Ethics
The research consists of secondary analyses of de-identified primary datasets. Information regarding informed consent of participants (or guardians) in primary studies can be found in the references in Supplementary Table S2.T2. Secondary analysis of these data was also approved by the Vanderbilt University Medical School Institutional Review Board (IRB no. 210968).
Software reporting
All analysis visualizations and statistics were generated using Python v3.9.20. Notable Python libraries used are pandas (v2.2.3), numpy (v2.0.2), seaborn (v0.13.2), matplotlib (v3.9.2), rpy2 (v3.5.11) and scipy (v1.13.1). For creating 3D qualitative visualizations of WM tracts, we used the mrview tool inside the MRtrix3 (version 3.0.4). Alternatively, for qualitative region of interest 3D visualizations, the OpenDIVE visualization library (v0.3.0) was used with a different Python (v3.10.18) for library compatibility purposes. Regarding the image preprocessing and processing software, please refer to ‘MRI processing pipeline’.
Data
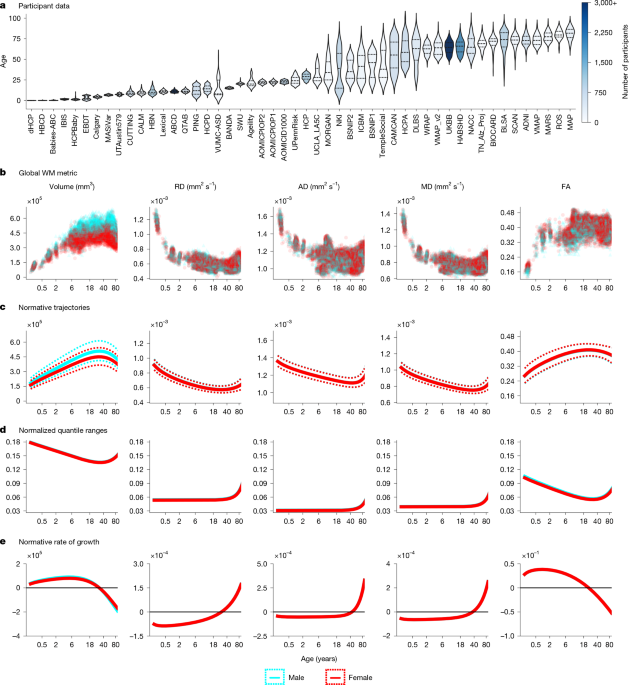
We aggregated diffusion-weighted imaging (DWI) and T1-weighted (T1w) data from 50 independent studies spanning 0 to 100 years of age (Extended Data Table 1), encompassing 75,036 DWI scans from cognitively normal and clinical participants. All data were converted from DICOM to NIfTI using dcm2niix and organized in BIDS format in accordance with previous work44. Notably, the 50 included datasets were harmonized through standardized preprocessing, postprocessing, and feature-wise statistical harmonization using GAMLSS.
MRI processing pipeline
DWI data were preprocessed using the PreQual pipeline45, which corrects for susceptibility-induced, motion-related and eddy current distortions, and performs slice-wise signal imputation. Specifically, image denoising was performed using the MRtrix3 toolkit46 (v3.0.4) implementation of Marchenko–Pastur principal component analysis. Following this, TOPUP from the FSL (v6.0.4) software library47 is used for susceptibility-distortion correction. For DWI without reverse phase encoding scans acquired, TOPUP is run using a synthetic b0 image created from a T1w scan from the same imaging session via the Synb0-DISCO48 algorithm (v3.1). FSL’s EDDY is then used for motion and eddy current-distortion correction, also performing slice-wise signal imputation with the flag ‘–repol‘. Following preprocessing, DTI models were fit to all volumes with b-values ≤ 1,500 s mm−2 (refs. 40,49) using dwi2tensor from MRtrix3. DTI-derived scalar maps—including FA, MD, AD and RD—were computed using tensor2metric from MRtrix346.
To enable consistent tract segmentation, all diffusion data were resampled to 1 mm isotropic resolution using MRtrix350. Tractography was performed using TractSeg51 (v2.8), which automatically segments 72 anatomically defined cerebral WM pathways (see Extended Data Table 2 for a list of tracts and abbreviations). For each tract, we computed streamline density-weighted averages of DTI metrics (FA, MD, AD and RD) as well as macrostructural features—volume, length and surface area—using the Scilpy toolkit52 (v1.5.0) scripts scil_compute_bundle_mean_std.py and scil_evaluate_bundles_individual_measures.py, respectively. Both raw and ratio-normalized macrostructural features are provided, where volume, surface area and average length of tracts are normalized to total brain volume (excluding ventricles), estimated total intracranial volume and cerebral WM volume.
