Serial EM reconstructions
EM reconstructions were based on a single animal. We previously imaged and aligned a 770 × 750 × 53-μm3 volume of lobule V of the mouse cerebellum for EM reconstructions composed of 1,176 45-nm-thick parasagittal sections. We used automated image segmentation to generate neuron boundaries36. We used the neuron segmentation to reconstruct ten CFs that were identified based on their characteristic morphology such as their large axons projecting into the molecular layer and running along individual PCs. All of these CFs were reconstructed in their entirety within the molecular layer, and the dendritic arbours of the associated PCs were contained within the EM volume.
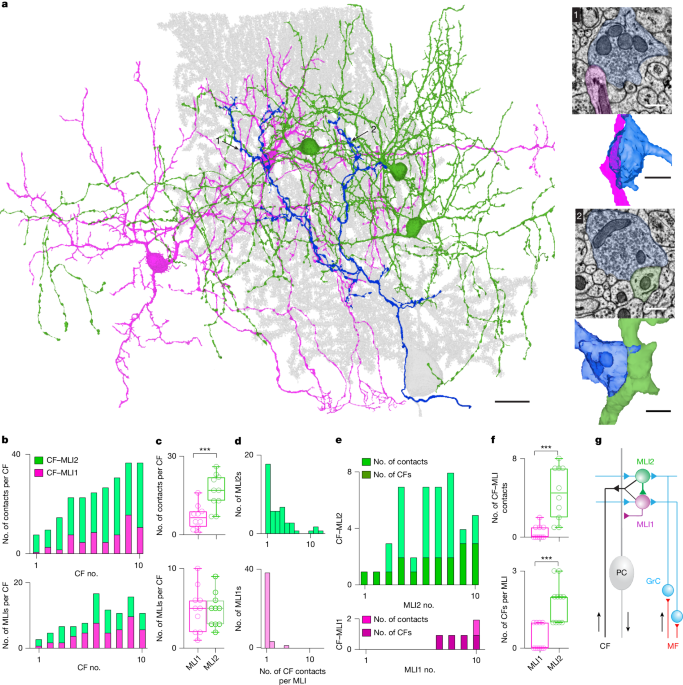
CF spillover contacts onto MLIs were identified by the physical contact of the CF bouton to an MLI dendrite. Vesicles are present in these boutons but they were not clustered in proximity to presynaptic active zones and PSDs were not associated with these contact sites. MLI contacts were analysed for ten CFs and subtyped into MLI1 or MLI2 on the basis of previous identification methods8. The surface areas of CF contacts onto MLIs were determined by tracing the contact area using annotation tools in Neuroglancer, retrieving their three-dimensional (3D) coordinates and through MATLAB calculating the 3D surface area. Parallel fibre synapses onto MLIs were determined by their small-diameter boutons and completely parallel morphology which was traced through the depth of the dataset.
CF–MLI1–PC and CF–MLI2–MLI1–PC synapses onto local and neighbouring PCs were identified using automated synapse prediction47. The neighbouring PC was determined to be the PC immediately adjacent to a PC that has an identified CF. A total of 608 CF–MLI1–PC synapses were made onto local PCs and 388 onto neighbouring PCs whereas a total of 2,799 CF–MLI2–MLI1–PC synapses were made onto local PCs and 2,583 were onto neighbouring PCs. Pathway analysis was done using Python. Reconstructions were used to estimate the fraction of GrC–MLI contacts that are influenced by CF contacts (Extended Data Table 1).
Slice experiments
Animal procedures were performed in accordance with the National Institutes of Health and Animal Care and Use Committee guidelines and protocols approved by the Harvard Medical School Standing Committee on Animals. C57BL/6 mice were obtained from Charles River Laboratories. Animals of either sex were randomly selected for experiments. Animals were housed on a normal light–dark cycle with an ambient temperature of 18–23 °C with 40–60% humidity.
