Animal studies
All animal studies were performed according to protocols approved by the University Committee on Animal Resources at the University of Rochester Medical Center. Gclcf/f mice27 were crossed with the MMTV-PYMT (The Jackson Laboratory, 022974)61 and Rosa26creERT2 (The Jackson Laboratory, 008463)62 mouse strains. For the tumour allograft model, Gclcf/fRosa26creERT2 MMTV-PYMT mice and control C57BL/6 mice (The Jackson Laboratory, 000664) were used (female, at least 12 weeks old). For tumour xenograft experiments, athymic nude NU/J mice (The Jackson Laboratory, 002019) were used (female, at least 8 weeks old).
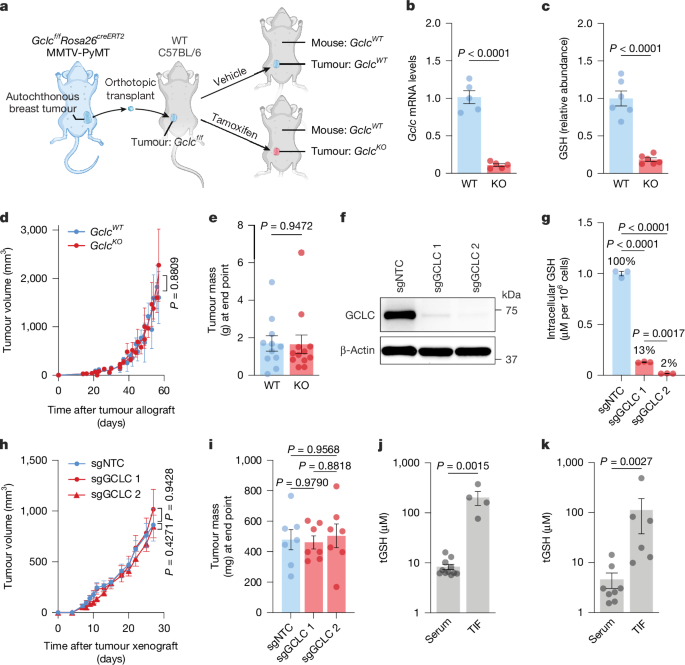
For the tumour allograft model, Gclcf/fRosa26creERT2 MMTV-PYMT mice were euthanized and autochthonous tumours were collected, cut into 8 mm3 fragments and cryopreserved in liquid nitrogen. These fragments were later thawed on ice and implanted into the right fourth mammary fat pad of immunocompetent C57BL/6 mice. Once the grafted fragments had formed fully established tumours, they were collected, cut into 8 mm3 fragments and cryopreserved using the same procedure. Finally, these tumour fragments were thawed and grafted into recipient immunocompetent C57BL/6 mice and used for the designated experimental treatments. To activate Cre recombinase and induce Gclc deletion in tumour pieces from Gclcf/fRosa26creERT2 MMTV-PYMT mice, female mice were intraperitoneally injected with tamoxifen (Sigma-Aldrich, T5648) at 50–160 mg kg–1 once daily for 5 consecutive days. For tumour xenograft models, 1 × 106 HCC-1806 cells or 5 × 106 PC3 cells were injected into the right fourth mammary fat pad or hind flank of athymic nude NU/J mice (The Jackson Laboratory, 002019) in 100 µl sterile PBS or a 1:1 mixture of cell suspension in PBS and Matrigel (Corning, 356237) using a 29-gauge 1 ml insulin syringe. In all cases, animals received only one cell injection at a single location per experiment.
Tumour volume was estimated using the formula for an oblate spheroid \(v=\frac{\pi }{6}\times ({l}^{2})\times w\), where volume (v) is calculated using caliper measurements of the longest (l) and shortest (w) sides of the tumour. For pharmacological interventions, treatment of mice was initiated once the average tumour volume in the groups reached approximately 300 mm3. Mice were treated with pharmacological agents by either intraperitoneal (i.p.) injection or inclusion in the drinking water. For treatment via i.p. injection, GGsTop (Tocris, 4452; WuXi, 926281-37-0; or MCE, HY-108467) was diluted in sterile saline and injected at the indicated concentrations. For treatment via drinking water, GGsTop (WuXi, 926281-37-0) was diluted to 62.5 µg ml–1 (189 µM) and provided to mice ad libitum. Based on an average daily water consumption of 4 ml, the estimated daily dose of GGsTop for a 25 g mouse was 10 mg kg−1 per day (equivalent to 5 mg kg−1 twice a day i.p. injections). NAC (Sigma-Aldrich, A7250) was diluted in drinking water at a concentration of 30 mM, the pH was adjusted to 7.00 and it was provided to mice ad libitum.
Mice of desired strains were age-matched and assigned randomly to their treatment groups. For xenograft studies, animals were allocated into groups ensuring the mean, median and standard error of tumor size was similar across all groups. Investigators were not blinded to group allocation during experiments owing to technical limitations. Statistical methods were not used to chose sample sizes. The number of animals assigned per condition was selected to account for the variability of the examined phenotypes based on pilot experiment and past experience with the animal models (≥6 animals per experimental condition). Mice were euthanized at humane end points (when the tumour diameter exceeded a total of 20 mm in length, when ulceration occurred or when weight loss exceeded 20%). Tumours and tissues were snap-frozen on dry ice and stored at –80 °C or placed in 10% neutral buffered formalin (Fisher Scientific, 22-110-761) for 24 h and then stored in 100% methanol (Fisher Scientific, A412P-4).
