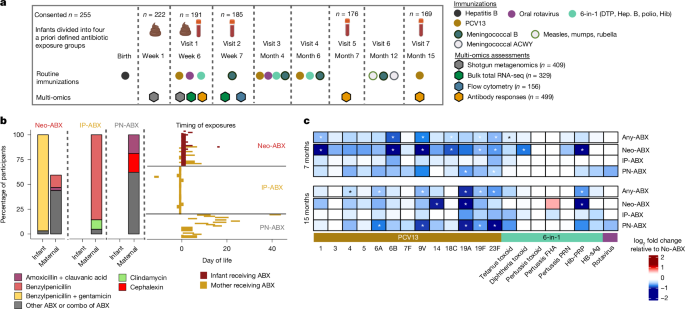
Study design
Infants were enrolled in the study at the Women’s and Children’s Hospital Adelaide, Australia, between April 2017 and March 2021 as part of the Antibiotics and Immune Responses (AIR) study, a prospective, observational clinical study (ACTRN12617000856314). Potential participants were approached by study research nurses in postnatal wards and invited to participate if the mother and their infant met the study eligibility criteria. Study information had also been previously provided to women delivering at the Women’s and Children’s Hospital through pamphlets in antenatal care packs. Following written informed consent, stool collection packs were provided to families, and medical case notes were reviewed to confirm eligibility criteria and antibiotic exposure status. Ethics approval for the study was obtained from the Human Research Ethics Committee of the Women’s and Children’s Health Network (approval number HREC/17/WCHN/19), and the authors affirm that they have complied with all relevant ethical regulations.
Exclusion criteria were gestation of less than 37 weeks; maternal body mass index greater than 30 (at first-trimester antenatal visit); maternal sepsis (defined by laboratory-confirmed bacterial infection in blood cultures or cerebrospinal fluid) during pregnancy; infant delivered by caesarean section; confirmed sepsis in infant (defined by laboratory-confirmed bacterial infection in blood cultures or cerebrospinal fluid); known or suspected disorder of the immune system that would prevent an immune response to the vaccines, such as congenital or acquired immunodeficiency, or receiving systemic immunosuppressive therapy; infant with suspected or confirmed HIV, major congenital abnormality or serious illness; maternal or infant participation in a clinical study that could interfere with participation in this study; and anything that would place the individual at increased risk or preclude their full compliance with or completion of the study. For immunogenicity analyses, only participants who received at least one vaccine at study visit 1 were included.
Participants were allocated to an antibiotic exposure group for immunogenicity analyses on the basis of medical record review and discussion with the mother to ascertain any antibiotic exposure before vaccination at study visit 1 (approximately 6 weeks of age). Antibiotic exposure groups were defined a priori as follows: infants with no direct or maternal antibiotic exposure (n = 80, No-ABX); (1) infants who received at least 48 h of antibiotic treatment in the neonatal period (the first 28 days after birth), with or without maternal antibiotic exposure (n = 32, Neo-ABX); (2) infants whose mothers received intrapartum antibiotics (within 28 days before or during delivery) and no direct infant antibiotic exposure up to 6 weeks of age (n = 49, IP-ABX); and (3) infants who did not meet the criteria for inclusion in the No-ABX, Neo-ABX or IP-ABX group. Upon study completion, all infants in group 3 were those who could have been exposed to maternal antibiotics during breastfeeding in the postnatal prevaccination period, so we called this group the postnatal exposure group (PN-ABX, n = 30). Researchers conducting the serological assays, omics analyses and other data analyses were blinded to antibiotic exposure group allocations. Participant data were recorded in a secure REDCap (https://project-redcap.org/) database.
Sample collection
Faecal samples for metagenomics were collected from the nappies (diapers) of each infant by parents at week 1 (days 2–10 of life) and week 6 (days 38–56 of life) and placed into Zymo DNA/RNA Shield faecal collection tubes (Zymo Research) before being stored at −80 °C. Blood (3–5 ml) was obtained from infants at approximately 6 weeks, 7 weeks, 7 months and 15 months. At 6 weeks, 1–3 ml of whole blood was placed into BD Vacutainer serum tubes, and 2 ml was placed in a PAXgene blood RNA tube (BD) for RNA preservation and stored at −80 °C. At 7 weeks, up to 1 ml of blood was collected in a BD Vacutainer K3 EDTA tube, 2 ml was collected for flow cytometry analysis, and the remainder (up to 2 ml) was placed into a PAXgene (BD) tube. At 7 months and 15 months, 3–5 ml of blood was collected into BD Vacutainer serum tubes.