T1w images were included only when acquired in the same session as DWI data. Brain segmentation was performed using the recon-all command from FreeSurfer (v7.2.0)53, yielding estimates of cerebral WM volume, brain volume excluding ventricles and total intracranial volume. For participants aged ≤2 years, we employed infant_recon_all from the infant FreeSurfer pipeline54 (v0.0) to account for age-specific brain morphology. Cerebral WM masks were defined using MRtrix346 5TT labels, excluding the cerebellum and brainstem. T1w images and corresponding WM segmentations were rigidly registered to DWI space using FSL’s47 epi_reg. Whole-brain WM DTI metrics were then computed by averaging values within the WM mask.
The PreQual preprocessing pipeline can be downloaded from https://zenodo.org/records/14058394. We also make the entire postprocessing pipeline available at https://zenodo.org/records/17144460.
Data selection
Following the approach of Bethlehem et al.6, we restricted our analysis to cross-sectional data. Participant scans were excluded if key demographic information—age, sex or cognitive status—was missing or not reported for that participant. For each dataset, sex was used as reported and encoded as a binary variable, whereas age was a continuous value in years. Regarding infants in the BABIES-ABC, HBCD and dHCP datasets, we used corrected age based on the due date to better account for prematurity and provide a more accurate developmental context.
Quality control (QC) procedures followed our previously established framework for large-scale dMRI analysis55. QC metrics were visually reviewed across PreQual preprocessing outputs, FreeSurfer segmentations, and T1w–DWI registration results. Scans were excluded if any preprocessing step failed or was deemed unusable. TractSeg outputs were excluded if more than 12 of the 72 tracts failed to reconstruct. Outlier removal was performed separately for each tract-level feature within each dataset, with observations excluded if exceeding ±4 standard deviations from the mean. We note it is possible for FreeSurfer to have failed with successful tractography, and vice-versa, in which cases we retained only data from the contrast that passed quality assessment. The number of sessions and participants retained after QC is detailed in Extended Data Table 1.
Normative trajectories were modelled using only participants labelled as ‘typically developing’ or ‘control’ within their respective studies to reflect patterns of healthy WM development and ageing. Participants who were initially labelled as typically ageing but later transitioned to a clinical diagnosis were also excluded when fitting normative trajectories. At the tract level, features were retained only if the tract was reconstructed with the full target of 2,000 streamlines—the default setting in TractSeg.
Normative modelling of features
We utilized GAMLSS15, a flexible regression framework endorsed by the World Health Organization for constructing normative growth curves12,16. GAMLSS extends generalized linear and additive models by allowing the simultaneous estimation of multiple distribution parameters—not only the mean, but also variance, skewness and kurtosis—through functions that vary with age and other covariates. This flexibility enables precise modelling of lifespan trajectories for WM features.
Formally, GAMLSS defines each parameter of the assumed response distribution through additive predictors:
$$g_k(\theta _k)=X_k\beta _k+\mathop\sum \limits_j=1^J_kZ_kj\gamma _kj,$$
(1)
where \(g_k(\cdot )\) is the link function for the \(k\)th parameter, \(X_k\) and \(\beta _k\) are the design matrix and fixed effects. The summation incorporates \(J_k\) smooth functions, where \(Z_kj\) are the design matrices for the basis expansions and \(\gamma _kj\) are the corresponding smoothing coefficients. This formulation allows flexible, non-linear modelling of the entire distribution as a function of covariates such as age, sex and dataset.
Previous work has shown dMRI-derived features to have skewed distributions. Following Bethlehem et al.6, we used the generalized gamma (GG) distribution with fractional polynomial (\(\textfp\)) fitting to estimate lifespan trajectories. The GG distribution offers substantial flexibility, accommodating a broad range of distributional shapes, and is therefore suitable for modelling both microstructural and macrostructural imaging features. The model parameters, location (μ), scale (σ) and shape (ν), were defined as:
$$g_1(\mu )=x_\mathrmsex\beta _1,\mathrmsex+\rmfp_1(x_\mathrmage)+\gamma _1,D$$
(2)
$$g_2(\sigma )=x_\mathrmsex\beta _2,\mathrmsex+\rmfp_2(x_\mathrmage)+\gamma _2,D$$
(3)
$$g_3(\nu )=\alpha ,$$
(4)
where \(g_1(\cdot )\) and \(g_2(\cdot )\) denote log link functions, whereas \(g_3(\cdot )\) is an identity link. Age was modelled as a continuous predictor using fractional polynomial transformations \(\textfp_k(\cdot )\), enabling non-linear representation of age-related effects. Sex was included as a binary fixed effect (\(x_\mathrmsex\)) and dataset-specific variability was modelled via random intercepts (\(\gamma _1,D\) and \(\gamma _2,D\)). The GG distribution was parameterized as described in Rigby et al.56 to enable compatibility with GAMLSS framework. Model selection was performed by the Bayesian information criterion57 across all combinations of 1–3 fractional polynomial terms for the \(\mu \) and \(\sigma \) parameters. The shape parameter \(\nu \) was treated as a global offset without age- or dataset-specific effects, consistent with Bethlehem et al.6.