Immunohistochemistry
Formalin-fixed, paraffin-embedded tissue sections (5 µm) were used for haematoxylin and eosin staining and immunohistochemical analyses. The tissues were dewaxed and rehydrated through a series of xylene and ethanol changes. For antibody staining, antigen retrieval was performed on the slides by incubating them in a steamer for 40 min in citrate buffer (Vector Labs, H-3300-250). Tissues were permeabilized with PBS supplemented with 0.1% Tween-20 for 10 min, followed by a peroxidase block with 3% H2O2 (Sigma-Aldrich, 216763) for 20 min. The slides were washed in PBS and then blocked using 10% goat serum in PBS for 1 h at room temperature before adding primary antibodies diluted in the blocking buffer. Primary antibody (anti-GCSc 1:100 (SantaCruz, sc390811), anti-CD45 1:100 (Santa Cruz, sc1178), anti-F4/80 1:100 (Cell Signaling Technology, 70076), anti-DNA/RNA Damage 1:200 (Abcam, 62623), anti-NRF2 1:100 (Abcam, 31163) and anti-GGT1 1:200 (donated from M. H. Hannigan)63) incubation was carried out overnight at 4 °C. Biotinylated goat anti-mouse IgG or goat anti-rabbit IgG (Vector Labs, BA-9200 1:200 and BA-1000 1:200, respectively) secondary antibodies were added in blocking buffer and incubated for 1 h at room temperature. Stains were developed using VectaStain Elite ABC-HRP Peroxidase and DAB Substrate kits (Vector Labs, PK-7100 and SK-4100, respectively). Tissues were counterstained with haematoxylin, mounted with Permount mounting medium (Fisher Scientific, SP15500) and coverslipped for imaging. Images were taken using an Olympus VS120 virtual slide microscope and Visiopharm image analysis system.
Immunofluorescence
Formalin-fixed, paraffin-embedded tissue sections (5 µm) were used for immunofluorescence. The tissues were dewaxed and rehydrated through a series of xylene and ethanol changes. For antibody staining, antigen retrieval was performed on the slides by incubating them in a steamer for 40 min in citrate buffer (Vector Labs, H-3300-250). The slides were washed in water and then blocked using 10% goat serum in PBS for 1 h at room temperature before adding primary antibodies diluted in the blocking buffer. Primary antibody (anti-GCSc 1:10 (SantaCruz, sc390811) and anti-CD45 1:200 (Proteintech, 31243-1-AP)) incubation was carried out overnight at 4 °C. Secondary antibodies (AlexaFluor 594 anti-mouse 1:500 (Invitrogen, A11005) and AlexaFluor 488 anti-rabbit 1:500 (Invitrogen, A11034)) were added to blocking buffer and incubated at room temperature for 1 h. For immunofluorescence, the second-to-last wash included 1 µg ml–1 4′,6′-diamidino-2-phenylindole dihydrochloride (DAPI; Sigma-Aldrich D9542) in PBS to stain cell nuclei. Immunofluorescence tissues were mounted with Immuno-Mount (Thermo Fisher, 9990402) and coverslipped for imaging. Immunofluorescence-stained slides were imaged using a Zeiss Axioimager.M2m with an AxioCam MRm camera and the AxioVision program for imaging.
Cell culture
Cell lines were maintained in RPMI 1640 (Thermo Fisher, 11875119) with 5% FBS (Sigma-Aldrich, SH30396.03) and 1% penicillin and streptomycin (Thermo Fisher, 15070063). Cell lines were obtained from the American Type Culture Collection (ATCC), the National Cancer Institute Division of Cancer Treatment and Diagnosis (NCI-DCTD) or donated from J. Brugge, S. Mello, B. Altman and P. Rappold. PC3 wild-type and PC3 GGT1+ cell lines were a gift from M. Hannigan40. The authenticity of the HCC-1806, MDAMB231, MDAMB468, JIMT-1 and A549 cell lines was confirmed by STR profiling. The authenticity of the remaining cell lines was not tested. All cell lines tested negative for mycoplasma contamination using a MycoAlert Mycoplasma Detection kit (Lonza, LT07-418). For experiments involving alterations to the culture medium, cells were seeded in regular RPMI 1640 with 5% dialysed FBS (Gibco, 26400044) and 1% penicillin and streptomycin. RPMI 1640 medium without l-glutamine, l-cysteine, l-cystine and l-methionine was used as the base medium for different medium conditions (MP Biomedicals, 1646454). l-glutamine (Fisher Scientific, 25030081), l-methionine (Sigma-Aldrich, M5308), l-cystine (Sigma-Aldrich, C7602), GSH (Sigma-Aldrich, G4251) or cysteinylglycine (Bachem, 4002969) was added to each condition at the indicated concentrations. GGsTop (Tocris, 4452; WuXi, 926281-37-0; or MCE, HY-108467), acivicin (MCE, HY-W016586), OU749 (Cayman, 13804), auranofin (MCE, HY-B1123), erastin (MCE, HY-15763), vincristine sulfate (MCE, HY-N0488), ferrostatin-1 (Fer-1; MCE, HY-100579), RSL3 (MCE, HY-10018A) and MK-571 (MCE, HY-19989) were diluted in dimethyl sulfoxide (DMSO; Sigma-Aldrich, 472301) at 10–20 mM and added at the indicated concentrations. dl-Buthionine-(S,R)-sulfoximine (BSO; MCE, HY-106376) and trolox (Sigma-Aldrich, 238813) were diluted in water at 100 mM and added at the indicated concentrations. Cells were seeded at 50,000 per well in 6-well plates and at 10,400 per well in 24-well plates. After allowing the cells to attach for 24 h, the medium was aspirated, the wells were washed once with PBS and the indicated medium was added. For experiments in 384-well plates, 500 cells were seeded per well, and after 24 h, the medium was switched using a series of 4 aspiration–wash steps with the desired final medium using Multidrop Combi (Thermo). Co-culture experiments with PC3 WT and GGT1+ cell lines were performed using 6-well (24 mm) PET inserts with 0.4 µm pores (VWR, 76313-902 or Corning, 354570). For these experiments, cells were seeded at the same density (12,992 cells per cm2) in both the bottom well and the insert. For cell number measurements in 6-well plates, the medium was aspirated, the wells were rinsed with PBS and the cells were detached by incubating with 500 µl of 0.25% trypsin (Thermo Scientific, 25200056) for 20 min at 37 °C. Trypsin was neutralized with 500 µl complete medium, the cells were resuspended and 100 µl of the suspension was mixed with 6.9 ml Beckman Coulter Isoton II diluent (Fisher Scientific, NC2470899) for counting with a Beckman Coulter Z1 Coulter Cell Particle counter. For cell number measurements in 24-well plates, unstained plates were scanned in bright field using SparkCyto (Tecan) at 37 °C under 5% CO2. For experiments in 384-well plates, wells were aspirated, rinsed with PBS and incubated with a fixative and staining solution containing 3.7% formaldehyde (Fisher Scientific, F75P-1GAL) and 5 µg ml–1 Hoechst 33342, trihydrochloride, trihydrate (Invitrogen, H1399). After 30 min, wells were aspirated, washed with PBS and plates were sealed with adhesive foil (VWR, 60941-124). Plates were imaged on a CellInsight CX5 HCS platform (Thermo), and counts of nuclei were used as the readout.