Vaccine schedule
Infants were vaccinated with PCV13 (Prevenar 13, Pfizer) and the combination 6-in-1 vaccine (Infanrix Hexa, GSK) at 6 weeks and 4 and 6 months of age and with the oral rotavirus vaccine (Rotarix, GSK) at 6 weeks and 4 months of age as per the NIP Schedule for South Australia as of 2017. Most of the infants (86%) were also vaccinated with the hepatitis B vaccine at birth (either H-B-Vax II Paediatric or Engerix B Paediatric). Infants in this study were also vaccinated with the meningococcal B vaccine (4CMenB/Bexsero, GSK) at 7 weeks, 4 months and 6 months of age, with a booster dose at 12 months of age. The meningococcal ACWY (Nimenrix, Pfizer) and the measles, mumps, rubella (M-M-R II, Merck Sharp & Dohme; or Priorix, GSK) vaccines were administered at 12 months of age as part of the standard NIP schedule (although antibody responses were not evaluated for these vaccines in this study). In this study, the booster dose of PCV13 was provided at 15 months rather than the current NIP scheduled time point of 12 months of age. All vaccines were administered intramuscularly except for the oral rotavirus vaccine. All vaccines were administered in line with Australian Immunisation Handbook recommendations.
Pneumococcal polysaccharide-specific IgG
Antibodies against PCV13 serotypes were measured as previously described13,15,16, with modifications. In brief, Bio-Plex COOH-microspheres (6.25 × 106, 500 µl, Bio-Rad) were conjugated overnight with optimized doses of DMTMM-modified pneumococcal polysaccharides (ATCC) in 1× PBS pH 7.2 (Life Technologies) as per Schlottmann et al.41. The optimal coating doses were a 1:1 dose, 500 µl microspheres with 500 µl DMTMM-modified polysaccharides for serotypes 1, 3, 5, 7F and 9V; and a 1:0.5 dose, 500 µl microspheres with 250 µl DMTMM-modified polysaccharides for serotypes 4, 6A, 6B, 14, 18C, 19A, 19F, 23F and 11A.
A ten-step, three-fold serial dilution of the international reference standard for human antipneumococcal capsule, 007sp (NIBSC), two in-house quality controls and test samples were diluted in adsorbent buffer containing 0.05% Tween-20, 2% N-butyl benzenesulfonamide (Sigma-Aldrich) and 2.5 µg ml−1 CWPS Multi (SSI Diagnostica) for 30 min to remove non-specific antibodies before incubation with polysaccharide-conjugated microspheres. Equal volumes (25 µl) of diluted samples and bead-mix containing 4,000 microspheres per region were combined on a MultiScreen filter plate (Merck) prewetted with 50 µl of 0.05% PBS with Tween-20 (PBS-T). Plates were incubated at room temperature in the dark on an orbital shaker at 500 rpm for 20 min. They were then washed twice with 100 µl 0.05% PBS-T, followed by addition of 100 µl (1:150) RPE-conjugated goat anti-human IgG Fc secondary antibody (Jackson ImmunoResearch Laboratories Inc.) and incubation for a further 20 min. Following washing, the beads were resuspended in 125 µl 0.05% PBS-T and read on a Bio-Plex-200 machine (Bio-Rad).
Diphtheria, tetanus and pertussis-specific IgG
Antibody concentrations specific to pertussis toxoid, pertactin, filamentous haemagglutinin, tetanus toxoid and diphtheria toxoid were measured as previously described14,42.
Hib-PRP- and HB-sAg-specific IgG
Antibodies against Hib-PRP and HB-sAg were measured as previously described for diphtheria, tetanus and pertussis (DTP)14,42, changing only the antigen-coated microspheres, quality controls and reference sera used. In brief, DMTMM-modified41 Hib-PRP 12/306 (NIBSC) was conjugated to Bio-Plex COOH-microspheres (Bio-Rad) as outlined above for pneumococcal polysaccharides, with an optimal dose of 1:1, 500 µl microspheres with 500 µl DMTMM-modified polysaccharide. Recombinant HB-sAg subtype adr (Prospec Bio) was conjugated as previously described for DTP antigens, with an optimal coating concentration of 10 µg ml−1. A ten-step, three-fold serial dilution of the international standard for anti-H. influenzae type b, 09/222 (NIBSC), was used as the reference serum, after anti-HB-sAg titres had been determined in that serum. Anti-HB-sAg titres in 09/222 were quantified against the second international standard for anti-HB-sAg, 07/164 (NIBSC), using the final Bio-Plex protocol; they were further validated using the current gold-standard method, the Alinity i Anti-HBs kit (PathWest), which showed excellent concordance (coefficient of variation less than 5%).
IgA Serology
Rotavirus IgA antibodies were measured by validated enzyme-linked immunoassay, as previously described17,18.
High and low vaccine responders
High and low vaccine responders were defined as infants with IgG titres in the 75th percentile and 25th percentile, respectively, for each of the antigens assessed.
