Mice
All experiments involving animals were approved by the Yale University Institutional Animal Care and Use Committee and conducted in compliance with all relevant university, state and federal guidelines. The day of vaginal plug detection was designated as PCD 0.5. The day of birth was designated as PD 0. Fezf2–Gfp (000293-UNC) and Arpp21–Gfp (011848-UCD) transgenic mice were obtained from the Gene Expression Nervous System Atlas55. Cux2-E1–Gfp, Cux2-E2–Gfp and Cux2-E3–Gfp transgenic mice were generated by delivering linearized DNA constructs by pronuclear injection. Three to seven founders from each line were examined for reproducible GFP expression. Zbtb18 KO mice were generated by the Masai laboratory28, and Zbtb18fl/fl mice were generated by the Heng laboratory. Emx1–Cre (The Jackson Laboratory; 005628) mice, Cux2–Cre mice, Neurod6–Cre (also known as Nex1–Cre) mice and CAG–Cat–Gfp (The Jackson Laboratory; 024636) mice were previously generated and described56,57,58,59 (Cre Driver Network at the National Institutes of Health (NIH) Blueprint for Neuroscience Research). Genotyping primers are shown in Supplementary Table 15.
The mice were provided with food and water ad libitum, maintained on a 12-h light/12-h dark cycle, provided veterinary care by the Yale Animal Resources Center and housed in clean groups. All mice used or bred for these experiments were in good health, as approved by the Yale Animal Resources Center and confirmed through regular veterinary monitoring. To maintain genetic diversity, multiple concurrent breeding pairs were maintained and siblings were never mated. Both males and females were used randomly throughout this study.
Although blinding was not relevant for the primary mutant versus control comparison, other aspects ofthe study required careful design to minimize bias. Randomization was implemented during data acquisition. Littermates (WT, HET and KO) were housed together to avoid confounding housing effects on statistical analyses. The experimental cohort comprised age-matched male and female littermates. Including samples from multiple litters further enhanced reproducibility.
Tissue preparation and fluorescence-activated cell sorting
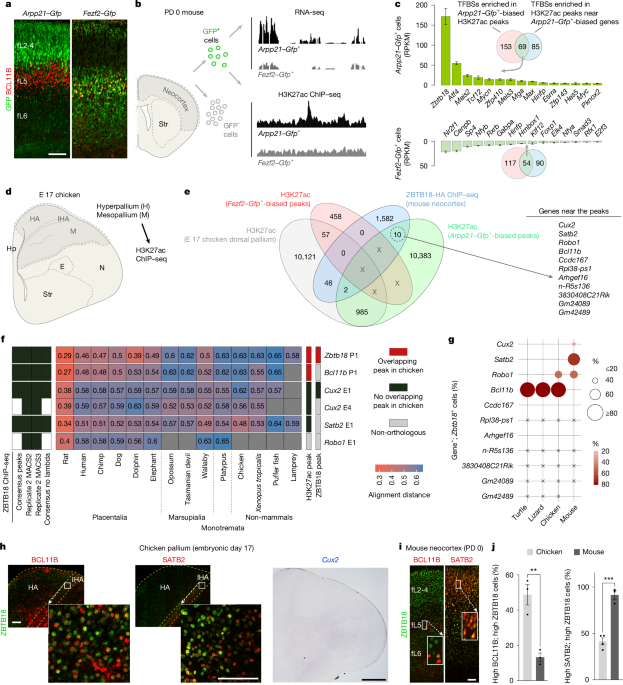
Neocortices from PD 0.5 Fezf2–Gfp, with GFP-expressing neurons enriched in the deep layer (L5–6) predominantly ET neurons (7), and Arpp21–Gfp, with GFP-expressing neurons enriched in predominantly IT neurons55, were dissected under a dissection microscope and minced with a sterile blade. Tails were collected for determining sex and genotyping. Single-cell suspensions from neocortical tissue were prepared by dissociation with a papain-based solution60 and incubated at 37 °C for 15 min with intermittent trituration using autoclaved, fire-polished glass Pasteur pipettes. Cells were then pelleted by centrifugation at 4 °C for 5 min, washed with sterile 1× phosphate-buffered saline (PBS) and filtered through a 40-μm strainer. Cells were then sorted to collect GFP+ cells by fluorescence-activated cell sorting (FACS) using FACSAria II (BD Biosciences) sorter or Beckman Coulter MoFlo sorter. Hibernate solution (Gibco) supplemented with 2% fetal bovine serum (Gibco) was used to collect FACS-sorted cells. Cell preparations were maintained at 4 °C during the entire process. For RNA-seq, immediately after FACS, cells were pelleted by centrifugation at 350g at 4 °C for 10 min, washed with PBS, pelleted and flash-frozen in liquid nitrogen and stored at −80 °C. For ChIP–seq, FACS-sorted GFP+ cells were pelleted by centrifugation at 350g for 10 min at 4 °C, washed with PBS and immediately crosslinked with formaldehyde solution at a final concentration of 1% for 10 min at room temperature. Glycine (AmericanBio) was added at a final concentration of 125 mM, and samples were incubated for 5 min at room temperature to quench crosslinking. Cells were washed again with PBS, pelleted, flash-frozen in liquid nitrogen and stored at −80 °C.
RNA-seq and initial analysis
Total RNA was extracted from FACS-purified cells or neocortical tissue using TRIzol reagent, according to the manufacturer’s instructions. DNase I (Invitrogen) was added to the extracted total RNA, incubated for 15 min at 37 °C to eliminate DNA contaminants and then inactivated according to the manufacturer’s instructions. RNA concentration and integrity were measured using a NanoDrop spectrophotometer (Thermo Fisher Scientific) and TapeStation 2200 (Agilent). Samples with RNA integrity number of 8 or higher were used for subsequent experiments. Libraries were prepared using the TruSeq Stranded Total RNA with Ribo-Zero preparation kit (Illumina), according to the manufacturer’s instructions. Libraries were quality controlled using the TapeStation 2200 (Agilent) and sequenced on the HiSeq 2000 platform (Illumina) at the Yale Center for Genome Analysis (YCGA) to generate 75-bp single-end reads. Sequencing data were quality controlled using FastQC and aligned to the mouse genome (NCBI38/mm10) using TopHat (v.1.0.13) with up to two mismatches61. An average of 40 million uniquely mapped reads were obtained for each sample. Differential expression analysis was performed using the R package DESeq, and principal component analysis was performed using the R package prompt. Differential expression of transcripts was detected using FDR < 0.01.