Acute parasagittal slices (220-μm thick) were prepared from postnatal day (P)28–45 C57BL/6 mice. Mice were anaesthetized with an intraperitoneal injection of 100 mg kg−1 ketamine and 10 mg kg−1 xylazine and perfused transcardially with an ice-cold solution containing (in mM): 110 choline chloride, 7 MgCl2, 2.5 KCl, 1.25 NaH2PO4, 0.5 CaCl2, 25 glucose, 11.6 sodium ascorbate, 3.1 sodium pyruvate and 25 NaHCO3, equilibrated with 95% O2 and 5% CO2. Slices were cut in the same solution, and then transferred to artificial cerebrospinal fluid (ACSF) containing (in mM): 125 NaCl, 26 NaHCO3, 1.25 NaH2PO4, 2.5 KCl, 1 MgCl2, 1.5 CaCl2 and 25 glucose, equilibrated with 95% O2 and 5% CO2. As indicated, some experiments were performed in elevated external calcium (2.5 mM). Following incubation at 34 °C for 30 min, the slices were kept up to 6 h at room temperature until recording.
Recordings were performed at 32 °C with an internal solution containing (in mM): 35 CsF, 110 CsCl, 10 HEPES, 10 EGTA and 2 QX-314 (pH adjusted to 7.2 with CsOH, osmolarity adjusted to 300 mOsm kg−1). Visually guided whole-cell recordings were obtained with patch pipettes of ∼3-MΩ resistance pulled from borosilicate capillary glass (BF150-86-10, Sutter Instrument). Electrophysiology data were acquired using a Multiclamp 700B amplifier (Axon Instruments), digitized at 20 kHz and filtered at 4 kHz using Igor Pro (Wavemetrics) running mafPC (courtesy of M. A. Xu-Friedman). Acquisition and analysis of slice electrophysiological data were performed using custom routines written in Igor Pro (Wavemetrics). The following receptor antagonists were added to the ACSF solution to block GABAergic and glycinergic synaptic currents (in μM): 10 SR95531 (gabazine), 1.5 CGP and 1 strychnine. All drugs were purchased from Abcam and Tocris. No power analysis or other statistical methods were used to pre-determine sample sizes. Sample sizes were similar to previous publications.
We recorded MLIs in voltage clamp at −65 mV. Recordings were made from lobules IV–V of the vermis. We recorded from MLIs in the inner two-thirds of the molecular layer and determined the identity of MLI1s and MLI2s by their characteristic electrical properties as previously described8. MLI1s and MLI2s are molecularly and functionally distinct interneuron classes intermingled in the molecular layer8,17. MLI2s have relatively uniform properties regardless of their location in the molecular layer, whereas MLI1s comprise a continuous molecular gradient spatially distributed from nearest to the PC layer to the top of the molecular layer. MLI1s and MLI2s identified by cell fills and post hoc in situ hybridization exhibited distinct electrical properties17. MLI1s express connexin 36, they are electrically coupled to each other and spontaneous action potentials in neighbouring MLI1s produce spikelets, whereas MLI2s do not express connexin 36, they are not electrically coupled to other cells and spikelets are not present17. We classified MLIs with spikelets as MLI1s. Although spikelets are present in most MLI1s, a lack of spikelets is insufficient to identify MLI2s because spikelets in MLI1s require intact spontaneous firing in neighbouring MLI1s, which can be disrupted by poor slice health or by slicing damaging MLIs near the surface of the slice. MLI1s have input resistances ranging from ∼200 MΩ near the PC layer to up to ∼1 GΩ near the top of the molecular layer, whereas most MLI2s have resistances of greater than 1 GΩ (refs. 8,17). We found that resistances of both MLI1s and MLI2s were slightly higher with our caesium-based internal solution than with the potassium-based internal solutions used in previous studies. MLI2s were identified by a lack of spikelets and input resistances of greater than 1 GΩ. Experiments were not performed on neurons that had resistances in the 500-MΩ to 1-GΩ range and lacked spikelets (fewer than 5% of all MLIs), because they could not be unambiguously categorized based on electrical properties alone. We avoided cells in the upper third of the molecular layer where MLI1s tend to have higher input resistances and more MLI1s lack spikelets. This approach yields high-confidence classification of MLI1s and MLI2s based on passive electrical properties in real time without needing post hoc in situ hybridization, but it excludes some MLI1s that lack spikelets, excludes a small number of MLI2s with intermediate resistances and excludes MLIs in the upper third of the molecular layer.