Multiplex opsonophagocytosis method
Multiplex opsonophagocytosis assays were performed on three serotypes (6B, 9V and 18C) using a previously published method43,44. Serum samples were heat-inactivated at 56 °C for 30 min; then, serial dilutions were made, and 20 μl per well of each dilution was tested in duplicate in 96-well u-bottomed microtitre plates (Interpath Services). Serial three-fold dilutions were used. Frozen aliquots of target pneumococci were thawed, washed with opsonization buffer (HBSS with magnesium and calcium, and 0.1% gelatin), and diluted to approximately 5 × 104 CFU ml−1 of each serotype. All serotypes were then pooled in one bacterial suspension, and 10 μl of the bacterial suspension was added to each well. Plates were incubated at room temperature with shaking (mini Orbital Shaker; Bellco Biotechnology) at 700 rpm for 30 min. Then, 10 μl of baby rabbit complement and 40 μl of HL60 cells (approximately 1 × 107 cells ml−1) were then added to each well (the HL60 cells had been washed twice with HBSS before use). Plates were incubated in a tissue culture incubator (37 °C in a 5% CO2 atmosphere) with shaking at 700 rpm. After 45 min, plates were placed on ice to stop the reaction, and 10 μl of the final reaction mixture was spotted onto Todd-Hewitt agar plates supplemented with 0.5% yeast extract. An overlay agar containing one of the selective antibiotics (selective markers for target bacteria) was added to one of each of the Todd-Hewitt agar plates, which was then incubated overnight at 37 °C with 5% CO2. The number of bacterial colonies was enumerated using a ProtoCol3 automatic colony counter (Synbiosis). Results were expressed as opsonization indices, defined as the interpolated dilutions of serum that killed 50% of bacteria. The lower limit of detection in the assay was 4. The opsonization indices of samples that did not kill 50% of bacteria were reported as 2 for purposes of analysis. A cut-off of 8 was used to define a positive response.
Human serum bactericidal activity assays
Human serum bactericidal activity assays were performed on 140 matched study participant serum samples (from blood draws at 6 weeks, 7 months and 15 months of the study) as described previously45. Briefly, heat-inactivated study participant sera were serially diluted two-fold in assay buffer and inoculated with approximately 1 × 103 CFU of N. meningitidis strain NZ98/254. Exogenous human complement (25% v/v, Pel-Freez Biologicals) was added, and assay microtitre plates were incubated at 37 °C with 5% CO2 for 60 min; then, samples were plated on GC agar base plates prepared using the tilt method and incubated at 37 °C with 5% CO2 overnight to enable CFU enumeration. Bactericidal titres are reported as the reciprocal of the lowest dilution of participant serum that induced at least 50% killing relative to the no-serum control. The serologic correlate of protection for N. meningitidis is considered to be a human serum bactericidal activity titre greater than or equal to 4 (ref. 22). Statistical analyses were performed using one-way analysis of variance and two-tailed paired Student’s t-test.
Infant blood preparation for flow cytometry
For flow cytometry analysis, 450 µl of infant blood was transferred from a BD Vacutainer K3 EDTA tube into a sterile 10-ml tube within 3 h of collection, and red blood cells were lysed by addition of 4.5 ml of 1× BD Pharm Lyse for 15 min at room temperature. Cells were then pelleted by centrifugation at 200g for 5 min, the supernatant was aspirated, and the cell pellet was resuspended in 10 ml flow cytometry buffer and centrifuged again. Cells were subsequently resuspended in 150 µl flow cytometry buffer and divided equally into three tubes for staining. Fc receptors were first blocked using Human TruStain FcX (BioLegend) in Brilliant Stain Buffer (BD) for 10 min on ice. Three panels were designed for same-day flow cytometry analysis (see Supplementary Table 8 for a complete list of cell subsets identified in each panel). Panel 1 was a pan-leukocyte panel enabling identification of neutrophils, monocytes, dendritic cells, eosinophils, basophils, T cells, B cells and natural killer cells. Cells were stained with a cocktail of anti-CD3 FITC (HIT3a, 1:20, BD Biosciences), anti-CD11c PE (B-ly6, 1:20, BD Biosciences), anti-HLA-DR APC-H7 (G46-6, 1:40, BD Biosciences), anti-CD15 V500 (HI98, 1:80, BD Biosciences), anti-CD45 BUV395 (HI30, 