On the Future of Species: Authoring Life by Means of Artificial Biological Intelligence Adrian Woolfson Bloomsbury (2026)
Biology is undergoing a transformation. After centuries of studying life as it evolves naturally, researchers are now using a combination of computation and genome engineering to intervene, generating new proteins and even whole bacteria from scratch. The use of artificial-intelligence tools to design biological components, an approach known as generative biology, is set to turbocharge this area of research. Just last year, scientists used AI-assisted design to produce artificial genes that can be expressed in mammalian cells and, for the first time, an AI program was used to create an entirely synthetic virus.
A new vision for how evolution works is long overdue
This approach is much more than just a series of technical feats. It could transform how life on Earth develops, as biochemist Adrian Woolfson describes in his latest book. On the Future of Species provides a sweeping account of the history and science behind this transformational technology, from the first gene-sequencing efforts to the rise of AI-powered techniques. His vision of what biology is becoming gives the book a coherence that is sometimes lacking in broad surveys of synthetic biology, which often represent only the current science.
One of the central themes in Woolfson’s book is that, thanks to computational tools, life’s patterns (whether the order in which genes are arranged on chromosomes or the physical traits that are encoded by genes) are becoming increasingly predictable and manipulable. This is making it ever simpler for researchers to tinker with an organism’s DNA to achieve a desired result — be it correcting a disease-causing mutation, designing proteins that have never existed in nature or engineering organisms that can clean up pollution.
Brave new world
AI systems can redesign genomes in silico as if they were software, rearranging bases like code. These programs use modelling to predict how the sequence of a gene relates to the structure of the protein it encodes, and therefore the protein’s function. They can also run simulations to test how a redesigned genome might behave.
It’s time to admit that genes are not the blueprint for life
One day, Woolfson theorizes, AI-assisted genome design might enable scientists to produce a computer system that he calls a species catalogue. This repository would contain all of the information needed to design a whole host of possible life forms. Such a catalogue would enable researchers to create organisms for a specific purpose, such as producing innovative drugs and making crops resistant to pests to eliminate the need for pesticides in agriculture.
Eventually, Woolfson notes, researchers hope to achieve what he refers to as artificial biological intelligence — models that can propose complete, working genomes from scratch. Yet there are many limitations that prevent current models from achieving this. For one thing, it is still frustratingly difficult to predict how the expression of each individual gene affects the expression of others. Furthermore, organismal development is context-dependent and is therefore hard to consolidate into a neat algorithm. Bee larvae can develop into workers or queens depending on nutrition, for example, and people learn to speak with the accent of their parents and peers. Neither of these characteristics can be predicted by computer models.
And then, of course, there are the challenges that come with actually building the organism in the laboratory. Just because it is possible to design a unicorn genome doesn’t mean that people will get to ride one. The synthetic yeast genome project, which began in 2006, illustrates some of these difficulties. The genome is a modified version of a natural yeast, Saccharomyces cerevisiae, with tweaks to remove non-essential genetic regions and to make it easy to physically shuffle sections of the genome, among other things. But assembling the synthetic chromosomes has been labour-intensive, with the last few being completed in 2025. Moreover, DNA produced conventionally is condensed as it is created so that it fits into the nucleus; it is difficult to retrospectively stuff whole chromosomes into a cell. The researchers involved are still working on this last problem.
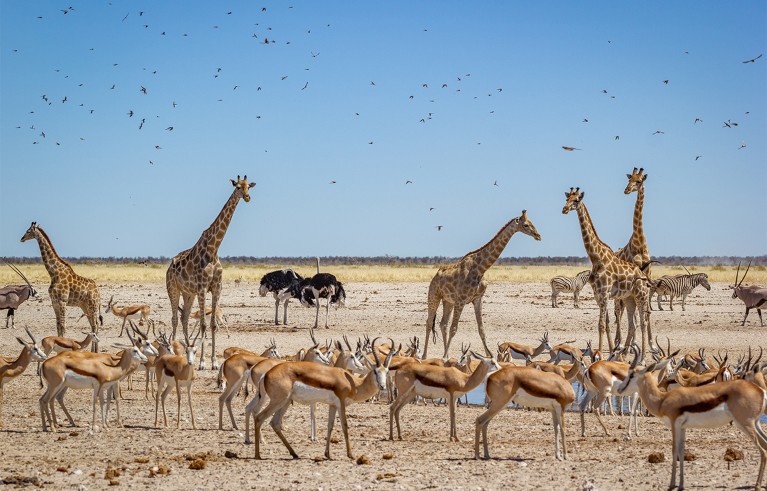

Once extinct, the unique constellation of traits a species carries can never be replicated.Credit: Francesco Riccardo Iacomino/Getty
Another complexity is how evolution limits the possible pathways down which an organism can develop, as a decades-long experiment highlights. Since 1988, researchers have tracked how cultures of Escherichia coli bacteria kept in identical conditions have evolved over many years. Most of the bacterial cultures have gradually accumulated genetic mutations to metabolize the glucose in their feedstock more efficiently. But in 2003, one culture surprised researchers by evolving to instead metabolize another molecule entirely: citrate. This shift required gene duplications and the rewiring of entire metabolic programs. It was probably written into the strain’s destiny early in the experiment, when the first of these mutations occurred, sending this one strain down a distinct evolutionary path from the others.
This shows, Woolfson warns, how small genome edits today could lock in irreversible biological futures. Some choices, once made, eliminate other options. If researchers want to freely design species, they need to watch out for these one-way streets. For example, if scientists edit out a biosynthesis pathway because they think that it isn’t necessary, all future organisms from that lineage will lack it. Too much genome editing might create organisms that cannot evolve, or be evolved, in response to changing environments.