We used a theta glass stimulus electrode to stimulate CFs in the GrC layer with pairs of stimuli (350 µs in duration, 50-ms interstimulus interval, 10–100 μA, 10-s intertrial interval) at different locations in an area around 50 µm into the GrC layer and around 100 µm on each side of the soma of the MLI being recorded. We recorded more than ten sites per cell in search of inputs that were all-or-none and with marked PPR (PPR < 0.6). We then reduced the stimulus intensity to threshold to evaluate the all-or-none nature of the response and contamination from GrC inputs. Stimulation at the threshold for CF activation stochastically evokes successes and failures, and slight increases in stimulus intensity eliminate failures. This indicates it is a single all-or-none input. To evaluate whether the CF input was isolated, failure trials were examined for GrC responses. Traces shown are averages of ten trials. For analysis of the kinetics, we calculated the 20–80 rise and decay times. We bath-applied TBOA (50 μM) while stimulating CF inputs with single stimuli every 10 s. The access resistance and leak current of the MLIs were monitored continuously. Traces shown are averages of ten trials for baseline and in the presence of TBOA. Parallel fibres were stimulated in the molecular layer with pairs of stimuli (350 µs in duration, 140–250 μA, 20-ms interstimulus interval, 5-s intertrial interval) within about 100 µm of the soma of the MLI being recorded. Parallel fibre inputs with paired-pulse facilitation were found for all MLIs recorded. Analysis of properties of CF and GrC synapses onto MLI2s and MLI1s in Fig. 2 was restricted to cells with isolated paired-pulse responses for ten trials.
We characterized the contributions of AMPARs and NMDARs to CF–MLI and PF–MLI synaptic currents with pharmacology. We recorded EPSCs from MLIs at +40 mV with a 10-s intertrial interval. Voltage steps were 3.5 s in duration, and single stimuli were delivered at 2.5 s. Voltage steps without stimulation were recorded each trial for subtraction. We then blocked the AMPA component with NBQX (5 µM) and followed this by bath application of the NMDAR antagonist CPP (2 µM). The access resistance and leak current of the MLIs were monitored continuously. Current traces are the average of approximately ten trials. The NMDA/AMPA ratio was calculated as the peak current of the NMDA component divided by the peak current of the AMPA component, with both components measured at +40 mV.
In situ hybridization
Hybridization chain reaction fluorescence in situ hybridization (HCR-FISH) experiments were performed using acute cerebellar slices from P28–P45 mice prepared as described, and fixed for 2 h in 4% paraformaldehyde in PBS (Biotium) at 4 °C. Slices were stored overnight in 70% ethanol in RNase-free water at 4 °C. A floating slice HCR-FISH protocol17 was performed with the following probes and matching hairpins (Molecular Instruments): sortilin-related VPS10 domain containing receptor 3 (Sorcs3), neurexophilin 1 (Nxph1) and glutamate receptor, ionotropic, NMDA2B (epsilon 2) (Grin2b). MLI1s express Sorcs3, and MLI2s express Nxph1. Amplification hairpins were B1–750, B2–488 and B3–647. Slices were mounted on slides (Superfrost Plus, VWR) with mounting medium (Fluoromount, ThermoFisher) and no. 1 coverslips. Images were acquired with a Leica Stellaris X5 confocal microscope using a ×63 oil immersion objective (1.4 numerical aperture, Olympus). The hybridization chain reaction probe/hairpin channels were imaged with 180-nm resolution in a 20-µm-thick, 1-µm-interval tiled z series in lobule IV/V. Sorcs3+ and Nxph1+ cell locations in the molecular layer were manually labelled using the multi-point tool in Fiji. Grin2b fluorescence in each cell was averaged within a 7-µm-diameter circular mask in a single z-plane through the centre of each cell in MATLAB (MathWorks).
In vivo electrophysiology and calcium imaging
Mice (P45–P90) for in vivo electrophysiology recordings (wild type C57BL/6J) and calcium imaging (Tg(PCP2-Cre)3555Jdhu) were used in accordance with approval from the Duke University Animal Care and Use Committee. Animals were housed on a normal light–dark cycle, and animals of either sex were randomly selected for experiments. No power analysis or other statistical methods were used to pre-determine sample sizes. Sample sizes were similar to previous publications.