ChIP–seq and initial analysis
Pooled GFP+ cells from FACS were used for ChIP–seq. A total of 2.5 × 107 cells per condition were crosslinked with a formaldehyde solution (Sigma-Aldrich) at a final concentration of 1% for 10 min at room temperature. L-Glycine (AmericanBio) was added at a final concentration of 125 mM and incubated for 5 min at room temperature to quench the crosslinking. Cells were washed with PBS and then disrupted using lysis buffer I (50 mM HEPES–KOH (pH 7.5), 140 mM NaCl, 1 M EDTA (pH 8.0), 10% glycerol, 0.5% Nonidet P-40 (NP-40), 0.25% Triton X-100 and 1× protease inhibitor) for 20 min at 4 °C and lysis buffer II (200 mM NaCl, 1 M EDTA (pH 8.0), 0.5 mM EGTA (pH 8.0), 10 mM Tris–HCl (pH 8.0) and 1× protease inhibitor) for 10 min at room temperature. Cells were centrifuged at 300g for 15 min at 4 °C, and pellets were dissolved in 400–600 μl of lysis buffer III (1 mM EDTA (pH 8.0), 0.5 mM EGTA (pH 8.0), 10 mM Tris–HCl (pH 8.0), 0.5% sarkosyl and 1× protease inhibitor) before being sheared into 200–500 bp fragments with a sonicator (Bioruptor; Diagenode). Dynabeads Protein G (Invitrogen) was pre-blocked with 5 mg ml−1 of ice-cold bovine serum albumin (BSA) and incubated with 5 μg anti-H3K27ac antibody (Abcam) at 4 °C with constant rotation for 12 h. Chromatin (25 μg) was added to the bead–antibody complex mixture per reaction and incubated with constant rotation for 16 h at 4 °C. Beads were washed with ice-cold radioimmunoprecipitation assay (RIPA) buffer (Thermo Fisher Scientific) eight times, rinsed with 1× Tris–EDTA (1× Tris–EDTA) solution, eluted by adding 200 μl of ChIP elution buffer (1% SDS; 1× Tris–EDTA) and incubated in a shaker for 20 min at 65 °C. ChIP DNA was incubated for 12 h at 65 °C for reverse crosslinking, treated with RNAse A (Thermo Fisher Scientific; 1 h; 37 °C) and Proteinase K (Sigma-Aldrich; 2 h; 55 °C) and then purified on PCR purification columns. For input control, 5 μg of chromatin from whole-cell extract of each sample was subjected to reverse crosslinking, RNase A treatment (Thermo Fisher Scientific; EN0531) and Proteinase K treatment (Sigma-Aldrich; 3115887001), together with immunoprecipitated samples, and purified using PCR purification columns. DNA amounts were quantified using the PicoGreen assay (Thermo Fisher Scientific; P7589). Immunoprecipitated DNA (5 ng) and input from each sample were used to prepare ChIP libraries with TruSeq ChIP Library Preparation Kit (Illumina; IP-202-1012), according to the manufacturer’s instructions. Libraries were size selected to enrich 300–400 bp size fragments, quality controlled and sequenced on Hiseq 2000 platform (Illumina) (YCGA). Approximately 20–25 million reads were obtained from each sample. FASTA files were mapped to the mouse genome (NCBI37/mm10) using TopHat v.1.0.13 (http://tophat.cbcb.umd.edu/) and Bowtie 2 (http://bowtie-bio.sourceforge.net/bowtie2/index.shtml)62. Peaks were identified using MACS2 (SCR_013291)63.
H3K27ac ChIP–seq in the chicken
Embryonic day 15 or Hamilton–Hamburger stage 41 (ref. 64) chicken eggs were obtained from Charles River Laboratories and incubated in the laboratory for 2 days at 37 °C in a humidified chamber. The dorsal pallial regions (hyperpallium apicale and mesopallium) were microdissected from the embryonic day 17 (Hamilton–Hamburger stage 43) chicken embryos. Ten pallia were pooled and crosslinked with a formaldehyde solution (Sigma-Aldrich) at a final concentration of 1% for 10 min at room temperature per sample. L-Glycine (AmericanBio; 56-40-6) at a final concentration of 125 mM was added and incubated for 5 min at room temperature to quench the crosslinking. Cells were washed with PBS thrice and lysed in a hypotonic solution (50 mM Tris–HCl (pH 7.5), 0.5% NP-40, 0.25% sodium deoxycholate, 0.1% SDS and 150 mM NaCl) on ice for 10 min to obtain the nuclei. The nuclei were centrifuged at 600g for 5 min at 4 °C, and pellets were resuspended in the SDS lysis buffer (1% SDS, 10 mM EDTA and 50 mM Tris–HCl (pH 8.1)) before being sheared into 200–500 bp size fragments using a sonicator (M220 Focused-ultrasonicator; Covaris). The sheared DNA was diluted with the ChIP dilution buffer (0.01% SDS, 1.1% Triton X-100, 1.2 mM EDTA, 16.7 mM Tris–HCl (pH 8.1) and 167 mM NaCl) and pre-cleared with Magnetic Protein A/G Beads (MilliporeSigma) for 1 h at 4 °C. The beads were discarded, and anti-H3K27ac antibody (Abcam; ab4729) was added. The samples were incubated on constant rotation for overnight at 4 °C. Magnetic Protein A/G Beads (MilliporeSigma) blocked with 1 mg ml−1 BSA (Sigma-Aldrich) and transfer RNA were added to the chromatin–antibody complexes for 4 h at 4 °C. The beads were washed with low salt (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris–HCl (pH 8.1) and 150 mM NaCl), high salt (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris–HCl (pH 8.1) and 500 mM NaCl), LiCl (0.25 M LiCl, 1% IGEPAL CA-630, 1% deoxycholic acid (sodium salt), 1 mM EDTA and 10 mM Tris–HCl (pH 8.1)) and 1× Tris–EDTA (AmericanBio; AB14033-01000), sequentially for 3 min each. ChIP DNA was incubated overnight at 65 °C for reverse crosslinking and subjected to RNase A (Thermo Fisher Scientific; EN0531) treatment (37 °C; 1 h) and Proteinase K (Sigma-Aldrich; 3115887001) treatment (55 °C; 2 h) and then purified on PCR purification columns. For input control, 5 μg of crosslinked chromatin from each sample was also treated for reverse crosslinking, RNase A (Thermo Fisher Scientific; EN0531) and Proteinase K (Sigma-Aldrich; 3115887001), together with immunoprecipitated samples and purified by PCR purification columns. DNA amounts were quantified using PicoGreen assay (Thermo Fisher Scientific; P7589). Immunoprecipitated DNA (10 ng) and input from each sample were used to prepare libraries using the KAPA HyperPrep Kit (Roche; KK8500), according to the manufacturer’s instructions. Sample multiplexing was done using combinatorial dual indexes (Illumina) (YCGA). Libraries were size selected to enrich 300–400 bp size fragments and quality controlled and sequenced on a HiSeq 2000 platform (Illumina). Approximately 60 million reads were obtained per sample. Reads were mapped to the chicken genome (Galgal6) using Bowtie 2 v.2.4.2 (Research Resource Identifier (RRID): SCR_016368; http://bowtie-bio.sourceforge.net/bowtie2/index.shtml). Duplicates were removed, and unique reads with mapping quality > 20 were selected using SAMtools v.1.16 (RRID: SCR_003030; https://github.com/samtools/samtools). Peaks were called using MACS2 v.2.2.7.1 (RRID: SCR_008036).