1:80, BD Biosciences), anti-CD16 BV421 (3G8, 1:143, BD Biosciences), anti-CD14 AF647 (Mop-9, 1:143, BD Biosciences), anti-CD123 BV711 (9F5, 1:143, BD Biosciences), anti-CD56 PE-Cy7 (5.1H11, 1:40, BioLegend), anti-CD19 BV605 (HIB19, 1:80, BioLegend), anti-CD20 BV786 (2H7, 1:80, BioLegend), anti-FcεRIα PerCP Cy5.5 (AER-37, 1:80, BioLegend) and anti-Siglec8 PE Dazzle (7C9, 1:143, BioLegend). Panel 2 enabled characterization of around 20 functional subsets of B and T cells including naive, central memory and effector memory CD4+ and CD8+ T cells; regulatory T cells; and naive, memory and plasma B cells. The antibody cocktail consisted of anti-CD19 PE (HIB19, 1:10, BD Biosciences), anti-CCR7/CD197 AF647 (150503, 1:10, BD Biosciences), anti-CD127 BV421 (A019D5, 1:40, BioLegend), anti-CD38 BV510 (HIT2, 1:40, BD Biosciences), anti-HLA-DR APC-H7 (G46-6, 1:40, BD Biosciences), anti-CD3 BUV395 (UCHT1, 1:40, BD Biosciences), anti-CD4 BV605 (RPA-T4, 1:80, BD Biosciences), anti-CD8 Cy7 (RPA-T8, 1:80, BD Biosciences), anti-IgD PerCP Cy5.5 (IA6-2, 1:80, BD Biosciences), anti-CD27 BV711 (L128, 1:143, BD Biosciences), anti-CD25 PE Dazzle594 (M-A51, 1:80, BioLegend), anti-CD20 BV786 (2H7, 1:80, BioLegend), and anti-CD45RA FITC (HI100, 1:143, BioLegend). Panel 3 was designed to assess specific T helper (TH) subsets, including TH1, TH2, TH9, TH17, TH22 and natural killer T cells, using anti-CD3 BUV395 (UCHT1, 1:50, BD Biosciences), anti-CD4 BV605 (RPA-T4, 1:100, BD Biosciences), anti-CD8 PE Cy7 (RPA-T8, 1:100, BD Biosciences), anti-CXCR3 BV421 (1C6/CXCR3, 1:100, BD Biosciences), anti-CCR10 APC (1B5, 1:100, BD Biosciences), anti-CCCR6 BV711 (11A9, 1:50, BD Biosciences), anti-CCR4 PECF594 (1G1, 1:50, BD Biosciences), anti-CD45RA FITC (HI100, 1:100, BioLegend) and CD1d-Tet PBS57 PE (1:5,000, NIH). See Supplementary Table 8 for a comprehensive list of antibody sources, catalogue numbers and fluorophores.
After each antibody cocktail had been added, samples were incubated in the dark on ice for 30 min before being washed and resuspended in 300 µl of flow cytometry buffer. Panel 1 was added to a BD Trucount FACS tube (BD), whereas panels 2 and 3 were added to round-bottomed 5-ml FACS tubes alongside 10 µl liquid counting beads (BD). Counts were generated as previously described25. Dead cells were stained by addition of DNA binding dye DAPI and were excluded from analysis. Samples were run on a FACSymphony (BD) flow cytometer using FACSDiva software, and results were analysed using FlowJo v.10.6.1 (Tree Star).
Extraction of RNA from blood
RNA extraction and elimination of genomic DNA were carried out using a PAXgene Blood RNA kit (Qiagen) per the manufacturer’s instructions, with final elution into 80 μl RNase-free water. A further RNA precipitation reaction was carried out. Briefly, RNA was resuspended in 2.5 × 100% ethanol and 10% sodium acetate and spun at 12,000g for 30 min at 4 °C. Samples were washed in 75% ethanol. Pellets were air-dried and resuspended in 40 µl of RNase-free water. The total RNA yield was determined using a Bioanalyzer 2100/TapeStation (Agilent) and Qubit (Thermo Fisher Scientific). Before library preparation, 0.2 ng of synthetic reference RNA standards (Sequins, Garvan Institute) were added to 250 ng of RNA.
RNA library preparation
Total RNA was converted to strand-specific Illumina-compatible sequencing libraries using a Nugen Universal Plus mRNA-Seq library kit from Tecan, with Any Deplete probes and custom gamma globin depletion per the manufacturer’s instructions (MO1442 v.2). Briefly, 250 ng of total RNA was polyA-selected, and the mRNA was fragmented before reverse transcription. Any deplete and custom gamma globin depletion was performed before strand selection II complementary DNA (cDNA) synthesis with pooled gamma depletion oligos, per the manufacturer’s instructions. The resulting cDNA was end-repaired, followed by ligation of Illumina-compatible barcoded sequencing adaptors. The cDNA libraries were strand-selected and PCR amplified for 12 cycles before quality control assessment using a TapeStation (Agilent Technologies). Library concentrations were assessed using a Qubit fluorescence assay (Thermo Fisher Scientific). Sequencing pools were generated by mixing equimolar amounts of compatible sample libraries on the basis of the Qubit measurements. The library pool was sequenced using an Illumina NovaSeq 6000 S1 flow cell with 2 × 100-bp paired-end reads.
