm{drugType}:{\mathrm{days}}_{\mathrm{treat}}\\ \,+\,\mathrm{dose}:{\mathrm{days}}_{\mathrm{treat}}+\mathrm{dose}:{\mathrm{days}}_{\mathrm{treat}}:\mathrm{drugType}\end{array}$$
(1)
where ‘drugType’ is an indicator variable that equals 1 for individuals using semaglutide and 0 for tirzepatide, ‘dose’ represents the dose in milligrams, daystreat represents the total days on the relevant drug and ‘:’ represents an interaction term between two or more variables. Note that semaglutide and tirzepatide typically have different standard dosing levels, which is handled in the regression model by the ‘drugType:dose’ interaction term.
Genotyping and SNP imputation
DNA extraction and genotyping were performed on saliva samples by Clinical Laboratory Improvement Amendments-certified and College of American Pathologists-accredited clinical laboratories of Laboratory Corporation of America. Samples were genotyped on one of five genotyping platforms. The V1 and V2 platforms were variants of the Illumina HumanHap550 BeadChip and contained a total of about 560,000 SNPs, including about 25,000 custom SNPs selected by 23andMe. The V3 platform was based on the Illumina OmniExpress BeadChip and contained a total of about 950,000 SNPs and custom content to improve the overlap with our V2 array. The V4 platform was a fully custom array of about 950,000 SNPs and included a lower redundancy subset of V2 and V3 SNPs with additional coverage of lower-frequency coding variation. The V5 platform was based on the Illumina Global Screening Array, consisting of approximately 654,000 preselected SNPs and approximately 50,000 custom content variants. Participant genotype data were imputed against a reference panel composed of data from the Haplotype Reference Consortium34 and augmented with additional sequences to boost imputation performance (Supplementary Information).
Association testing
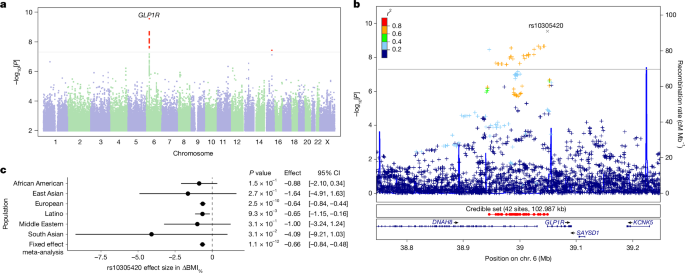
We performed a GWAS of ΔBMI% in people of European ancestry using methods that have been described previously35. In brief, unrelated participants were included in the GWAS analyses on the basis of European ancestry as determined by a genetic ancestry classification algorithm36. The GWAS was performed including covariates as described in equation 1 above, with the addition of five genetic principal components to account for fine-scale genetic ancestry, and indicator variables to account for variation in the genotyping platform. Among 21,822 people of European ancestry, we required participants to have complete data needed to construct the target phenotype and GWAS covariates (that is, data available for pre-treatment weight, post-treatment weight, drug type, dosage, time on treatment and factors such as age, sex and height), resulting in 18,488 participants. Finally, participants were filtered on relatedness such that no two people shared more than 700 cM identity by descent37, which corresponds approximately to the minimal expected sharing between first cousins in an outbred population, resulting in a final GWAS sample size of 15,237. An equivalent procedure was used for a GWAS of side effect phenotypes. For the purposes of testing drug-specific associations, we repeated the GWAS procedure for the semaglutide and tirzepatide-treated populations separately, removing the drug-type covariate and interaction terms as appropriate. All GWASs were adjusted for inflation using genomic control, with the inflation factor being no more than 1.035 in all phenotypes.
Given the smaller sample sizes available in non-European populations, we did not perform genome-wide association testing in these populations, and instead focused analyses on variants discovered as associated in the European GWAS. For these variants, we tested for association in non-European populations following a similar approach to that described above.
Replication
We performed replication of the identified efficacy association in the All of Us cohort38, using Controlled Tier Dataset v.8. We extracted genomic data, EHR data and a drug code referring to either semaglutide or tirzepatide from 9,579 participants. After filtering to retain participants with information regarding pre-treatment and post-treatment BMI and genotype data passing quality control, we obtained 4,889 participants, of which 3,948 had complete data when incorporating covariates akin to those used in the GWAS. For the replication analysis, we tested for association between the EHR-derived ΔBMI% and the genotype, including covariates. We repeated the replication analysis having performed mean-imputation of missing drug dose data, allowing a larger sample size of 4,855 to be analysed.
We also attempted replication analysis in the UK Biobank cohort, although the available data predate the availability of semaglutide or tirzepatide, and hence relied on earlier variants of GLP1 receptor agonists. Full details of the replication analysis methodology is provided in Supplementary Information.
Genetic and non-genetic risk modelling
To construct combined genetic and non-genetic models of ΔBMI% and risk of treatment-related side effects, we selected treatment, clinical, demographic, disease diagnosis and genetic variables as predictors. In addition to the covariates included in the GWAS, we also included years of education as a proxy for socio-economic status, and binary indicators of previous disease diagnosis for T2D, hypertension and non-alcoholic fatty liver disease. All continuous predictor variables were standardized before modelling to allow for the comparison of effect sizes.
We used a linear multi-variable model to fit ΔBMI%. Given the binary nature of side effect phenotype definitions, we fitted multi-variable logistic regression models (equivalent to a generalized linear model with a binomial family and logit link function). The dataset was partitioned randomly into training (70% of the sample) and held out test (30%) sets, with the test set being used to assess model performance. Further details are outlined in Supplementary Information.
Model performance of efficacy was further assessed by applying the model derived from our self-report data in a sample of 642 people who had provided HealthKit EHR data but had not completed the GLP1 survey, and hence were not used in the construction of the model. To replicate the situation where efficacy predictions are made before treatment, we assumed the dose, treatment duration and drug type variables were unknown, and imputed these values in the model to an arbitrary constant value for all participants.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.