Mice were anesthetized and placed on a stereotaxic apparatus. Isoflurane was delivered at 5% for induction and at 1.5% for maintenance throughout surgery. A titanium headpost (HE Palmer) was secured to the skull with Metabond (Parkell). Body temperature was maintained with a heating pad (TC-111 CWE). Bupranex (0.05 mg kg−1, subcutaneously) was administered for 24 h after surgery and animals were monitored for 4 d.
For in vivo electrophysiology experiments, after 2+ weeks of recovery, mice were anesthetized with 2% isoflurane and a small craniotomy (1 mm) was drilled over the primary fissure of lobule simplex. The craniotomy was covered with Qwik-Cast (WPI) and a thin layer of Metabond between recording days. At 1 week post-surgery, mice were head-fixed and placed on a motorized circular treadmill for 45 min over 4 d. The rotation speed was initially set at 0.06 m s−1 and was increased in steps of 0.02 m s−1 per day until reaching a final speed of 0.12 m s−1. After this initial phase, 2 more days were added in which the motor was turned off and on in alternating epochs of approximately 10 s while mice were on the treadmill. Following habituation, the craniotomy was opened and a Neuropixels 1.0 probe coated with dye (tetramethylindocarbocyanine perchlorate (DiI) or dioctadecyl-tetramethylindodicarbocyanine (DiD)) was lowered at a rate of 3 μm s−1 into the right primary fissure of the cerebellum (antero-posterior: −5.82 mm, medio-lateral: 2.0 mm from bregma), reaching a final placement of around 2,000 μm below the cortical surface. The probe was allowed to settle for 45 min before data collection. A blunted 23-gauge needle (PrecisionGlide) was placed 3 mm from the right eye of the mouse to deliver corneal airpuffs (25 psi). Air flow was controlled with a solenoid (The Lee Company) and an air pressure regulator (SNS). A total of 100 airpuffs were delivered each session. Mice were recorded for 2 or 3 d (1 session each day, n = 3 mice, 5 recording sessions). All Neuropixels recordings were performed with the motorized treadmill set to 0.12 m s−1. After the last day of recording, mice were euthanized and perfused for histology. Then, 50-μm brain sections were collected to visualize and confirm the probe location. Electrophysiology data were collected with SpikeGLX.
For imaging experiments, a craniotomy (3 mm) was drilled over lobule simplex of the right cerebellar hemisphere. For GCaMP expression targeted at PC dendrites, a glass micropipette was filled with virus AAV1.CAG.Flex.GCaMP6f.WPRE.SV40 (UPenn vector core, titre = 1.2 × 1013) or AAV1.CAG.Flex.GCaMP7f.WPRE.SV40 (Addgene 104496, titre = 1.5 × 1013) diluted 1:4 to 1:20 in ACSF (titre for injection: 7.5 × 1011). Then, 150–200 nl of virus was injected at 2–4 sites in the craniotomy. The injection rate was 30–50 nl min−1.
2P imaging was performed using a resonant scanning microscope (Neurolabware) with a ×16 water immersion objective (Nikon CFI75 LWD 16x W, 0.80 numerical aperture) (n = 4 mice, 4 imaging sessions). To stabilize the immersion solution for imaging, a polymer (MakingCosmetics, 0.4% Carbomer 940) was used. For GCaMP imaging, a 920-nm-wavelength Ti:Sapphire laser was raster scanned with a resonant galvanometer (8 kHz; Cambridge Technology) onto the cerebellum at a frame rate of 30 Hz with a laser power of less than 40 mW and a field of view of 1,030 × 581 μm2 (796 × 264 pixels). Emitted photons were collected through a green filter (510 ± 42-nm band filter (Semrock)) onto a GaAsP photomultiplier (H10770B-40, Hamamatsu) and the microscope was controlled using Scanbox software (Neurolabware). Calcium imaging experiments were conducted on a setup with a freely moving wheel. Mice were engaged in locomotion, which was in the forward direction because the position of the animal on the wheel strongly favoured forward running (they are placed behind the centre point of the wheel so that gravity promotes forward locomotion). Speed was not monitored in these experiments.