RNA sequencing analysis
Sequence read quality was assessed using FastQC v.0.11.4 (ref. 46) and summarized with MultiQC v.1.8 (ref. 47), followed by quality control with Trimmomatic v.0.38 (ref. 48) with a window size of 4 and an average quality score of 25. Reads of fewer than 50 nucleotides after trimming were discarded. Reads that passed all quality control steps were then aligned to the human genome (GRCh38) using HISAT2 v.2.1.0 (ref. 49). The gene count matrix was generated with FeatureCounts version 1.5.0-p2 (ref. 50) using the union model with Ensembl v.93 annotation. Data were then imported into R v.4.2.0 for further analysis. Counts were normalized using the trimmed mean of M values method in EdgeR v.3.38 (ref. 40), followed by multidimensional scaling analysis and differential gene expression analysis performed with the glmLRT function. For differential expression analysis comparing week 6 and week 7, a paired design was used, fitting each participant into the model as well as each time point. As each participant’s samples were processed together, this intrinsically controlled for batch effects. For non-paired comparisons, processing batch and infant sex were fitted into the model to adjust for those effects. Unknown sources of variation were identified using svaseq v.3.46.0 (ref. 38) and adjusted for before multidimensional scaling and gene set variation analysis. The gene count matrix was filtered to remove genes with a count per million of less than 5 in 15 samples before differential expression analysis. The Benjamini–Hochberg method was used to correct for multiple comparisons. Gene set enrichment analysis was carried out using the fgsea R package v.1.22.0 (ref. 39) with predefined BTMs24, with input ranks as fold changes calculated by edgeR. Gene set variation analysis was used to calculate a per-sample activity score for each of the BTMs (excluding unannotated modules labelled ‘TBA’) using R Bioconductor package GSVA v.1.44.1 (ref. 51). Spearman correlation analysis was performed using the Hmisc v.4.7-0 package in R v.4.2.
DNA extraction from faecal samples
Two millilitres of the faecal suspension from the Zymo DNA/RNA Shield faecal collection tube (Zymo Research) was transferred into a sterile microcentrifuge tube and pelleted by centrifugation at 16,200g for 20 min at 4 °C. The supernatant was then removed, and DNA extraction was performed on the faecal pellet using the PowerLyzer PowerSoil Kit protocol (Qiagen) as described previously52.
Quantitative real-time PCR analysis to determine bacterial load in infant stool samples
Total bacterial load in the samples was determined by quantitative PCR (qPCR) of the 16S ribosomal RNA (rRNA) gene using a SYBR-based assay, as described previously53. Each qPCR reaction comprised 1× PowerUP SYBR Green Master Mix (Applied Biosystems by Thermo Fisher Scientific), 0.2 µM of each forward and reverse primer, and 1 µl of DNA in a 35-µl total reaction volume. Each reaction was then aliquoted into three technical replicates of 10 μl for qPCR analysis. Numbers of 16S rRNA gene copies were calculated on the basis of a standard curve generated by serial dilutions of Escherichia coli genomic DNA and normalized against the faecal weight (in mg) of each sample.
Metagenomic library preparation
Shotgun metagenomics libraries were generated as previously described with modifications54. Briefly, libraries were generated for samples with a DNA concentration of at least 0.2 ng µl−1 using a Nextera DNA XT Library Preparation Kit and indexed using a Nextera XT Index Kit v.2 (Illumina). Quality control checks for DNA fragment size following PCR enrichment were performed using a DNA 1000 chip on a Bioanalyzer (Agilent Technologies). All libraries with a concentration of 5 nM and above were pair-end sequenced (2 × 151 bp) on the NovaSeq 6000 platform (Illumina).
Metagenomic data analysis
Read quality was assessed using FastQC v.0.11.4 (ref. 46) and summarized with MultiQC v.1.8 (ref. 47). Adaptors were removed with cutadapt v.1.18, and quality-filtering and trimming were performed using Trimmomatic v.0.38 (ref. 48) with a window size of 4, average quality score of 25 and minimum length of 50 nucleotides. The percentage of human DNA per sample was estimated using Kraken2 v.2.1.1 (ref. 55). MAG bins were generated by assembling each of the 409 samples individually using metaSPAdes v.3.15.2 (ref. 56). Reads were then aligned back to this assembly catalogue, and the mapped output was used for binning with MetaBAT2 v.2.15 (ref. 57). Counts per bin were then calculated on the basis of SAM files output from bowtie2 v.2.5.0. Each MAG bin was taxonomically classified using GTDB-Tk v.2.1.0 and reference data release 207 (ref. 58). This MAG bin count matrix was then imported into R for further analyses. Count data were transformed to relative abundances before being visualized using bar plots and were normalized by CLR elsewhere as implemented in R library PhyloSeq v.1.28. Differential abundance analysis was performed with LinDA as implemented in R library MicrobiomeStat v.1.18 with raw count data as input. Counts were filtered for taxa detected in at least 5% of samples before testing with the prev.filter flag in LinDA59. Spearman correlation analyses were performed using functions in base R with the CLR-normalized data as input. Alpha and beta diversity analyses were performed in R using PhyloSeq v.1.28 (ref. 60). Metagenomic samples were clustered into CSTs with Dirichlet multinomial mixture modelling (R package DirichletMultinomial v.1.38.0), with the optimal number of clusters calculated using the lowest LaPlace approximation. Quality-filtered reads were also used as input to HUMAnN v.3.0 (ref. 61) for generation of microbial pathway data and summed to represent counts for each MetaCyc pathway and gene ontology term (biological processes only). Pathway count data were also CLR-normalized. To account for variation in 16S rRNA copy numbers between samples, which could have biased qPCR estimates of bacterial load, a mean copy number per sample was calculated on the basis of the species-level relative abundance profiles from the shotgun metagenomic data and the copy number from rrnDB62. Antimicrobial resistance genes were analysed by prediction of the encoded protein sequences from MAGs using prodigal v.2.6.3, clustering at 95% identity with cd-hit v.4.8.1, and conversion to amino acid sequences using transseq to generate a non-redundant predicted protein catalogue. The non-redundant catalogue was aligned to the Comprehensive Antibiotics Resistance Database (v.3.2.7) using the Resistance Gene Identifier (v.5.2.1) with default parameters except the addition of ‘–exclude_nudge’. Per-gene counts were generated by aligning reads using bowtie2 against the gene catalogue and generating a per-sample count with SAMtools v.1.17. Reads per kilobase per million mapped reads values were generated in R, summed by antibiotic class and compared between groups using a Wilcoxon rank sum test, controlling for multiple comparisons with the Benjamini–Hochberg method. Causal mediation analysis was performed in R with the mediation R package (v.4.5.0)28.