ZBTB18 ChIP–seq in the mouse
Epitope-tagged ChIP was performed to identify ZBTB18 binding sites. Cortices from PD 0 were isolated and subjected to enzymatic dissociation. Cells from 15 cortices were pooled and treated as one sample. Cells were washed and transfected with HA-tagged Zbtb18 using Amaxa Basic Nucleofector Kit (Lonza Bioscience; VPI-1003) for primary mammalian neurons following the manufacturer’s instructions. After 48 h of culture, the medium was removed, and the cells were crosslinked with a formaldehyde solution at a final concentration of 1% for 10 min at room temperature. L-Glycine (AmericanBio; 56-40-6) was added at a final concentration of 125 mM and incubated for 5 min at room temperature to quench the crosslinking. The cells were scraped and collected in a 50-ml conical tube and lysed in the hypotonic solution (50 mM Tris–HCl (pH 7.5), 0.5% NP-40, 0.25% sodium deoxycholate, 0.1% SDS and 150 mM NaCl) on ice for 10 min to obtain the nuclei. The nuclei were centrifuged at 600g for 5 min at 4 °C, and pellets were resuspended in the SDS lysis buffer (1% SDS, 10 mM EDTA and 50 mM Tris–HCl (pH 8.1)) before being sheared into 200–500 bp size fragments using a sonicator (M220 Focused-ultrasonicator; Covaris). The sheared DNA was diluted with the ChIP dilution buffer (0.01% SDS, 1.1% Triton X-100, 1.2 mM EDTA, 16.7 mM Tris–HCl (pH 8.1) and 167 mM NaCl) and pre-cleared with Protein A/G Magnetic BEads (Thermo Fisher Scientific) for 1 h at 4 °C. For epitope-tagged ChIP, 5 μg of HA antibody (MilliporeSigma; 11867423001) was used. Samples were incubated on constant rotation for overnight at 4 °C. Magnetic Protein A/G Beads (MilliporeSigma; 88803) were blocked with 1 mg ml−1 BSA (Sigma-Aldrich; A2153), and transfer RNAs were added to the chromatin–antibody complexes for 4 h at 4 °C. The beads were washed with low salt (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris–HCl (pH 8.1) and 150 mM NaCl), high salt (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris–HCl (pH 8.1) and 500 mM NaCl), LiCl (0.25 M LiCl, 1% IGEPAL CA-630, 1% deoxycholic acid (sodium salt), 1 mM EDTA and 10 mM Tris–HCl (pH 8.1)) and 1× Tris–EDTA (AmericanBio) sequentially for 3 min each. ChIP DNA was incubated overnight at 65 °C for reverse crosslinking, treated with RNase A (37 °C for 1 h) and Proteinase K (55 °C for 2 h) and then purified using PCR purification columns. For input control, 5 μg of crosslinked chromatin from each sample was also treated by reverse crosslinking, RNase A (Thermo Fisher Scientific) and Proteinase K (MilliporeSigma), together with immunoprecipitated samples, and purified using PCR purification columns. DNA amounts were quantified by PicoGreen assay (Thermo Fisher Scientific; P7589). Immunoprecipitated DNA (10 ng) and input from each sample were used to prepare libraries using KAPA HyperPrep Kit (Roche; KK8500) following the manufacturer’s instructions and multiplexing using combinatorial dual indexes (Illumina). The libraries were size-selected to enrich 300–400 bp size fragments, quality controlled and sequenced on the HiSeq 2000 platform (Illumina) (YCGA). Approximately 60 million reads were obtained per sample. The reads were mapped to the mouse genome (mm10) using Bowtie 2 v.2.4.2. Duplicates were removed, and unique reads with mapping quality > 20 were selected using SAMtools (v.1.16). We explored different options to determine putative ZBTB18 peaks in terms of replicates and parameters, including using TRANSFAC65, PROMO66, MatInspector67 and JASPAR databases68. Hence, peaks were called using four different approaches: (1) MACS2 (ref. 63) with default parameters, requiring the peak to be present in at least two biological replicates; (2) MACS2 with the ‘–nomodel’ and ‘–nolambda’ parameters, requiring consistency across replicates; (3) MACS2 with default parameters but only requiring the peak to be present in replicate 2; and (4) MACS3 with the peaks of replicate 2.
TF binding site prediction of candidate CREs
We obtained the DNA sequence for all 62,448 H3K27ac peaks using the twoBitToFa69 tool (https://genome.ucsc.edu/goldenPath/help/twoBit.html). We then ran the tool Find Individual Motif Occurrences (https://meme-suite.org/meme/doc/fimo.html) with default parameters to predict TFBSs in those sequences70. We used the JASPAR 2016 CORE dataset71 in MEME format containing 635 motifs. For the ET neuron-specific TFBS enrichment analysis, we calculated the number of bases present in the binding sites for a particular TF but not in the binding sites within IT neuron-associated peaks and compared them with the same numbers in ET neuron-associated peaks through Fisher’s exact test. Motifs with an FDR-corrected P value < 0.05 were considered significantly enriched.
For the six key CREs (Cux2-E1, Cux2-E4, Satb2-E1, Robo1-E1, Bcl11b-P1 and Zbtb18-P1), we further used the regular expression (Regex) method with the ZBTB18 consensus motif sequence 5′-CAGATGT-3′, allowing 1-nt mismatch to scan for any non-canonical or low-affinity motifs. Next, we used PROMO66 to test whether there are more sites in all six CREs. Finally, we reanalysed public ZBTB18 ChIP–seq data from HEK293 cells72 to derive an empirical motif using MEME–ChIP, obtaining the top motif significantly enriched in centrality through CentriMo, and searched for the top motif (MEME–TOP) in all six CREs.