Analysis and statistics
The numbers of replicates and statistical tests for all experiments are summarized in Extended Data Table 2.
For in vivo Neuropixels recordings, units were sorted using Kilosort 2.0 and manually curated with Phy as described previously8. After unit curation, layer and cell type were identified using a deep-learning classifier48. Units were separated into PCSS (mouse 1 session 1 (m1s1), n = 19; m1s2, n = 26; m2s1, n = 58; m2s2, n = 8; m3s1, n = 15), PCCS (m1s1, n = 13; m1s2, n = 37; m2s1, n = 48; m2s2, n = 76; m3s1, n = 70), MFs (m1s1, n = 80; m1s2, n = 110; m2s1, n = 48; m2s2, n = 13; m3s1, n = 9), MLIs and unclassified units. To obtain the initial population of putative MLIs, we pooled together units in the molecular layer that the classifier labels as MLIs or unclassified units. After this initial step, manual MLI1 (m1s1, n = 4; m1s2, n = 26; m2s1, n = 32; m2s2, n = 15; m3s1, n = 35) and MLI2 (m1s1, n = 0; m1s2, n = 3; m2s1, n = 1; m2s2, n = 4; m3s1, n = 10) classification was performed as previously described in detail8.
For spontaneous CS CCGs, the peak changes in spike rates following the complex spike were measured as the averages of t1 = 1–2 ms and t2 = 3–6 ms for MLIs, and t1 = 0–1 ms and t2 = 4–7 ms for PCs. The cumulative changes in spike rates were measured as the average of 20–30 ms following the complex spike of the integrals of the CCGs. All responses were measured relative to baseline averaged 30 ms before the complex spike. Pairs were determined to be connected if the response had a Z score > 4 or <−4. Latencies were measured as the half-max times for t1 and t2 for connected pairs with response Z scores > 10 or <−10.
PSTHs were constructed by convolving the trial-averaged activity of each unit with a 6-ms Gaussian kernel and then averaging across units. Traces then were aligned to airpuff onset. MF and PCSS heatmaps were cross-validated using half of the trials to calculate the max airpuff response for each unit and then used to sort the averaged Z-scored responses of the remaining trials. MLI1 and MLI2 response heatmaps were constructed by averaging the Z-scored activity of all MLIs connected to the units in the PCCS heatmap. MLIs without a connected PCCS are blanked (white lines). Significantly airpuff-driven cells were obtained by comparing the activity 100 ms before and after the airpuff using the Wilcoxon signed rank test. Data are reported as mean ± s.e.m., and statistical significance was defined as P < 0.05. *P < 0.05; **P < 0.01; ***P < 0.001.
For in vivo calcium imaging analysis, raw data were motion-corrected on the x and y planes using sub-pixel image registration. PC dendrite isolation was performed with a post hoc processing pipeline consisting of principal component analysis followed by individual component analysis49. Final dendrite segmentation was achieved by applying a Gaussian filter and thresholding pixels of individual components. A binary mask was created by combining highly correlated pixels (correlation coefficient > 0.80) and removing overlapping regions between segmented dendrites. Any PC somas present in the field of view were avoided. Raw calcium traces were filtered (high-pass Butterworth filter, 0.035 Hz, order 3) and the MATLAB OASIS package50 was used to identify events corresponding to complex spikes. The ‘FOOPSI’ method was used to account for the shape of the calcium transients using GCaMP as a calcium indicator. In addition, a threshold of 3 s.d. above baseline was used to identify transients.