Generation of KO and knockdown cell lines
Optimized sgRNA oligonucleotides64 were cloned into lentiCRISPRv2 (ref. 65) (Addgene, 52961) for CRISPR–Cas9 KO or into pLV hU6-sgRNA hUbC-dCas9-KRAB-T2a-Puro66 (Addgene, 71236) for CRISPRi knockdown, as previously described41. HEK293T cells were transfected with 1 µg lentiCRISPRv2 containing the sgRNA insert, 0.67 µg psPAX2 (Addgene, 12260) and 0.33 µg pVSVg (Addgene, 8454) using 6 µg polyethylenimine (Sigma-Aldrich, 919012) in Opti-MEM (Thermo Fisher, 31985062). Supernatants containing lentiviral particles were collected and replaced with fresh medium daily for 3 consecutive days. The pooled lentiviral supernatant was filtered through 0.45 µm PVDF syringe filters (Whatman, 6746-2504), aliquoted and stored at –80 °C. For transduction, 200 µl lentiviral supernatant was thawed on ice and added to HCC-1806 cells together with 8 mg ml–1 Polybrene (hexadimethrine bromide; Sigma-Aldrich, H9268) in 6-well plates. Cells were then allowed to recover for 24 h in fresh medium before antibiotic selection with 2 µg ml–1 puromycin for at least 4 days, after which they were expanded or used for subsequent experiments. To isolate single-cell clones, antibiotic-selected polyclonal GGT1 KO cells were seeded at low density (50 cells per 15 cm dish) and grown for 21–28 days. Then colonies were individually collected by trypsinization using 8 mm cloning cylinders (Pyrex, 09-552-21) and transferred to individual wells for expansion. The following sgRNAs for GGT1 CRISPRi were used: sgGGT1 1 (HL-ID: 3) forward: 5′-CACCGGTGCCTCCCACTGTCCGCCG-3′; sgGGT1 1 (HL-ID: 3) reverse: 5′-CACCGGTCTGGGCTCACCCGACGCC-3′; sgGGT1 2 (HL-ID: 5) forward: 5′-AAACCGGCGGACAGTGGGAGGCACC-3′; and sgGGT1 2 (HL-ID: 5) reverse: 5′-AAACGGCGTCGGGTGAGCCCAGACC-3′. The following sgRNAs for GGT1 CRISPR–Cas9 KO were used: sgGGT1 1 (HL-ID: 6) forward: 5′-CACCGTGCGGGACGGTGGCTCTG-3′; sgGGT1 1 (HL-ID: 6) reverse: 5′-AAACCAGAGCCACCGTCCCGCAC-3′; sgGGT1 2 (HL-ID: 10) forward: 5′-CACCGCCCAGTGCGCCGCTCAG-3′; and sgGGT1 2 (HL-ID: 10) reverse: 5′-AAACCTGAGCGGCGCACTGGGC-3′. The following sgRNAs for GCLC CRISPR–Cas9 KO were used: sgNTC forward: 5′-CACCGGAGGCTAAGCGTCGCAA-3′; sgNTC reverse: 5′-AAACTTGCGACGCTTAGCCTCc-3′; sgGCLC-a forward: 5′-CACCGCATACTCACCTGAAGCGA-3′; sgGCLC-a reverse: 5′-AAACTCGCTTCAGGTGAGTATGC-3′; sgGCLC-b forward: 5′-CACCGAAATATCCGACATAGGAG-3′; and sgGCLC-b reverse: 5′-AAACCTCCTATGTCGGATATTTC-3′.
Flow cytometry
To quantify cell proliferation, 300,000 cells were plated in 100 mm dishes and treated as specified. At the end point, attached cells were pulsed with 10 mM BrdU (Sigma-Aldrich, B9285) for 30 min in their culture medium. Labelled cells were then detached by trypsinization, pelleted, washed once with PBS, resuspended in 300 µl ice-cold PBS and fixed by adding 700 µl ice-cold absolute ethanol while gently vortexing. Fixed cells were stored at –20 °C. For labelling, the fixed cells were thawed and washed for 5 min with 1 ml of 1% BSA (Sigma-Aldrich, A7906) in PBS. The cells were then centrifuged at 10,000g for 2 min and incubated with 1 ml of 2 M HCl and 0.5% Triton X-100 (Sigma-Aldrich, 93443) for 30 min with gentle agitation. Then the cells were centrifuged and washed once with 1 ml of 0.1 M borax (Sigma-Aldrich, 71997), followed by another wash with 1% BSA in PBS. After centrifugation and thorough removal of the supernatant, the pellets were incubated for 1 h in 60 µl staining solution (1% BSA and 0.5% Tween-20 in PBS) containing 3 µl anti-BrdU–FITC monoclonal antibody (BioLegend, 364104, clone 3D4). The samples were subsequently washed with 1 ml of 1% BSA and 0.5% Tween-20 in PBS, centrifuged and the pellets were resuspended in 1 ml of PI solution (20 µg ml–1 propidium iodide (Sigma-Aldrich, P4170), 0.5 mg ml–1 RNase A (Sigma-Aldrich, R5503) and 0.1% Triton X-100 in PBS). The cells were incubated overnight (approximately 16 h), and then 10,000 events were analysed by flow cytometry using a BD Accuri C6 Plus flow cytometer (BD Biosciences) and FCS Express 7 Research (DeNovo Software).