Generation of enhancer reporter transgenic mice
Putative enhancers (Cux2-E1, Cux2-E2 and Cux2-E3) were amplified from mouse genomic DNA, cloned into pBgn–Gfp vector (Sestan laboratory) and sequence verified by Sanger sequencing (YCGA). The enhancers were placed upstream of the human BGN minimal promoter73 to drive GFP expression. The primers and oligonucleotides used for cloning DNA constructs are shown in Supplementary Table 15. Enhancer-inserted plasmids were linearized using appropriate restriction enzymes, size-selected by gel electrophoresis and then purified by phenol/chloroform extraction. A final concentration of 2.5 ng μl−1 of DNA was used for pronuclear injection. At PD 0, pups were examined for GFP expression under a fluorescence microscope, and tail samples were collected for genotyping to confirm the presence of Gfp loci. For GFP+ founders, brains were collected and proceeded with immunohistological analysis to examine the GFP expression pattern. Three to seven founders from each transgenic line with stable GFP expression patterns were acquired and analysed.
Quantitative RT–PCR
Total RNA was extracted using TRIzol reagent (Invitrogen; 15596018) from freshly isolated neocortical tissue from PCD 15.5 and PD 0 wild-type, Neurod6–Cre; Zbtb18 fl/fl cKO and Neurod6–Cre; Zbtb18fl/+ (control) mice and subjected to DNase I (Invitrogen; AM1907) treatment as previously described. Complementary DNA (cDNA) was synthesized using reverse transcriptase (Invitrogen; 18080093) following the manufacturer’s instructions, and quantitative PCR was performed in triplicate for each sample using an RT–PCR machine (iQ5 system; Bio-Rad) with primer sets spanning exon junctions of the targeted transcripts. Identical or near-identical-sized transcript fragments from the mouse Tbp gene were used as an internal control, and the expression level of each gene was normalized to wild type for relative fold changes. Sequences of primers used are provided in the Supplementary Table 15. Unpaired two-tailed t-test was used to detect differences between samples.
Single-cell RNA-seq analysis across species
To assess the expression patterns of Zbtb18 across different species, we reanalysed public single-nucleus transcriptome datasets for amygdala in turtle (Trachemys scripta elegans), lizard (Pogona vitticeps), chicken (Gallus gallus) and mouse (Mus musculus). We checked the expression level of Cux2, Satb2, Robo1, Bcl11b, Ccdc1167, Rpl38-ps1, Arhgef16, n-R5s136, 3830408C21Rik, Gm24089 and Gm42489 across major cell types, as identified by the original studies12,19,20,21. For cross-species comparison, we included only the excitatory neurons.
Postmortem human tissue
De-identified postmortem human brain tissue was acquired from the Sestan laboratory collection or the NIH NeuroBioBank (https://neurobiobank.nih.gov), as previously published30. All tissue was collected with informed consent from parents or next of kin and with approval of all relevant review boards or committees of the Yale University School of Medicine and the NIH. Similarly, tissue was handled in accordance with the appropriate constraints, regulations and ethical guidelines for the research use of human brain tissue set forth by the NIH (http://bioethics.od.nih.gov/humantissue.html) and the World Medical Association Declaration of Helsinki (https://www.wma.net/policies-post/wma-declaration-of-helsinki-ethical-principles-for-medical-research-involving-human-subjects/). The samples used in this study were analysed without regard to gender, and the influence of sex-specific characteristics of male versus female samples was not considered in this study.
The samples were fixed in 4% paraformaldehyde (PFA) (Electron Microscopy Sciences) for 2 days at 4 °C. Tissue sections were mounted and dried overnight. Antigen retrieval was performed using R-Buffer A pH 6.0 (Electron Microscopy Sciences) and the bench-top Antigen Retriever device (Electron Microscopy Sciences) according to the manufacturer’s instructions. Sections were washed three times in PBS, each for 15 min, and incubated in blocking solution containing 5% (v/v) normal donkey serum (Jackson ImmunoResearch Laboratories), 1% (w/v) BSA and 0.4% (v/v) Triton X-100 (Sigma-Aldrich) in PBS for 1 h at room temperature. Primary antibodies were diluted in blocking solution as follows: ZBTB18 (Proteintech; 12714-1-AP), 1:1,000; BCL11B (Abcam; ab18465), 1:500; and SATB2 (Genway Biotech; GWB-9F2D9F), 1:200. Tissue sections were incubated with these antibodies for two nights at 4 °C. Sections were washed three times with PBS, each for 15 min, before being incubated with the appropriate fluorescent secondary antibodies (Jackson ImmunoResearch Laboratories) for 1.5 h at room temperature. All secondary antibodies were raised in donkey and diluted at 1:250 in blocking solution. Finally, sections were washed with PBS with 0.3% Triton X-100, treated with the Autofluorescence Eliminator Reagent (MilliporeSigma; 2160) according to the manufacturer’s instructions and coverslipped with aqueous mounting medium (VECTASHIELD; Vector Laboratories). These sections were digitized using confocal microscope (ZEISS; LSM 510 Meta), and images were assembled in ZEISS ZEN, ImageJ, Adobe Photoshop and Adobe Illustrator.
Cell line and transfection
Neuro-2a cells (American Type Culture Collection; CCL-131) were maintained in Dulbecco’s modified Eagle’s medium (Gibco) supplied with 10% fetal bovine serum (Gibco), L-Glutamine (Thermo Fisher Scientific), penicillin (Gibco) and streptomycin (Gibco). The cell line was authenticated by morphology or genotyping, and no commonly misidentified lines were used. All lines tested negative for mycoplasma contamination, checked monthly using the MycoAlert Mycoplasma Detection Kit (Lonza). The cells were passaged every 2–3 days upon reaching 80% confluency. For transfection experiments, the Neuro-2a cells were seeded at an appropriate density the day before transfection; when the cells reached 60–70% confluency, transfection was performed 20–24 h later, with Lipofectamine 2000 (Thermo Fisher Scientific; 11668027) mixed with appropriate vectors at a DNA:lipid ratio of 1:3. Either 24 h or 48 h after transfection, the cells were dissociated with 0.25% trypsin (Gibco), washed with PBS and processed for corresponding assays.
In utero electroporation
IUE was performed on PCD 14.5 and PCD 15.5 timed-pregnancy embryos (n = 3–6 for each condition), and littermates were used as controls. A 0.5-μl DNA preparation (4 μg μl−1 of DNA mixed with 0.05% Fast Green FCF Dye (Sigma-Aldrich) was injected into the lateral ventricle of the embryos and electroporated using a square-wave pulse electroporator (Harvard Apparatus; BTX) at 35–38 V, five pulses, 50 ms ON and 950 ms OFF to deliver DNA constructs to the ventricular zone. At PD 0, pups were screened for GFP or RFP expression under a fluorescence-attached dissection microscope. Pups with the fluorescence signal were euthanized, and brains were analysed as previously described.