ΔF/F signals were measured on a trial-by-trial basis by using the mean of a 500-ms window before each airpuff as F0. Spikes that occurred in a 500-ms window 2 s before the airpuff were classified as spontaneous spikes. Spikes that occurred between 0 and 500 ms after airpuff onset were classified as airpuff-evoked spikes. In both conditions, when there was an extra spike within 500 ms of the first spike, the instance was classified as two (or more) spikes. Active dendrites were defined as dendrites that fired at least one spike in the ‘spontaneous’ window or the airpuff window and their distributions were compared using the Mann–Whitney U test. PSTHs were constructed by aligning ΔF/F traces to the first spike after airpuff onset and averaging them across trials. For spontaneous spikes, ΔF/F traces were aligned to the first spike in the equivalent 500-ms window 2 s before airpuff onset. Spiking heatmaps were constructed using cross-validated deconvolved spike PSTHs aligned to airpuff onset. Cross-validation was performed using half of the trials to calculate the latency to the max airpuff response for each dendrite and then using that order to sort the averaged Z-scored responses of the remaining trials. For spike synchrony analysis, synchrony was defined as the co-firing of two or more dendrites within a temporal window of 66 ms (one frame before or after the reference spike). To assess the effect of distance between synchronous dendrites on calcium influx, a sliding window of 50 µm with 10-µm steps was used. Trials were classified as synchronous when two or more dendrites co-fired in the time and distance windows, and they were classified as not synchronous otherwise. Trials in which two or more dendrites closer than the target distance window co-fired were removed from analysis. For assessing statistical significance, ΔF/F traces in the 500-ms window were integrated for each condition and compared using Kruskal–Wallis with Dunn’s test post hoc. Data are reported as mean ± s.e.m. and statistical significance was defined as P < 0.05. *P < 0.05; **P < 0.01; ***P < 0.001.
Adjusting ROIs to avoid fluorescence contamination
To quantify calcium-dependent increases in PC dendrites, it is important to evaluate the spatial extent of fluorescence changes associated with individual PCs (Extended Data Fig. 8). To do so, we constructed a contamination kernel that describes how fluorescence decays outside each dendritic ROI in a way that is independent of dendritic shape or size. This shape-invariant approach was essential because dendrites vary widely in geometry, and contamination primarily occurs at the edges where the imaging point spread function overlaps with adjacent structures. Measuring contamination relative to the ROI perimeter therefore provides a robust, geometry neutral estimate of signal spread.
To implement this approach, we began by evaluating spontaneous PCCS activity using only imaging frames in which a single dendrite fired an isolated spike within a 100-µm spatial and 100-ms temporal window (Extended Data Fig. 8a,b). This ensured that the final contamination profile reflected fluorescence spread directly attributable to the activity of individual ROIs. Next, for each isolated event, we computed the change in raw fluorescence (ΔF) on a pixel-by-pixel basis by subtracting the mean fluorescence of the 15 frames preceding a PCCS. To compare ΔF changes across dendrites, we rotated each ROI to align them in their longest vertical axis (Extended Data Fig. 8c,d). Then, to construct a shape-invariant contamination kernel, we first associated each pixel outside an ROI with both its closest distance to the ROI and its angle from the centre of the ROI. These measurements established a shape-invariant pixel map of the space outside of each ROI that could be combined across dendrites.
To combine these maps into a contamination kernel, we needed to normalize ΔF changes across events of different amplitudes. We therefore established a contamination ratio as the ΔF of each pixel outside the ROI divided by the mean ΔF inside the ROI on the same frame. This ratio expresses contamination as a fraction of the signal magnitude within the active dendrite, allowing comparison across ROIs and events. The final contamination kernel was then constructed by pooling contamination ratios across ROIs and frames into 1-µm radial bins and 20° angular bins. The median ratio within each bin defined a two-dimensional, shape-invariant contamination kernel representing the average fractional fluorescence spread as a function of distance and angle from an ROI (Extended Data Fig. 8e). This approach captures asymmetric contamination patterns that would be averaged out in a one-dimensional radial analysis. Indeed, we found that contamination is not isotropic around ROIs (Gaussian fits), with greater spread along the vertical axis. This finding could be due to dendrite local geometry or motion-related artefacts in our experiments.