For quantifying cell death, 50,000 HCC-1806 cells were seeded into 6-well plates and treated as indicated. At the end point, cells were collected by trypsinization (the culture medium and PBS washes were collected and re-added to each sample) and washed once with assay buffer (10 mM HEPES, 140 mM NaCl and 3.3 mM CaCl2, pH 7.4). Then the pellets were stained for 20 min in 100 µl assay buffer containing 5 µg ml–1 propidium iodide (Sigma-Aldrich, P4170) and 3 µl Annexin V–FITC (BioLegend, 0640906). Then cells were diluted with 0.5–1 ml assay buffer, and 10,000 events were analysed by flow cytometry using a BD Accuri C6 Plus flow cytometer (BD Biosciences) and FCS Express 7 Research (DeNovo Software).
For quantifying lipid peroxidation, 95,000 cells were seeded into 12-well plates and treated as indicated. At the end point, 2.5 µM Bodipy 581/591 C11 (Cell Signaling, 95978) was added directly to each well and incubated for 30 min. Wells were then washed with PBS and collected by trypsinization. Pellets were resuspended in 300 µl PBS, and 10,000 events were analysed by flow cytometry using a BD Accuri C6 Plus flow cytometer (BD Biosciences) and FCS Express 7 Research (DeNovo Software).
Immunoblot assays
Cell lysates from HCC-1806, PC3 WT and PC3 GGT1+ cells were obtained by placing PBS-washed cell plates on ice and scraping with RIPA buffer (Thermo Scientific, 89900) containing Halt Protease and Phosphatase Inhibitor cocktail (Thermo Scientific, 1861280). Cell lysates were incubated for 30 min on ice and then centrifuged at 12,000g for 15 min at 4 °C. The supernatant was collected and stored at –80 °C. Extracted proteins were quantified using a Pierce BCA Protein Assay kit (Thermo Scientific, 23225). Lysates were combined with 6× Laemmli SDS sample buffer (Boston BioProducts, BP-111R) containing 5% 2-mercaptoethanol (VWR Life Science, M131), then loaded onto 4–20% Criterion TGX pre-cast gels (Bio-Rad, 5671093). Each lane received 40 µg protein for HCC-1806 samples or 75 µg for PC3 samples. Separated proteins were transferred onto Immobilon-P Transfer membranes (MilliporeSigma, IPVH00010), blocked for 1 h using 5% BSA in TBS–0.5% Tween-20 and stained overnight with primary rabbit-anti-GCLC (Sigma-Aldrich, HPA036359) or affinity-purified rabbit-anti-GGT1 (GGT129, as previously described63) 1:1,000 in BSA 2.5% overnight at 4 °C. Stained membranes were washed three times with TBS-T and stained with HRP-linked secondary antibody donkey-anti-rabbit (Amersham, NAV934) 1:5,000 in BSA 2.5% for 1 h. Membranes were washed three more times with TBS-T, and then the antibody-stained protein signal was amplified and visualized using SuperSignal (Thermo Fisher, 34577). Blots were imaged with a ChemiDoc MP Imaging system (Bio-Rad). Membranes were stripped for 20 min at room temperature using a Blot Restore Membrane Rejuvenation kit (Sigma-Aldrich, 2520-M), then washed twice with TBS-T, re-blocked with BSA 5% for 30 min and incubated with mouse-anti-β-actin (Sigma-Aldrich, A1978) 1:3,000 in BSA 2.5% for 4 h at room temperature and then incubated with a sheep-anti-mouse (Amersham, NA931) secondary antibody at 1:5,000 in BSA 2.5% for 2 h.
Drug screening
MAPS was performed as previously described41. In brief, 500 cells were seeded per well in 384-well plates, and after 24 h, the medium was switched using a series of 4 aspiration–wash steps with the desired final medium. Compound libraries were gifts from J. Brugge and the ICCB-L. Next, 100 nl of compounds were transferred using a JANUS MDT Workstation (Revvity) and a 384-well pin tool (V&P Scientific). After 96 h, wells were aspirated, washed with PBS, and a fixative/staining solution containing 3.7% formaldehyde (Fisher Scientific, F75P-1GAL) and 5 µg ml–1 Hoechst 33342, trihydrochloride, trihydrate (Invitrogen, H1399) was added. After 30 min, wells were aspirated, PBS was added and plates were sealed with adhesive foil (VWR, 60941-124). Plates were imaged using a CellInsight CX5 HCS platform (Thermo), and nuclei were identified as a readout of cell counts. Data post-processing was conducted using R scripts.