Preclinical mouse experiments
C57BL/6J mice (The Jackson Laboratory; RRID: IMSR_JAX:000664) were bred and maintained under specific and opportunistic pathogen free (SPF) conditions or gnotobiotic conditions at the South Australian Health and Medical Research Institute (SAHMRI). SPF mice were kept in cages with free access to commercial pelleted food and water and were housed under standardized conditions with regulated daylight, humidity and temperature. Gnotobiotic (GF) C57BL/6 mice (Translational Research Institute) were housed in positively pressurized HEPA-filtered isolators at the SAHMRI Preclinical, Imaging and Research Laboratories (PIRL) with access to autoclaved commercially pelleted food and sterilized water ad libitum, with regulated 12 h day/night and regulated humidity (45–75%) and temperature (18–24 °C). All experiments were carried out in accordance with protocols approved by the SAHMRI Animal Ethics Committee. Each vaccine experiment was conducted as an independent experiment. Equal numbers of male and female mice were used. Young mice (aged approximately 3 weeks) or dams (aged 8–16 weeks) and pups were randomized to the treatment groups. Investigators were not blinded to the mouse treatment groups. The appropriate sample size for the murine experiments was determined on the basis of data from our previous mouse vaccine experiments.
GF mouse dams and their pups remained GF (as determined by culturing of faecal samples and 16S rRNA gene qPCR) or were colonized as previously described63 by means of oral gavage with a consortium of Bifidobacterium species consisting of B. breve (DSM 20213, Leibniz Institute), B. longum subspecies longum (DSM 20219) and B. longum subspecies infantis (DSM 20088) at approximately 14 days postcoitus (E14) and at day 7 and day 14 postbirth. Successful colonization was confirmed by culturing and Sanger sequencing. For culturing, Bifidobacterium isolates were plated on MRS agar (Thermo Fisher Scientific) enriched with 1 mg ml−1 cysteine, and single colonies were isolated, confirmed using a MALDI Biotyper (Bruker) and cultured for 48 h at 37 °C using BD GasPak pouch systems (BD). To assess whether administration of a commercially available probiotic could restore antibody responses to PCV13, GF mice were colonized by means of oral gavage at day 21 of life with 2 × 107 CFU L. acidophilus and B. bifidum isolated from commercial probiotic Infloran (Laboratorio Farmaceutico). Further GF mice were similarly colonized with 2 × 107 CFU with each of the strains individually. L. acidophilus and B. bifidum cultures were prepared by serially diluting Infloran on to MRS agar plates (Thermo Fisher Scientific) enriched with cysteine for L. acidophilus or reinforced clostridial agar (Thermo Fisher Scientific) for B. bifidum, under anaerobic conditions. Single colonies were identified using a MALDI Biotyper (Bruker). Further GF mice, aged 21 days, were colonized with Lactobacillus murinus (DSM 20452), Enterococcus gallinarum (DSM 24841), Akkermansia muciniphila (DSM 22959), Bacteriodes acidifaciens (DSM 15896), Blautia producta (DSM 2950), Lactobacillus plantarum (ATCC 202195, The Global Bioresource Centre) or Enterobacter cloacae1.
To produce littermate wild-type and knockout mice for the experiments in Myd88−/−, Tlr2−/− and Tlr4−/− mice, pairs of heterozygous Myd88+/−, Tlr2+/− and Tlr4+/− dams (8–12 weeks of age) were bred with heterozygous studs. Equal numbers of wild-type and knockout offspring were cohoused for studies.
Vaccination of mice
Mice were vaccinated with 3 μg PCV13 (Pfizer), 8 μg Infanrix Hexa combination vaccine (GSK) or 17.5 μg Bexsero meningococcal serogroup B vaccine (GSK) intraperitoneally at day 28 postbirth and boosted with the same dose 2 weeks later. Control mice were mock-vaccinated with 100 μl of PBS. Four to eight sex-matched mice per group were vaccinated or mock-vaccinated in each experiment. Sera were collected 2 and 4 weeks postimmunization in each experiment and also at 6 and 8 weeks in selected experiments.