Neuronal tracing and imaging
Neonatal mice injections were performed with a motorized microinjector (RWD Life Science; MM-500 and R480). In brief, mice at PD 3 were anaesthetized through hypothermia, and approximately 50 nl of retrograde AAV viral particles carrying pCAG–Gfp cassette (Addgene; 37825-AAVrg; titre ≥ 7 × 1012 vg ml−1) was injected into the medial prefrontal cortex visually with the guide of a stereo binocular microscope. At PD 7, the injected mice that were properly targeted were perfused with ice-cold PBS, followed by 4% PFA (Electron Microscopy Sciences). The collected brains underwent postfixation at 4 °C overnight and washed with PBS before cutting coronally at 60-μm thickness. The sections were stained with anti-GFP antibody (Abcam; ab13970) before imaging. Images were taken using an automated slide scanner (Olympus; VS200) with ×20 objective. For analysis, the images from wild-type mice were aligned with the Allen Brain Atlas coordinates at PD 6, and the Neurod6–Cre Zbtb18 cKO was approximated with the wild-type counterpart. The cell numbers from each bin were counted using the ImageJ tool (RRID: SCR_003070). The total cell number from the neocortex, as shown in Fig. 3c from three mice of each genotype, was compared. For the percentage of distribution, we divided the cell number from each bin by the total number of cells in the neocortex, and comparisons between each bin were made with an unpaired t-test.
BrdU/CIdU/IdU birth-date labelling
Either 5′-bromo-2′-deoxyuridine (BrdU) (Sigma-Aldrich; B5002) or CIdU (Sigma-Aldrich; C6891) was dissolved in distilled water at 100 mg ml−1, and IdU (Sigma-Aldrich; I7125) was dissolved in distilled water at 50 mg ml−1 and stored at −20 °C in the dark. At the proscribed ages, a single dose of IdU, or CIdU was redissolved by intermittent vortexing and given to the timed-pregnant mice by intraperitoneal injection at a dosage of 1 mg (20 g)−1 body weight (10 μl of CIdU or 20 μl of IdU). Pups were euthanized within the PD 1, and brains were fixed in 4% PFA (Electron Microscopy Sciences) for 12 h at 4 °C. Heterozygous littermates were used as controls. The brains were embedded with 4% agarose and sectioned at 50-μm thickness on a vibratome (Leica; VT1000S). Sections were treated with 2 M HCl at room temperature for 30 min and co-immunostained with BrdU (Sigma-Aldrich) and other markers for immunofluorescence.
Recombinant DNA
DNA constructs used for making transgenic mice were subcloned from the pBgn–GFP (Sestan laboratory) backbone with Cux2-E1, Cux2-E2 and Cux2-E3 inserted. Vectors for overexpression experiments were subcloned from the pCAGIG backbone (Addgene; 11159), with inserts of the full coding sequences of the following genes: mouse Zbtb18 (also known as Rp58, Znf238 and Zfp238), BC054529; mouse Pou3f2, NM_008899; mouse Trim28 (also known as Rnf96), BC058391; mouse Lhx2, BC055741; mouse Satb2, BC138626; mouse Hdac2, BC138517; mouse E2F1, BC052160; mouse Sin3A, BC052716; mouse Foxn2, NM_001355743; mouse Coup-tf1 (also known as Nr2f1), BC108408; and mouse Robo1, NM_019413.2. Coding sequences in DNA constructs used for IUE experiments were cloned into an expression construct with a Neurod1 promoter74 and verified by Sanger sequencing (YCGA). Other DNA constructs used in this study included pCAG–Cre, pCALNL–GFP, pCAGIG, pCALSL–RFP, pNeurod1–GFP and subcloned plasmids. The plasmids pCAG–Cre, pCALNL–GFP and pCAGIG were gifts from C. Cepko (Addgene; 11159, 13770 and 13775, respectively), whereas pCALSL–RFP and pNeurod1–GFP have been previously described74,75. Vectors carrying enhancers for luciferase assay were subcloned from pGL4.24 backbone (Promega). For enhancer mutagenesis, desired sites within specific enhancers were mutated using a site-directed mutagenesis kit (New England Biolabs; Q5 Site-Directed Mutagenesis Kit) following the manufacturer’s instructions. The mutated products were validated by Sanger sequencing. Oligos and primers used for cloning are listed in Supplementary Table 15. For neurite outgrowth analysis in chicken primary neurons, pCagen (empty vector; a gift from C. Cepko; Addgene; 11160), pCag–mCherry (a gift from Phil Sharp laboratory, through Addgene; 41583) and pCag–Cux2 (mouse Cux2 expression construct, cloned in pCAG backbone using NsiI and NotI restriction enzymes, with Cux2 open reading frame subcloned from MGC clone ID 30532644) were used.
Cux2-E1 enhancer activity assay and Cux2 overexpression in chicken hyperpallial neurons
Primary hyperpallial cells were isolated from chick embryos at embryonic days 7 and 17. After carefully opening the eggshells, the embryos were collected and immediately transferred into ice-cold artificial cerebrospinal fluid supplemented with the following components: 92 mM N-methyl-D-glucamine (Sigma-Aldrich; M2004), 20 mM HEPES (Sigma-Aldrich; H3375), 5.5 mM glucose (Sigma-Aldrich; G7021), 30 mM sodium bicarbonate (Sigma-Aldrich; S5761), 5 mM sodium L-ascorbate (Sigma-Aldrich; A4034), 2.5 mM potassium chloride (Sigma-Aldrich; P9541), 1.25 mM sodium phosphate (Sigma-Aldrich; S0751), 2 mM thiourea (Sigma-Aldrich; T7875), 3 mM 196 sodium pyruvate (Sigma-Aldrich; P2256), 5.5 mM urea (Sigma-Aldrich; U5128), 10 mM magnesium sulfate (Sigma-Aldrich; M7506) and 0.5 mM calcium chloride (Sigma-Aldrich; 21115). Following dissection, tissues were enzymatically dissociated by incubation in 2 mg ml−1 of papain (Transnetyx; PAP) for 20–30 min at 37 °C. After enzymatic digestion, 0.1 mg ml−1 of DNase I (STEMCELL Technologies; 07900) was added, and the tissue was gently triturated using a 1-ml glass pipette to obtain a single-cell suspension. Dissociated cells were transfected with various plasmids using the Chicken Neuron Nucleofector Kit (Lonza; VPG-1002) according to the manufacturer’s instructions. A total of 4 µg of each plasmid was used in all experiments. Post-transfection cells were seeded at a density of 4 × 105 cells per well in 24-well plates (ibidi; 82406) precoated with poly-D-lysine (Gibco; A3890401) and laminin (Gibco; 23017015). Cultures were maintained in neurobasal medium (Gibco; 21103049) supplemented with 1× B27 (Gibco; 17504044), 1% GlutaMAX (Gibco; 35050061), 30 nM sodium selenite (Sigma-Aldrich; S5261), 5 µg ml−1 of human insulin (Sigma-Aldrich; I9278), 1% penicillin–streptomycin (Gibco; 15140122), 10 ng ml−1 of brain-derived neurotrophic factor (PeproTech; 450-02) and 10 ng ml−1 of NT-3 (PeproTech; 450-03). Media were changed every 2 days. After 48 h, primary chicken neuronal cultures were fixed with 4% PFA for 10 min, followed by three washes with PBS. Cells were permeabilized with 0.3% Triton X-100 (Sigma-Aldrich; T8787) in PBS for 10 min at room temperature and blocked with 10% (v/v) normal donkey serum (Jackson ImmunoResearch Laboratoires; 017-000-121) in PBS for 1 h at room temperature. The primary antibodies were diluted in the same blocking solution and incubated with the cells overnight at 4 °C. The primary antibodies used were anti-SATB2 (1:100; Abcam), anti-ZBTB18 (1:250; Proteintech), anti-GFP (1:2,000; Abcam) and anti-mCherry (1:1,000; Aves Labs). After three washes with PBS, cells were incubated with the respective secondary antibodies prepared in blocking solution for 2 h at room temperature. Nuclei were stained with DAPI (1 μg ml−1; Sigma-Aldrich; D9542) for 5 min at room temperature. Fluorescent images were acquired using a confocal microscope (LSM880; Zeiss). Post hoc, the GFP intensity was quantified using the ImageJ tool (RRID: SCR_003070), and the neurite length was traced following mCherry signal using the NeuronJ plugin of the ImageJ tool. Because deletion constructs used as controls may alter the local DNA context and possibly disrupt cryptic overlapping motifs, we further performed in silico analysis of the 30-bp DNA sequencing flanking the deleted region and found no high-affinity binding motifs for other relevant TFs.