GGT activity assay
Cell cultures were washed and scraped with PBS. The use of trypsin for cell detachment was avoided to prevent proteolytic degradation of cell-surface GGT. Detached cells were collected and centrifuged at 300g for 5 min at room temperature. Then pellets were resuspended in lysis buffer (Tris-HCl 100 mM and 0.5% Triton X-100, pH 8) containing Halt Protease and Phosphatase Inhibitor cocktail (Thermo Scientific, 1861280), thoroughly vortexed for 1 min and incubated for 15 min on ice. Lysates were cleared by centrifugation at 12,000g for 15 min at 4 °C and the supernatants were stored at –80 °C until analysis. Tissues were lysed by transferring them to pre-filled bead mill tubes (Fisher Scientific, 15-340-154) containing lysis buffer (Tris-HCl 100 mM and 0.5% Triton X-100, pH 8) containing Halt Protease and Phosphatase Inhibitor cocktail and homogenized using a bead mill (VWR) for 10 s. Tissue lysates were then rotated for 15 min at 4 °C and centrifuged at 12,000g for 15 min at 4 °C. Supernatants of cells or tissues were stored at –80 °C until analysis. Extracted proteins were quantified using a Pierce BCA Protein Assay kit. GGT reaction assays were carried out by using the GpNA method67. In brief, cell or tissue protein lysates were assayed in transparent 96-well plates (Greiner, 655101) in the presence of 1 mM of the GGT substrate l-glutamic acid γ-p-nitroanilide (GpNA; Cayman, 36209) and 20 mM of the transpeptidation acceptor glycyl-glycine (Sigma-Aldrich, G1002) in Tris-HCl 100 mM, pH 8, buffer. GGT catalyses the breakdown of the γ-glutamyl bond in the substrate and generates p-nitroanilide (pNA), which is monitored kinetically by an increase in absorbance at 418 nm. P-nitroaniline (Sigma-Aldrich, 185310) was used to generate a standard curve (0–40 nmoles) for the calculation of GGT activity. Absorbance (418 nm) was assessed using a plate reader set at 37 °C with a total reading time of 45–60 min and 1–5 min intervals between reads. GGT activity was calculated by obtaining the slope of the linear range of each sample (Abs per min) and dividing by the slope of a 4-nitroaniline standard curve (Abs per nmoles pNA) and by the protein mass (mg) to determine the reaction rate of the sample (nmoles pNA min−1 mg−1). The optimal protein mass for the GGT assay was experimentally determined for each cell line or mouse tissue (HCC-1806 lysates or tumour xenografts: 200 µg; PC3 cell line: 100 µg; PC3 GGT1+: 1 µg; kidney: 0.5 µg; spleen: 150 µg; liver: 250 µg; seminal vesicle: 10 µg; pancreas: 2.5 µg; lungs: 2.5 µg; muscle: 200 µg; testis: 50 µg; epididymis: 10 µg; PyMT breast tumour: 50 µg; heart: 150 µg; mammary fat pad: 150 µg; brain: 80 µg; large intestine: 80 µg; brown adipose tissue: 300 µg).
GGT histochemical assay
Histochemical staining of GGT was performed as previously described68, with modifications. Cells were seeded into 6-well plates and cultured in complete medium. At the end point, cells were washed 3 times with saline and stained for 20 min at room temperature using a saline solution containing 0.2 mM l-glutamic acid γ-(4-methoxy-β-naphthylamide) (GMNA; Sigma-Aldrich, G0141), 20 mM glycyl-glycine (Sigma-Aldrich, G1002), 25 mM Tris (Sigma-Aldrich, 93362) and 1.2 mM FastBlue (Sigma-Aldrich, F3378). For negative controls, the staining solution included 5 mM serine (Sigma-Aldrich, S4311) and 10 mM sodium borate (Sigma-Aldrich, 71997). After incubation, cells were rinsed with saline and incubated with 100 mM CuSO4 (Sigma-Aldrich, C1297) for 2 min. Following a final saline wash, 50% glycerol was added to the wells, and bright-field images were taken using an inverted microscope.
GSH quantification
For some experiments, the GSH concentration was determined using a GSH-Glo Glutathione Assay (Promega, V6912). In brief, 6,250 cells were seeded into 96-well white cell culture-treated plates (Falcon, 353296), treated for 24 h as indicated and labelled following the manufacturer’s instructions. The luminescence of samples and of the GSH standard curve was detected using a Spark Multimode microplate reader (Tecan).
RNA analysis
mRNA was isolated from cells or tissues using an E.Z.N.A. total RNA kit I (Omega Bio-Tek, R6834-02). For gene expression analysis, 1 µg RNA was used for cDNA synthesis using a qScript cDNA Synthesis kit (Quanta Bio, 66196756). The expression of target genes was analysed via quantitative real-time PCR with a QuantaStudio 5 qPCR machine (Applied Biosystems, Thermo Fisher Scientific).
Mouse serum analysis
Mice were anaesthetized with isoflurane, after which blood was collected via the retro-orbital venous sinus into BD microtainer tubes (BD, 365967). Serum was isolated from the blood by centrifuging blood samples at 10,000g for 5 min and stored at –80 °C. VRL Animal Health Diagnostics carried out the analysis of biomarkers of liver damage.
Metabolite analysis
For metabolite analysis of cultured cancer cells, cells were seeded in 6-well plates and treated as specified. At the end point, cells were rapidly washed with ice-cold PBS and incubated at –80 °C for 30 min with extraction solvent at a ratio of 1 ml per 1 × 106 cells. The cells were then scraped on ice, transferred to pre-chilled microcentrifuge tubes, centrifuged at 15,000g for 20 min and the supernatants were collected. Conditioned medium was extracted by taking 10 µl of the medium supernatant and mixing it with 390 µl of the extraction solvent. Control medium samples were generated by incubating the treatment medium in cell-free 6-well plates under the same conditions and duration as the cell culture samples. Extracts of the medium were incubated at –80 °C for at least 15 min before being centrifuged at 15,000g for 20 min. All extracts were stored at –80 °C. Metabolite profiling of reduced thiol-containing metabolites was carried out in the presence of NEM. The data shown reflect the detection of their corresponding NEM adducts (for example, NEM-cystine, NEM-cysteinylglycine and NEM-GSH). tGSH was calculated as the sum of reduced GSH and two times the amount of oxidized glutathione (tGSH = GSH + 2 × GSSG). The extraction solvent consisted of 80% methanol and 20% aqueous solution (pH 7.00) containing 10 mM ammonium formate (Sigma-Aldrich, 70221-25G-F) and 25 mM NEM (Thermo Scientific, 040526.06). The final concentrations of ammonium formate and NEM in the extraction solvent were 2 mM and 5 mM, respectively. For inverse tracing experiments, HCC-1806 cells were seeded in T75 flasks and cultured for >7 days in cystine-free medium supplemented with heavy isotope-labelled 13C2-cystine (Cambridge Isotope Laboratories, CLM-520-0). At the end of this >7-day labelling period (t = 0 h), samples were collected to verify complete incorporation of the heavy label into cellular metabolites. The remaining cells were collected by trypsinization, washed three times with PBS and seeded into 6-well plates using cystine-free medium containing unlabelled GSH. After 3 days in the presence of unlabelled GSH (t = 72 h), cells were extracted for metabolomic analysis as described above.