Murine faecal sampling
Faecal samples were collected from mice at the start of each experiment and at each experimental time point to confirm the GF or colonization status of the mice. Faecal pellets were aseptically collected, immediately snap-frozen and stored at −80 °C.
DNA extraction from murine faecal samples
Faecal samples were individually weighed, resuspended in 1 ml of phosphate-buffered saline (PBS) (pH 7.2) by vortexing and pelleted by centrifugation at 13,000g for 5 min. Microbial DNA extraction was performed on the faecal pellets using a DNeasy PowerSoil Kit (Qiagen) with the following minor modifications: faecal pellets were resuspended in 750 μl of PowerSoil bead solution and 60 μl solution C1 in a PowerSoil bead plate and incubated at 65 °C for 10 min before bead beating as per previously described protocols63.
16S rRNA gene sequence analysis
Murine faecal DNA extracts were used to generate amplicons of the V4 hypervariable region of the 16S rRNA gene as described previously63. Sequencing of the amplicon library was performed using an Illumina MiSeq system (2 × 300-bp run). Paired-end 16S rRNA gene sequences were demultiplexed and imported into QIIME2 (release 2023.2) for processing64. Sequences were error-corrected, and counts of error-corrected reads per sample, which we refer to herein as exact sequence variants, were generated with DADA2 (ref. 65). Taxonomy was assigned to sequences with the sklearn plugin for QIIME2 with an 80% confidence threshold, using the SILVA v.138 database66. Exact sequence variant counts were converted to relative abundance, which was plotted in R v.4.2.0 with ggplot2 v.3.3.6.
Quantitative real-time PCR analysis to determine bacterial load
Real-time PCR was performed on a Quant Studio 7 Flex Real-time PCR system (Thermo Fisher Scientific) using primers that targeted conserved regions of the 16S rRNA gene as described in ref. 67. Each reaction comprised 5 μl of SYBR Green qPCR (Thermo Fisher Scientific), 0.2 μM of each forward and reverse primer, 1 μl of DNA template and sterile water to a total reaction volume of 22 μl. A no-template DNA control, with the DNA substituted with 1 μl sterile water, was included in each run. Real-time PCR reactions for each sample were performed in duplicate, consisting of 10 μl per reaction. The amplification program was 50 °C for 2 min, 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. Melting curve analysis was performed at the end of the program. The total bacterial load in each sample was quantified against a standard curve of serial dilutions of E. coli genomic DNA, which were performed in the same run, and was detectable within a range of 3 to 3 × 106-equivalent E. coli cells. Numbers of cells were then calculated against the faecal weight used for each sample. A Mann–Whitney test was used to assess statistical significance.
Assessment of antibody responses in mice
Levels of antigen-specific IgGtotal IgG1, IgG2c, IgG3 and IgM were measured by enzyme-linked immunosorbent assay (ELISA) in serum collected from each vaccine experiment. Briefly, Nunc MaxiSorp ELISA plates (Sigma) were coated either with CRM197 (0.5 μg ml−1, Finabio), PCV13 vaccine (0.05 μg ml−1, Pfizer), pneumococcal polysaccharide (4 μg ml−1 PPS1, 161-X, ATCC), PPS3 (4 μg ml−1 169-X, ATCC), PPS6B (4 μg ml−1 105-X, ATCC), PPS9V (4 μg ml−1 305-X, ATCC) or diphtheria toxoid (1 μg ml−1, List Biological Laboratories), tetanus toxoid (1 μg ml−1, List Biological Laboratories), pertussis toxoid (1 μg ml−1, List Biological Laboratories) or Bexsero vaccine (2 μg ml−1, GSK). Plates were washed with 0.5% Tween-20/PBS and blocked with 1× ELISA buffer (eBioscience). Mouse serum samples were diluted in 1× ELISA buffer and incubated on blocked plates. Antigen-specific serum antibodies were detected using horseradish peroxidase (HRP)-conjugated antibodies: anti-mouse IgG (Novex) or anti-mouse IgG1 (Abcam) at a dilution of 1:1,000 or IgG2c, IgG3 or IgM (Abcam) at 1:500. HRP activity was detected using tetramethylbenzidine substrate (Thermo Fisher Scientific) and stopped with 2 N H2SO4. Developed plates were recorded using a Synergy HTX microplate absorbance reader (BioTek) at 450 nm with correction at 595 nm by subtraction. Data were analysed using Prism 10 (GraphPad).