Luciferase assay
Neuro-2a (American Type Culture Collection; CCL-131) cells were plated into 96-well or 24-well plates at a density of 10,000 or 50,000 cells per well, respectively. Sixteen hours after plating, cells were transfected with a DNA mixture consisting of 100 ng and 500 ng overexpression plasmids, 30 ng and 500 ng of firefly luciferase (pGL4.24; Promega) plasmids containing candidate enhancers, together with 20 ng and 25 ng pRL-SV40 plasmid (Promega) as a control for 96-well and 24-well plates, respectively, using Lipofectamine 2000 (Thermo Fisher Scientific). Transfected cells were lysed and assayed 24–48 h after transfection using the Dual-Luciferase Reporter Assay System (Promega) according to the manufacturer’s instructions. Relative luciferase activity was calculated by normalizing the firefly luciferase luminescence to the Renilla luciferase luminescence value. Luciferase activity was measured and quantified by GloMax-Multi Detection System (Promega). Primers used for amplifying the candidate enhancer and promoter regions are shown in Supplementary Table 15. Experiments were performed in triplicate or quadruplicate, and the data were normalized to the empty vector control (pCAGEN versus the pZBTB18). P values were calculated using unpaired two-tailed t-tests to compare the overexpression and control. To perform the luciferase assay on multiple mutated enhancers, a one-way ANOVA test was used to detect differences among more than two enhancers; alpha value = 0.05.
In situ hybridization for the human, mouse and chicken brain sections
RNA probes for mouse and human were generated using the respective neocortical tissue cDNA as template (mouse Zbtb18, ENSMUSG00000063659; mouse Cux2, ENSMUSG00000042589; mouse Satb2, ENSMUSG00000038331; mouse Robo1, ENSMUSG00000022883; mouse Robo2, ENSMUSG00000052516; mouse Robo3, ENSMUSG00000032128; mouse Rorb, ENSMUSG00000036192; mouse Dcc, ENSMUSG00000060534; human ZBTB18, ENSG00000179456; and human CUX2, ENSG00000111249) by TA Cloning Kit (Invitrogen) and followed by in vitro transcription (Roche) according to the manufacturer’s instructions. For chicken probes, the cDNA fragments coding for SATB2 (NM_001199110) and CUX2 (XM_415167), were amplified by RT–PCR and subcloned into the plasmid vector pTA2 (Toyobo). Templates purified by phenol/chloroform extraction and digoxigenin (DIG)-labelled probes were synthetized using T3 (Roche) and T7 RNA polymerases (Roche), respectively. RNA labelling was performed mix (Roche) according to the manufacturer’s instructions. Probes were purified by phenol/chloroform extraction, quantified, quality controlled and stored at −80 °C until hybridization. Primers used for generating probes are provided in Supplementary Table 15.
For single-colour ISH, free-floating or slide-mounted cryosections at 20–30 μm thickness were processed according to a previously described protocol76. In brief, brains were fixed overnight at 4 °C in 4% PFA (Electron Microscopy Sciences) diluted in Dulbecco’s phosphate-buffered saline (DPBS) (Thermo Fisher Scientific), equilibrated for 12 h at 4 °C in 10% sucrose and another 12 h at 4 °C in 30% sucrose (AmericanBio) in DPBS (Thermo Fisher Scientific). Fixed brains were then embedded in O.C.T. (Scigen) and sliced on a cryostat (Leica Biosystems; CM1800). Slides were stored at −80 °C until processed for ISH. Sections were first postfixed in 4% PFA (Electron Microscopy Sciences) in PBS for 15 min at room temperature, washed with PBS and submerged in hybridization buffer (5× saline–sodium citrate, 50% formamide, 5× Denhardt’s solution, 500 µg ml−1 of salmon sperm DNA and 250 µg ml−1 of torula yeast RNA) supplemented with 1,000 ng ml−1 of the appropriate DIG-labelled probe at 70 °C overnight. Sections were washed twice for 60 min at 70 °C in 2× saline–sodium citrate, 50% formamide and 0.1% Tween, followed by washing with 100 mM Tris–HCl (pH 7.5), 150 mM NaCl and 0.1% Tween. These sections were then blocked with 10% sheep inactivated serum (Sigma-Aldrich) and incubated overnight at 4 °C with an anti-DIG antibody conjugated to alkaline phosphatase (1:2,000; Roche). The sections were then rinsed in the substrate buffer (100 mM Tris–HCl (pH 9.5), 100 mM NaCl, 50 mM MgCl2 and 0.1% Tween-20) before being overlaid with NBT–BCIP substrate (Roche). Revelation was done at room temperature in the dark until the desired signal was reached. Finally, the sections were rinsed in DPBS (Thermo Fisher Scientific), postfixed with 4% PFA (Electron Microscopy Sciences) in DPBS (Thermo Fisher Scientific), washed with water and mounted with Permount mounting medium (Electron Microscopy Sciences).