For quantification of sulfur-containing metabolites in cultured cancer cells and mouse serum, 10 µl samples were extracted with 90 µl of ice-cold extraction solvent (80% methanol and 20% H2O) containing isotope-labelled internal standards (36 µM 13C215N-GSH, 0.92 µM 13C415N2-GSSG and 10 µM D4-cystine). Following 30 min of incubation at 4 °C, extracts were cleared by centrifugation (17,000g, 20 min, 4 °C) and then supernatants were analysed by liquid chromatography and mass spectrometry (LC–MS). For the quantification of these metabolites in TIF samples, 5 µl samples were extracted in 45 µl ice-cold extraction solvent (75% acetonitrile, 25% methanol and 0.1% formic acid) containing isotope-labelled internal standards (36 µM 13C2,15N-GSH, 0.92 µM 13C4,15N2-GSSG and 10 µM D4-cystine). Extracted samples were vortexed at 4 °C for 10 min, cleared by centrifugation (17,000g, 10 min, 4 °C) and then supernatants were analysed by LC–MS. For chromatographic metabolite separation, a Vanquish ultra-performance liquid chromatography (UPLC) system was coupled to a Q Exactive HF (QE-HF) mass spectrometer equipped with heated electrospray ionization (HESI; Thermo Fisher Scientific). Samples were run on an Atlantis Premier BEH Z-HILIC VanGuard FIT column (2.5 µm, 2.1 mm × 150 mm, Waters). Mobile phase A was 10 mM (NH4)2CO3 and 0.05% NH4OH in H2O, whereas mobile phase B was 100% acetonitrile. The column chamber temperature was maintained at 30 °C. The mobile phase parameters were set according to the following gradient: 0–13 min of 80% to 20% of mobile phase B; 13–15 min of 20% of mobile phase B. The ESI ionization mode was negative, and the MS scan range (m/z) was set to 65–975. The mass resolution was 120,000 and the automatic gain control (AGC) target was 3 × 106. The capillary temperature and capillary voltage were maintained at 320 °C and 3.5 kV, respectively. A volume of 5 μl of each sample was loaded for metabolite detection. The LC–MS metabolite peaks were manually integrated and identified using El-Maven (v.0.12.0) by matching with a previously established in-house library33.
For metabolic analysis of cultured cancer cells and tumours, frozen tumour tissue was homogenized in 80% methanol containing 20 mM NEM using a Precellys cold tissue homogenizer (Bertin) at a ratio of 20 mg tissue to 1 ml solvent. Following homogenization, samples were transferred to –80 °C for 30 min and then placed on regular ice for 30 min with vortexing every 10 min. Next, samples were centrifuged at 17,000g for 10 min and 800 µl supernatant was dried in a vacuum evaporator (Thermo). Samples were reconstituted in 90 µl of 50% acetonitrile (Fisher Scientific, A955) and transferred to glass vials for LC–MS analysis. For mouse serum collection, the animals were anaesthetized with isoflurane, and blood was collected via the retro-orbital venous sinus into BD microtainer tubes (BD, 365967). Serum was obtained by centrifuging blood samples at 10,000g for 5 min and stored at –80 °C. Then 20 µl serum was mixed with 2 µl 200 mM NEM (Thermo Scientific, 040526.06) and extracted with 80% methanol. Next, 900 µl serum extract was dried in a vacuum evaporator (Thermo), reconstituted in 90 µl of 50% acetonitrile (A955, Fisher Scientific) and transferred to glass vials for LC–MS analysis. LC–MS analysis was carried out by the URMC Metabolomics Resource. For LC–MS analysis, metabolite extracts were analysed by high-resolution mass spectrometry with an Orbitrap Exploris 240 (Thermo) coupled to a Vanquish Flex liquid chromatography system (Thermo). A volume of 2 µl samples was injected into a Waters XBridge XP BEH Amide column (150 mm length × 2.1 mm i.d., 2.5 µm particle size) maintained at 25 °C, with a Waters XBridge XP VanGuard BEH Amide (5 mm × 2.1 mm i.d., 2.5 µm particle size) guard column. For positive-mode acquisition, mobile phase A was 100% LC–MS-grade H2O with 10 mM ammonium formate and 0.125% formic acid. Mobile phase B was 90% acetonitrile with 10 mM ammonium formate and 0.125% formic acid. For negative-mode acquisition, mobile phase A was 100% LC–MS-grade H2O with 10 mM ammonium acetate, 0.1% ammonium hydroxide and 0.1% medronic acid (Agilent). Mobile phase B was 90% acetonitrile with 10 mM ammonium acetate, 0.1% ammonium hydroxide and 0.1% medronic acid. The gradient was 0 min, 100% B; 2 min, 100% B; 3 min, 90% B; 5 min, 90% B; 6 min, 85% B; 7 min, 85% B; 8 min, 75% B; 9 min, 75% B; 10 min, 55% B; 12 min, 55% B; 13 min, 35%, 20 min, 35% B; 20.1 min, 35% B; 20.6 min, 100% B; 22.2 min, 100% B, all at a flow rate of 150 μl min−1, followed by 22.7 min, 100% B; 27.9 min, 100% B at a flow rate of 300 μl min−1, and finally 28 min, 100% B at flow rate of 150 μl min−1, for a total length of 28 min. The H-ESI source was operated in positive mode at spray voltage 3,500 or negative mode at spray voltage 2,500 with the following parameters: sheath gas 35 a.u., aux gas 7 a.u., sweep gas 0 a.u., ion transfer tube temperature 320 °C, vaporizer temperature 275 °C, mass range 70 to 1,000 m/z, full scan MS1 mass resolution of 120,000 FWHM, RF lens at 70% and standard AGC. Data-dependent MS2 fragmentation for compound identification and annotation was performed via the AquireX workflow (Thermo Scientific), which comprised three deep scans with MS1 resolution at 60,000 and MS2 resolution at 15,000. LC–MS data were analysed using Compound Discover (v.3.3, Thermo Scientific) and El-Maven software69 for peak area determination and compound annotation. Compounds were annotated by matching to LC–MS method-specific retention time values of external standards and MS2 spectral matching to external standards and the mzCloud database (Thermo Scientific).