By fitting each angular profile with an exponential decay, we defined the zone of contamination as the entire region where the exponential decay remained above 0. This produced a binary contamination mask, meaning that any pixel whose fitted contamination exceeded zero was considered contaminated (Extended Data Fig. 8). This deliberately conservative cutoff ensured that even minimal optical overlap was excluded from subsequent analyses. To correct for cross-contamination among neighbouring dendrites, for each ROI we calculated the distance and angle of every pixel relative to all other ROIs. Pixels falling within the binary contamination mask of any neighbouring ROI were excluded from further analysis (Extended Data Fig. 8f–h). All spike-triggered ΔF/F and event-averaged measurements (Fig. 5d,f–j and Extended Data Fig. 8i) were therefore computed exclusively from pixels classified as uncontaminated, providing a conservative yet reliable estimate of each dendrite’s intrinsic activity.
Simulations
The model of the molecular layer of the cerebellar cortex consisted of 840 MLI1s, 315 MLI2s and 170 PCs with their associated CF inputs (Fig. 4f). The model was constrained by EM reconstructions, slice recordings and in vivo recordings of firing properties. The distributions of MLI1, MLI2 and PC baseline firing rates, capacitances and input resistances were based on slice and in vivo studies8,17. Synaptic connections made by MLI1s and MLI2s were based on slice recordings and EM reconstructions8. MLI1s were electrically coupled to each other with properties based on slice recordings. Model neurons were placed in the volume, and dendritic and axonal arbours assigned as 3D right rectangular prisms centred at the somas, with dimensions based on EM reconstructions. Synaptic and electrical connections were then made between MLIs and PCs with probabilities and strengths based on the volume of arbour overlap (axodendritic for synaptic connections, dendrodendritic for MLI1–MLI1 electrical connections), scaled by measurements of electrical coupling in slice experiments. CF–MLI1 and CF–MLI2 synaptic connections were based on a combination of EM reconstructions and slice experiments. Inputs due to MF activity were modelled as GrC inputs to MLIs and PCs with spike times drawn as Poisson events and their kinetics based on slice recordings.
MLIs were modelled as single-compartment exponential integrate-and-fire neurons with membrane potential (\(V\)) dynamics obeying
$${C}_{{\rm{m}}}\frac{{\rm{d}}V}{{\rm{d}}t}={g}_{{\rm{GABA}}}({V}_{{\rm{I}}}-V)+{(g}_{{\rm{AMPA}}}+{g}_{{\rm{N}}})({V}_{{\rm{E}}}-V)+\frac{{V}_{{\rm{rest}}}-V+{\Delta }_{T}{e}^{\frac{V-{V}_{{\rm{T}}}}{{\Delta }_{T}}}\,}{{R}_{{\rm{m}}}}+\sum _{j}{g}_{{\rm{gap}}}^{j}({V}_{j}-V)$$
(1)
where \({C}_{{\rm{m}}}\) and \({R}_{{\rm{m}}}\) are the membrane capacitance and resistance, \({g}_{\mathrm{GABA}}\) and \({g}_{\mathrm{AMPA}}\) are the inhibitory and excitatory conductances with reversal potentials \({V}_{{\rm{I}}}\) and \({V}_{{\rm{E}}}\), \({g}_{{\rm{N}}}\) is the conductance noise, \({V}_{\mathrm{rest}}\) is the resting potential, \({V}_{{\rm{T}}}\) is the threshold, \({\Delta }_{T}\) is the spike slope factor and \({g}_{\mathrm{gap}}^{j}\) is the gap junction conductance with an electrically coupled cell \(j\) with membrane potential \({V}_{j}\). Parameter values and definitions are given in Extended Data Table 3. Simulations used the Euler methods with 10-μs time steps. Synaptic currents were modelled as a difference of exponentials with rise times, decay times and latencies determined from experimental data. For MLI1s an electrical coupling term mediated by dendrodendritic gap junctions was added that was proportional to the somatic voltage difference between electrically coupled cells. An exponential spike-generating current was added51 to reproduce quantitatively MLI1–MLI1 cross-correlations observed in slice experiments8. This current described the initiation and rising phase of action potentials using two parameters, a soft threshold \({V}_{{\rm{T}}}\) and a spike slope factor \({\Delta }_{T}\). When the membrane potential reached its peak \({V}_{\theta }\), the dynamics in the equation 1 were replaced with a linear downward ramp towards the reset potential lasting 1 ms. Following each action potential, there was a refractory period drawn randomly from a uniform distribution, during which the membrane potential evolved freely under the dynamics in equation 1 with the spike-generating current disabled.