For two-colour ISH, probes were synthesized by either DIG-labelled (Roche) or fluorescein-labelled (Roche) RNA labelling mixes in vitro transcription (Roche). Free-floating or slide-mounted cryosections (20–30 μm thickness) were postfixed in 4% PFA (Electron Microscopy Sciences) for 15 min, washed and hybridized overnight at 70 °C in 14-ml hybridization solution with 500 ng ml−1 of DIG-labelled (Roche) human ZBTB18 probe and 500 ng ml−1 of fluorescein-labelled (Roche) human CUX2 probe for human tissue. For mouse tissue, 500 ng ml−1 of DIG-labelled mouse Zbtb18 and 500 ng μl−1 of fluorescein-labelled mouse Cux2 probes were used. The signals were sequentially detected with an alkaline phosphatase-conjugated anti-DIG antibody (MilliporeSigma), NBT–BCIP substrate (Roche) and horseradish peroxidase-conjugated anti-fluorescein antibody. Horseradish peroxidase activity was initially amplified using TSA (Akoya) and subsequently strengthened with 3,3′-diaminobenzidine tetrahydrochloride hydrate (DAB) (Sigma-Aldrich).
Immunostaining and immunoblotting
Brains dissected from embryonic and neonatal mice were fixed by immersion in 4% PFA (Electron Microscopy Sciences) overnight at 4 °C. Adult brains were perfused with 10 ml PBS followed by 10 ml of 4% PFA (Electron Microscopy Sciences), isolated and postfixed by immersion in 4% PFA (Electron Microscopy Sciences) overnight at 4 °C. The brains were then sectioned at a thickness of 50 μm (postnatal) and 80 μm (embryonic) on a vibratome (Leica; VT1000S). The brain sections were blocked using blocking solution (5% BSA, 10% donkey serum and 0.3% Triton X-100 in 1× PBS solution) for 1 h at room temperature and incubated with appropriate primary antibodies for 12–24 h at 4 °C, washed three times with 1× PBS and incubated with appropriate secondary antibodies for 1 h at room temperature. DAPI was used to stain nuclei. The antibody dilutions used for immunostaining were as follows: anti-ZBTB18 (rabbit; 1:1,000; Proteintech), anti-GFP (chicken; 1:3,000; Abcam), anti-RFP (rabbit; 1:1,000; Abcam), anti-CUX1 (rabbit; 1:250; Santa Cruz Biotechnology), anti-CUX2 (rabbit; 1:250; Abcam), anti-SATB2 (mouse; 1:200; Genway), anti-BCL11B (rat; 1:500; Abcam), anti-TBR1 (rabbit; 1:250; Santa Cruz Biotechnology), anti-LHX6 (mouse; 1:300; Santa Cruz Biotechnology), anti-LHX6 (goat; 1:250; Santa Cruz Biotechnology), anti-RELN (mouse; 1:300; Millipore), anti-BrDU (mouse; 1:250; BD Biosciences) and anti-BrDU (rat; 1:250; Accurate Chemical). For the quantitative analysis of total number of BCL11B+ and SATB2+ cells, z-stack images from three per species were used to segment DAPI signals from individual nuclei, and fluorescence signals from the nuclei were obtained using Volocity (v.6.3.1). The positive signals for each marker protein were defined on the basis of the median nuclear intensity from all cells analysed using the Spotfire software (v.11.2.0).
Immunoblot analyses were performed using a previously described protocol77. In brief, neocortical tissue from PD 0 Zbtb18fl/fl; Emx1–Cre mice (n = 3), Zbtb18fl/+; Emx1–Cre mice (n = 3) and wild-type mice (n = 3) were isolated and snap-frozen in liquid nitrogen. The samples were then minced and mixed with lysis buffer (150 mM NaCl, 1.0% NP-40 and 50 mM Tris–HCl (pH 8.0)), including protease inhibitors (Roche). Equal amounts of protein from each sample were loaded for SDS–PAGE, followed by western blot analysis with signals detected using the enhanced chemiluminescence detection reagent. Antibody dilutions used for immunoblot were as follows: anti-GAPDH (rabbit; 1:5,000; Abcam; ab9485), anti-CUX2 (rabbit; 1:1,000; Abcam), anti-ZBTB18 (rabbit; 1:2,000; Proteintech) and anti-ZBTB18 (goat; 1:2,000; Santa Cruz Biotechnology). For detailed antibody information, refer to Supplementary Table 14.
Obtaining multispecies alignments of TF binding sites
Beginning with the lists of putative TFBSs identified by Find Individual Motif Occurrences, as described earlier70, we selected only those motifs that are recognized by TFs expressed (reads per kilobase of transcript per million mapped reads ≥ 1) in our RNA-seq samples derived from either IT or ET neurons for further analysis. This produced a list of 267 motifs. The total number of binding sites for those motifs ranged from 0 to 244,332.
For each of the 267 motifs, we obtained the multiple-species sequence alignment of 60 vertebrate species from the MULTIZ60 (ref. 78) track in the University of California, Santa Cruz (UCSC) browser using mafsInRegion69 from the UCSC tools. We stitched maf alignments and converted them to fasta using the script maf_to_concat_fasta.py from bx-python (https://github.com/bxlab/bx-python/blob/main/scripts/maf_to_concat_fasta.py).
Conservation of TF binding sites across species
We used an in-house script to analyse the multiple sequence alignment of each binding site. First, we calculated the information content for each position in each MEME motif using the R package TFBSTools79. For each motif, we identified positions with high information content (0.75 or higher) and considered them ‘core’ positions. Second, we divided the species in the 60-way multiple sequence alignment into four groups: ‘placental mammals’, ‘marsupials’, ‘monotremes’ and ‘non-mammals’. Finally, for each binding site, we counted the number of species in each group in which the motif was present without any single nucleotide substitutions in any of the core positions compared with the mouse reference sequence mm10. We repeated the analysis with one, two and three mutations. A motif was not considered conserved if it contained an indel spanning the core positions.
For each group of species, we produced a binary vector in which, for each binding site, a value of 1 was assigned if the binding site was conserved and 0 if it was non-conserved. For placental mammals, we required at least six species fulfilling the conservation criteria stated above. For marsupials, monotremes and non-mammalian species, we required at least one species.