Human biospecimens, clinical data and metabolite analysis of human samples
Human biospecimens and deidentified clinical data were provided by the Wilmot Cancer Institute Biobank Shared Resource (BSR) at the University of Rochester. All samples were collected under Institutional Review Board-approved protocols (STUDY61977 and STUDY7108), and all participants provided written informed consent. A total of 16 participants were enrolled, all of whom were women with a median age of 64 years (range of 25–88 years). Neoadjuvant treatment was administered in 4 participants (25%). The cohort included 81% white, 6% Black and 13% with race not reported. Most tumours were invasive ductal carcinoma (94%) with 6% classified as other histology. Oestrogen receptor positivity was observed in 81% of cases and progesterone receptor positivity in 62%. HER2 status was distributed as 0 in 56%, 1+ in 32%, 2+ in 6% and 3+ in 6%. Tumour staging showed 44% T1 and 56% T2 with no T3 or T4 disease. Nodal staging included 38% N0, 38% N1, 6% N2 and 18% NX. All participants were M0 at diagnosis with no metastatic disease. Tumour grade was 1 in 12%, 2 in 50% and 3 in 38%.
Peripheral whole blood was collected in the perioperative setting via a venous catheter immediately before surgery using K2-EDTA-coated tubes (BD Biosciences, 367899). Samples were centrifuged at 2,500g for 10 min at 4 °C to separate plasma. The plasma supernatant was aliquoted, snap-frozen in liquid nitrogen and stored at –80 °C before analysis.
Immediately following excision, tumour specimens were delivered to Surgical Pathology and reviewed by certified pathologists at the University of Rochester Medical Center, who allocated remnant malignant tissue for research. Research biospecimens were promptly transferred to the BSR laboratory, rinsed with ice-cold PBS and gently blotted dry. Tumour tissues (100–500 mg) were dissected into smaller fragments to cover the filter surface of 0.22 µm nylon-filtered microcentrifuge tubes (Corning, 8160), taking care to minimize mechanical cell lysis. Tubes were centrifuged at 300g for 5 min at 4 °C to collect TIF. The isolated TIF was snap-frozen in liquid nitrogen and stored at –80 °C before analysis.
Quantitative metabolite profiling of the TIF and serum human patient samples was performed based on a protocol adapted from a previous study28. In brief, chemical standard libraries of 149 metabolites in seven pooled libraries were prepared and serially diluted in high-performance liquid chromatography (HPLC)-grade water from in a dilution series from 5 mM to 1 µM to generate external standard pools, which are used for calibration of isotopically labelled internal standards and to quantify concentrations of metabolites for which internal standards were not available. We then measured 0.3 µl of each sample using a 1 µl Hamilton syringe (Hamilton, 80135) and extracted these volumes in 20 µl of a 75:25:0.1 HPLC grade acetonitrile–methanol–formic acid extraction mix with the following labelled stable isotope internal standards: 2H9-choline (DLM-549), 13C4-3-hydroxybutyrate (CLM-3853), 13C6,15N2-cystine (CNLM-4244), 13C3-lactate (485926, Sigma-Aldrich), 13C6-glucose (CLM-1396), 13C3-serine (CLM-1574), 13C2-glycine (CLM-1017), 13C5-hypoxanthine (CLM-8042), 13C2,15N-taurine (CNLM-10253), 13C3-glycerol (CLM-1510), 2H3-creatinine (DLM-3653), 13C10-kynurenic acid (80445, Sigma-Aldrich), 13C10,15N5-adenosine (CNLM-3806), 13C8-indole-3-carboxaldehyde (CLM-10745), 15N4-inosine (NLM-4264), 13C10-kynurenine (CLM-9884) and 13C4-methylmalonic acid (CLM-9426). All standards were obtained from Cambridge Isotope Laboratory unless stated otherwise. Samples in extraction mix were vortexed for 10 min at 4 °C and centrifuged at 15,000 rpm for 10 min at 4 °C to pellet insoluble material. Next, 18 µl of the soluble polar metabolite supernatant was transferred to sample vials for analysis by LC–MS as previously described4,28. Once LC–MS analysis was performed, Skyline software was used for metabolite identification. External standard libraries were used to confirm the m/z and retention time for each metabolite. For quantitative analysis, when internal standards were available, external standard libraries were used to quantify concentrations of isotopically labelled internal standards in the extraction mix. Once internal standard concentrations were obtained, the peak areas of the unlabelled metabolites in the samples of culture medium were compared with the peak area of the quantified internal standard to determine the metabolite concentration in the sample.