Each PC was modelled using an existing two-compartment model whose single-neuron properties were explored previously52. Its compartments were labelled ‘somatic’ and ‘dendritic’ with their associated membrane potentials \({V}_{{\rm{s}}}\) and \({V}_{{\rm{d}}}\) obeying
$${C}_{{\rm{s}}}\frac{{\rm{d}}{V}_{{\rm{s}}}}{{\rm{d}}t}={g}_{{\rm{s}}}{(V}_{{\rm{rest}},{\rm{s}}}-{V}_{{\rm{s}}})+({g}_{{\rm{s}}}+{g}_{j}){\Delta }_{T}{e}^{\frac{{V}_{{\rm{s}}}-{V}_{{\rm{T}}}}{{\Delta }_{T}}}+{g}_{j}({V}_{{\rm{d}}}-{V}_{{\rm{s}}})+{g}_{{\rm{GABA}},{\rm{s}}}({V}_{{\rm{I}}}-{V}_{{\rm{s}}})$$
(2)
$${C}_{{\rm{d}}}\frac{{\rm{d}}{V}_{{\rm{d}}}}{{\rm{d}}t}={g}_{{\rm{d}}}({V}_{{\rm{rest}},{\rm{d}}}-{V}_{{\rm{d}}})+{g}_{j}({V}_{{\rm{s}}}-{V}_{{\rm{d}}})+{g}_{{\rm{GABA}},{\rm{d}}}({V}_{I}-{V}_{{\rm{d}}})+{g}_{{\rm{AMPA}},{\rm{d}}}({V}_{{\rm{E}}}-{V}_{{\rm{d}}})$$
(3)
where \({C}_{{\rm{s}}},{C}_{{\rm{d}}}\) and \({g}_{{\rm{s}}},{g}_{{\rm{d}}}\) are the membrane capacitances and conductances of the two compartments, \({g}_{j}\) is the junctional conductance between the compartments and the remaining parameters are defined analogously to the MLIs. A linear downward ramp was added as with the MLIs but with duration 100 μs. MLI–PC connections were distributed among the somatic and dendritic compartments with the ratio chosen to approximate the distributions of somatic versus dendritic MLI1–PC synapses seen in EM reconstructions. This ratio depended linearly on the distance of the MLI1 from the PC layer, from 50/50 for MLI1s at the PC layer to 100% onto the dendritic compartment for MLI1s at the top of the molecular layer.
For each of the ten reconstructed CFs (Fig. 1), two sets of aggregate data were compiled from the verified contact locations obtained from the serial EM reconstructions. (1) The numbers of contacts onto each postsynaptic MLI were recorded along with its cell type (MLI1 or MLI2). (2) The spatial bounding boxes of these contacts along the three directions were recorded for each CF (min and max coordinates along the three directions, corrected to have the PCL at y = 0). Using this aggregate data, model CFs were chosen randomly from the ten reconstructed CFs with replacement and placed at the location of each PC soma. For each CF, we made two ranked lists for each postsynaptic cell type (MLI1 and MLI2), one containing number of contacts (from EM data) and the other containing a list of potential postsynaptic partners, in order of axo-dendritic overlap volume. The number of connections that a specific CF made with a specific MLI was assigned by matching the two lists. To convert from contact numbers to synaptic conductances, we assumed that the conductance was proportional to the number of contact sites. We estimate the conductance corresponding to an individual contact by dividing the maximal observed conductance by the maximum number of contacts observed in reconstructions. This yielded a spillover quantal size of 1.2 nS for MLI2s and 0.375 nS for MLI1s.
The network simulator along with its analysis was written in Python using the package ANNarchy53 and run on a 16-core machine. Simulations were prepared and run in batches with the same initial state and different run conditions. For example, during the stimulation of the ten individual CFs, a batch of identical networks was prepared, whereas in each network a different CF was stimulated along with other cases such as the co-activation of MFs. This allowed individual neurons to be compared across different runs in the batch. Simulations consisted of a 60-s baseline, followed by 5,000 300-ms-long airpuff trials, for a total of 1,560-s simulated time.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.