Enrichment in TF binding sites in sets of H3K27ac peaks
We selected seven sets of ChIP–seq peaks: peaks annotated to differentially expressed genes between IT and ET neurons (DESeq2 PAdj < 0.05; one set for upregulated genes and another for downregulated genes), peaks differentially enriched between IT and ET neurons (as determined by DESeq2 with FC > 2 and PAdj < 0.1; one set for IT neuron-biased peaks and another for ET neuron-biased peaks), peaks annotated to genes differentially expressed between the Zbtb18 knockout (KO) or Zbtb18 heterozygous mice (DESeq2 PAdj < 0.05; one set for KO and another for heterozygous-biased genes) and a final set composed by a manually curated list of genes important for corpus callosum formation extracted from the literature. In addition, we produced a list of background peaks composed of all H3K27ac peaks, which did not fall in any of the previous categories. We then grouped these CREs into four categories to evaluate the enrichment of TFBSs across species: (1) H3K27ac peaks associated with ET neuron-biased genes (n = 1,433), which include peaks annotated to ET neuron-enriched genes and peaks enriched in ET neurons; (2) peaks associated with IT neuron-biased genes (n = 1,409), which include peaks annotated to IT neuron-enriched genes and peaks enriched in IT neurons; (3) peaks associated with CREs near genes with expression that was downregulated in Zbtb18 KO (n = 8,187); and (4) peaks associated with CREs near genes with expression that was upregulated in Zbtb18 KO (n = 3,705). For associating peaks with genes in the identification of putative CREs, we associated peaks with the nearest genes.
For each group of species, we counted the number of conserved and non-conserved binding sites observed in each set of peaks. We compared the proportion of conserved versus non-conserved binding sites to the same ratio in the background peaks using Fisher’s exact test. Motifs with FDR-adjusted P values < 0.05 were considered significant.
The number of motifs tested in each category of enhancers, including those selected as background supplementary, is shown in Supplementary Table 17. To evaluate the robustness of these enrichments, we devised a bootstrap-based strategy (Extended Data Fig. 10d). For each comparison, we counted ZBTB18 motifs within each category and performed 1,000 random samplings, with replacement, of an equal number of ZBTB18 motifs from background H3K27ac regions (considered invariant putative regulatory elements). We then compared the observed ratios of conserved and non-conserved motifs at each phylogenetic depth to the distribution of 1,000 such ratios obtained from the background peaks. The percentile occupied by the observed ratio represents an empirical P value, indicating how likely it is to obtain this ratio by chance among the invariant peaks.
Conservation of H3K27ac and ZBTB18 peaks
We used mafsInRegion69 from the UCSC tools to obtain MULTIZ60 alignments for each selected enhancer. We stitched maf alignments and converted them to fasta using the script maf_to_concat_fasta.py from bx-python. The pairwise alignment distance between species of each of the selected H3K27ac peaks was obtained using the function dist.alignment from the seqinr package in R.
The intersection of chicken and mouse H3K27ac and ZBTB18 ChIP–seq peaks was conducted using the ‘IntersectBed’ function within bedtools. To facilitate this intersection, chicken peaks were initially converted into mouse coordinates (from gg6 to mm10) using LiftOver with -minMatch = 0.1. Orthologous regions in chicken of all ZBTB18 ChIP–seq peaks identified in mice were determined using LiftOver from mm10 to gg6 using the same minMatch value.
Evolutionary age of the ZBTB18 gene
To evaluate the evolutionary age of the ZBTB18 gene, we used protein sequence similarity (BLASTP80 and HMMER81) to search our reference database, which is adapted from UniProt82 and trimmed with a taxonomically informed procedure that optimizes database size while ensuring that species with well-resolved genomes are kept in all major branches of the phylogenetic tree of life83. We partitioned the similarity hits according to their evolutionary distance from humans and the species in which they appear. Here the evolutionary age of a gene is defined as the taxonomic restriction of the most ancient fragment of the protein encoded by the gene83,84,85,86. If an ancestral gene duplicated and expanded, all descendant genes are considered of the same evolutionary gene as the ancestral gene86. For any protein of at least 40 amino acids, the taxonomic restriction is indicated as the index of the NCBI taxonomic node87 at which the query species and subject species diverged. The evolutionary age of each taxonomic (phylogenetic) node is provided by the TimeTree database88. Thus, a human protein also present in bacteria is ancient; a human protein also present in sea squirts is restricted to the Chordata phylum; a human protein also present in platypus is restricted to the Mammalia class; a human protein present also in the Coquerel’s sifaka lemur is restricted to the Primates order; a human protein present only in humans is restricted to the human species. First, we found that few bacterial species in our database harbour proteins similar to the human ZBTB18 protein (four species with hits in 1,600 species of our database; 0.025%). Of the four bacterial proteins similar to ZBTB18, three of them have two or four zinc finger domains, whereas the remaining protein has a BTB domain. Most eukaryotic species harbour proteins similar to the human ZBTB18 (67 of 100 eukaryotic species; 67%). The three eukaryotic proteins most similar to ZBTB18 have two or four zinc finger domains. Second, we found that in all subsequent evolutionary nodes of the human lineage, all outgroup species for each node harbour proteins similar to ZBTB18. Thus, ZBTB18 is an ancient gene that probably arose at the origin of eukaryotes. The few bacterial hits may indicate an even more ancient origin in all cellular organisms. The presence of genes similar to ZBTB18 in only 0.025% of bacterial species is consistent either with an extremely ancient origin in all cellular organisms followed by many events of gene loss in most bacterial species, or with two to four events of horizontal gene transfer from eukaryotes to a few bacterial species. Although both explanations are possible, the second requires fewer events. In any case, ZBTB18 is an ancient gene that arose in eukaryotes approximately two billion years ago or even earlier at the origin of cellular organisms approximately 3.6 billion years ago.
Cell counting and data analysis
For each cell counting analysis, neocortical somatosensory regions from three different brains of each condition were used. For PD 0 mouse and late to midfetal human brain tissues, 200-μm-wide neocortical columns covering from layer 1 to subplate were used as standard fields to conduct counting. For PCD 14.5 and PCD 16.5 mouse tissues, 200-μm-wide neocortical columns covering from layer 1 to ventricular zone were used as standard fields. Unpaired two-tailed t-tests were used to detect differences between samples. P < 0.05 (α = 0.05) was set as the cutoff for significance.
Replicates
For ChIP–seq, two replicates were used for each condition. For RNA-seq, a minimum of three replicates were used for each condition. For ChIP–quantitative polymerase chain reaction analysis, a minimum of four replicates were used for each condition. For RT–PCR analysis, three replicates from each time point were used for each condition. For Cux2–E1, Cux2–E2 and Cux2–E3 transgenic mice, three to seven founders from each line were analysed. For IUE experiments, three to six successfully electroporated animals were examined for each analysis. For IDU/CIDU in vivo labelling, a minimum of two litters were used for each experiment, and three to five brains from each litter were examined. For ISH and immunofluorescence experiments using mouse tissue, a minimum of three animals were used for each experiment, and between 3 and 20 sections from each animal were examined. For ISH and immunofluorescence experiments using non-mouse mammals, two to five sections from two individuals were used. For cell culture experiments, cells were isolated from three different mouse brains per condition, and two replicates from each brain were used for subsequent observations. For luciferase assays, a minimum of three replicates were used for each transfection and analysis. Statistical methods were not used to determine sample size. For all experiments, n = 3 or >3 was used.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.