For metabolites for which an internal standard was not present in the extraction mix, external standard libraries were used to perform analysis of relevant metabolite concentrations. In brief, the peak area of the metabolite was normalized to the peak area of an isotopically labelled internal standard with similar elution time, both in samples from culture medium and external standard library dilutions. Using the external standard library dilutions, we created a standard curve based on the linear relationship of the normalized peak area and the concentration of the metabolite, excluding those metabolites with an r2 < 0.95. This standard curve was then used to interpolate the concentration of the metabolite in the samples.
Proteomic analysis
Cells were seeded in 10-cm dishes and cultured for 7 days in cystine-free medium supplemented with heavy isotope-labelled 13C6,15N2-cystine (Cambridge Isotope Laboratories, CNLM-4244-H). At the end of this 7-day labelling period (t = 0 h), samples were collected to verify complete incorporation of the heavy label into cellular proteins. The remaining cells were washed three times with PBS and transferred to 6-well plates containing cystine-free medium supplemented with unlabelled GSH. After an additional 4 days (t = 96 h), cells were collected by trypsinization, pelleted, snap-frozen and stored at –80 °C. Cell pellets were then lysed in 50 µl of 5% SDS, 100 mM TEAB and sonicated (QSonica) for 5 cycles, with a 1 min resting period on ice after each cycle. Samples were then centrifuged at 15,000g for 5 min to pellet cellular debris, and the supernatant was collected. Protein concentration was determined using a BCA kit (Thermo Scientific), after which samples were diluted to 1 mg ml−1 in 5% SDS and 50 mM TEAB. Next, 25 µg protein from each sample was reduced with 2 mM dithiothreitol, followed by incubation at 55 °C for 60 min. Iodoacetamide was added to 10 mM and incubated in the dark at room temperature for 30 min to alkylate the proteins. Phosphoric acid was added to 1.2%, followed by six volumes of 90% methanol, 100 mM TEAB. The resulting solution was added to S-Trap micros (Protifi) and centrifuged at 4,000g for 1 min. The S-Traps containing trapped protein were washed twice by centrifuging through 90% methanol and 100 mM TEAB. Then 1 µg of trypsin was brought up in 20 µl of 100 mM TEAB and added to the S-Trap, followed by an additional 20 µl of TEAB to ensure the sample did not dry out. Samples were incubated at 37 °C overnight. The next morning, the S-Trap was centrifuged at 4,000g for 1 min to collect the digested peptides. Sequential additions of 0.1% trifluoroacetic acid (TFA) in acetonitrile and 0.1% TFA in 50% acetonitrile were added to the S-trap, centrifuged and pooled. Samples were frozen and dried in a Speed Vac (Labconco), then re-suspended in 0.1% TFA before MS analysis.
Peptides were injected onto a 75 µm × 2 cm trap column (Thermo Fisher) before re-focusing on an Aurora Elite 75 µm × 15 cm C18 column (IonOpticks) using a Vanquish Neo UHPLC (Thermo Fisher) connected to an Orbitrap Astral mass spectrometer (Thermo Fisher). Solvent A was 0.1% formic acid in water, whereas solvent B was 0.1% formic acid in 80% acetonitrile. Ions were introduced to the mass spectrometer using an Easy-Spray source operating at 2 kV. The gradient began at 1% B and ramped to 5% B in 0.1 min, increased to 30% B in 12.1 min, increased to 40% in 0.7 min and finally increased to 99% B in 0.1 min and was held for 2 min to wash the column for a total run time of 15 min. After each run was completed, the column was re-equilibrated with 1% B before the next injection. The Orbitrap Astral was operated in data-independent acquisition (DIA) mode, with MS1 scans acquired in the Orbitrap at a resolution of 240,000, with a maximum injection time of 5 ms over a range of 380–980 m/z. DIA MS2 scans were acquired in the Astral mass analyser with a 3 ms maximum injection time using a variable windowing scheme, using 2 Da windows from 380 to 680 m/z, 4 Da windows from 680 to 800 m/z and 8 Da windows from 800 to 980 m/z. The HCD collision energy was set to 25%, and the normalized AGC was set to 500%. Fragment ions were collected over a scan range of 150–2,000 m/z. The cycle time was 0.6 s.
For data analysis, the raw data were processed with DIA-NN (v.1.9.2; https://github.com/vdemichev/DIANN)70. For all experiments, data analysis was carried out using library-free analysis mode in DIA-NN. To annotate the library, the Homo sapiens UniProt ‘one protein sequence per gene’ database (UP0000005640_9606, downloaded 7 April 2021) was used, with ‘deep learning-based spectra and RT prediction’ enabled. For precursor ion generation, the maximum number of missed cleavages was set to 1, the maximum number of variable modifications to 1 for Ox(M), the peptide length range to 7–30, the precursor charge range to 2–4, the precursor m/z range to 380–980 and the fragment m/z range to 150–2,000. The quantification was set to ‘Robust LC (high precision)’ mode with normalization disabled, MBR enabled, protein inferences set to ‘Genes’ and ‘Heuristic protein inference’ turned off. MS1 and MS2 mass tolerances, along with the scan window size, were automatically set by the software. To quantify light and heavy cysteine-containing peptides, the following parameters were included in the ‘Additional Options’ pane in the DIA-NN software: fixed-mod Carb, 57.021464, C, label; lib-fixed-mod Carb; channels Carb,L,C,0; Carb,H,C,4.007099; original-mods. Precursors were subsequently filtered at library precursor q value (1%), library protein group q value (1%) and posterior error probability (50%). Protein quantification was carried out using the MaxLFQ algorithm as implemented in the DIA-NN R package (https://github.com/vdemichev/diann-rpackage), and the number of peptides quantified in each protein group was counted as implemented in the DiannReportGenerator package (https://github.com/URMC-MSRL/DiannReportGenerator)71.
Statistical analysis
All statistical analysis was completed using either R or GraphPad Prism (v.10.6.1).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.