Tissue dissociation and flow cytometry analysis
Single-cell suspensions of murine spleen or lymph nodes were prepared by mechanical dissociation and filtration through a 70-µm nylon filter (Merck Millipore). Cells were stained with Fixable Viability Stain 780 (BD Biosciences) and murine Fc-blocking antibody clone 2.4G2 (1:100, BD Biosciences) in PBS for 15 min. Cells were washed and then stained with fluorochrome-conjugated antibodies in FACS buffer (PBS, 0.1% bovine serum albumin, 2 mM EDTA) against CD3 (142-C11, 1:300, BD Biosciences), CD4 (RM4, 1:300, BD Biosciences), CD8 (53-6.7, 1:600, BD Biosciences), CD19 (1D3, 1:300, BD Biosciences), CD38 (90, 1:600, BioLegend), CD44 (IM7, 1:600, BD Biosciences), CD62L (MEL-14, 1:600, BD Biosciences), CD95 (J02, 1:300,BD Biosciences), CD138 (281-2, 1:600, BD Biosciences), B220 (RA3-6B2, 1:600, BD Biosciences), CXCR5 (L138D7, 1:200, BioLegend), GL7 (GL7, 1:600, BD Biosciences), IgD (11-26 c.2a, 1:300, BioLegend), IgM (R6-60.2, 1:300, BD Biosciences), PD-1 (RMP1-30, 1:300, BD Biosciences) and streptavidin-PECF549 (1:400, BioLegend).
For assessment of CRM+ GC B cells, cells were stained with Fixable Viability Stain 780 (1:1000, BD) and murine Fc block (1:100, BD) in PBS for 15 min; then, they were washed and incubated with biotinylated CRM (1:100, Fina BioSolutions) in FACS buffer. Surface antibody cocktails were added directly after staining for 20 min at 4 °C. Data were acquired on a BD FACS Symphony and analysed using FlowJo v.10. For assessment of CD86, CD80 and MCHII expression, single-cell suspensions of spleen or lymph nodes were stained with Zombie Aqua Fixable Viability Stain 780 (1:1000, BioLegend) and murine Fc-blocking antibody clone 2.4G2 (1:100, BD Biosciences) in PBS for 15 min. Cells were washed and then stained with fluorochrome-conjugated antibodies in FACS buffer (PBS, 0.1% bovine serum albumin, 2 mM EDTA) against CD19 (ID3, 1:200, BD), CD3 (145-2C11, 1:150, BD Biosciences), CD8 (53-6.7, 1:200, BD), NK1.1 (PK136, 1:200, BD), Ly6G (1A8, 1:600, BD), B220 (RA3-6B2, 1:3000, BD), CD11b (M1/70, 1:500, BD), CD11c (HL3, 1:400, BD Biosciences), MHCII (M5/114.15.2, 1:150, Miltenyi), CD86 (PO3.3, 1:150, Miltenyi) and CD80 (16-10A1, 1:150, Tonbo Biosciences).
Immunofluorescence staining and imaging
Mediastinal lymph nodes were fixed in 1 ml 4% paraformaldehyde (Sigma) for 4 h and then incubated in 1 ml 30% sucrose solution overnight at 4 °C before being embedded in OCT (Tissue Tek) as described previously68. Slides were blocked in blocking buffer (normal goat serum 4% v/v (Southern Cross Science), normal donkey serum 4% v/v (Stratech Scientific APAC) and bovine serum albumin 15% v/v (AusGeneX)) for 2 h at room temperature. Slides were permeabilized with Triton X-100 2% v/v (Sigma) and stained with CD16/32 (1:100, BD Biosciences) and CD21/35 clone 7E9 (1:200), GL7 (1:200), CD3 clone 17A2 (1:200) and IgD clone 11-26c.2a (1:200) (all from BioLegend). Images were acquired using a Leica TCS SP8X confocal microscope and analysed using Fiji69 in ImageJ.
Statistical analysis
The primary objective of the AIR study was to determine whether antibody responses to the PCV13 vaccine at 7 months of age were different between antibiotic-exposed and unexposed groups, specifically comparing the following:
-
1.
The percentage of participants achieving a seroprotective antibody response (greater than or equal to 0.35 μg ml−1) against pneumococcal polysaccharide serotypes in the PCV13 vaccine at approximately 7 months of age.
-
2.
Geometric mean concentrations in μg ml−1 of anti-PPS IgG as measured by serum assay at approximately 7 months of age.
As secondary outcomes, we compared PCV13 vaccine responses at 15 months of age and antibody responses to the Infanrix Hexa 6-in-1 vaccine (6-in-1, DTPa-HepB-IPV-Hib) and the Rotarix oral rotavirus vaccine in antibody-exposed and unexposed infants at both time points. Multivariable linear regression adjusting for baseline prevaccination titres and sex was used to test for any association between antibiotic exposure group and IgG geometric mean concentrations for each of the vaccine antigens assessed. Further longitudinal analyses were conducted by fitting a mixed effects multivariable linear regression, including a random effect for participant ID, to assess the interaction between time point and antibiotic exposure group on IgG geometric mean concentrations for each of the vaccine antigens. Multivariable logistic regression was used to test for any association between antibiotic exposure group and the proportion of infants achieving a seroprotective response. Statistical significance was defined as P < 0.05. Further sensitivity analyses were conducted to assess the robustness of associations to adjustment for probiotic consumption, formula consumption, baseline IgG/IgA concentration, infant sex and gestation at delivery. The AIR study also included a range of exploratory systems immunology assessments for which analyses were undertaken in line with best practices for each data type and as described in the sections above. Measurements were taken from distinct samples. All statistical tests throughout were two-sided; the specific statistical analyses used are specified in the figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.