We modelled normative trajectories for 509 features, including 288 tract-level (72 tracts × 4 features) microstructural measures (mean FA, MD, AD and RD) and 216 macrostructural (72 tracts × 3 features) measures (volume, average length and surface area). An additional five global WM features—mean FA, MD, AD, RD and WM volume—were also modelled. Macrostructural features were further normalized using total intracranial volume, brain volume excluding ventricles, and global WM volume to account for anatomical scaling.
To derive scaling factors for normalization, we assumed a spherical brain geometry and estimated radius and surface area using:
$$r_\mathrmBrain=\left(\frac3V_\mathrmBrain4\rm\pi \right)^\frac13$$
(5)
and
$$\mathrmSA_\mathrmBrain=4\rm\pi r_\mathrmBrain$$
(6)
respectively, where \(V_\mathrmBrain\) denotes the normalizing volume, with separate scaling factors derived for estimated total intracranial volume, brain volume excluding ventricles, and global WM volume. VBrain, SABrain and rBrain (Equations 5 and 6) were used to normalize tract-level volume, surface area and average length metrics, respectively.
When considering both normalized and unnormalized features, we had 1,157 features in total for which we created lifespan curves: 509 unnormalized (288 microstructure, 216 macrostructure and 5 global) and 648 normalized (216 macrostructure normalized in 3 separate ways) features.
WM lifespan milestones
For assessing myelination, we studied the median RD trajectories of tract-specific brain charts. Following the work from Yeatman et al.17, we performed piecewise segmentations of the trajectories to assess whether the myelination time points align with proposed periods of myelin development and maturation derived in refs. 18,25,26. Specifically, we performed a piecewise segmentation of the median trajectories to identify three break points that minimize the sum of squared errors across the resulting linear segments (where the piecewise model was fit between ages 1 and 90 years). The segments correspond to myelination periods of rapid development in infancy, slower myelination in adolescence, the stable plateau of young adulthood, and the decline of ageing. For each segment, we calculated both the average slope and the per cent change per year (Supplementary Table SDT.T5).
Centile scores across cognitive groups
We used the fitted normative trajectories and dataset-specific random effects to compute individualized centile scores for non-control participants across all tract-level features. In total, 6,777 participants from 16 clinical groups passed QC and were included in the analysis (Extended Data Table 3).
As the analysis was restricted to cross-sectional data, we selected a single scan per participant. For individuals with longitudinal data, we retained the most recent scan corresponding to their most severe clinical diagnosis. For example, participants progressing from cognitively unimpaired to MCI and then to Alzheimer’s disease were classified based on their most recent scan with a clinical Alzheimer’s disease or dementia diagnosis.
For each clinical group, we compared the distribution of centile scores across tract metrics to the 50th percentile expected in the normative population. In addition, we quantified overall deviation from the normative distribution using the nCMD for each participant \(j\) (\(\mathrmnCMD_j\)):
$$\mathrmnCMD_j=\frac\sqrt(x-\mu )^TS^-1(x-\mu )N_j_\mathrmtot$$
(7)
where \(N_j_\mathrmtot\) is the number of reconstructed tract features for participant \(j\), \(x\) is a vector of centile scores (ranging from \(0\) to \(1\)), \(\mu =0.5\) denotes the median centile, and \(S^-1\) is the inverse of the covariance matrix for the features.
We computed nCMD separately for tract-level microstructural features, macrostructural features, and their combination. Group-level comparisons were then made by evaluating the median nCMD in each non-control group relative to the distribution observed in the cognitively normal population, where statistical significance was tested using a one-sample Wilcoxon test (two-sided) after Bonferroni correction for multiple comparisons.
Alignment for OOS datasets
A central utility of normative brain charts is their use as reference models for external, OOS datasets. To enable this, new datasets must be aligned to the fitted trajectories by estimating study-specific offsets for the distributional parameters. Within the GAMLSS framework, each dataset \(D\) is modelled with random effect terms \(\gamma _1,D\) and \(\gamma _2,D\), which account for dataset-specific variabilities in the location (\(\mu \)) and scale (\(\sigma \)) parameters, respectively. For an unseen dataset \(S\), alignment involves estimating these two unknown offsets. We demonstrate this process using the ADNI dataset, where we fit additional models without ADNI for the purpose of demonstrating OOS alignment.
For a given brain chart modelling a tract-level metric, the GAMLSS-based lifespan distribution is defined as:
$$g_1(\mu )=\mathrmfp_1(x_\mathrmage)+x_\mathrmsex\beta _1,\mathrmsex+\gamma _1,S$$
(8)
$$g_2(\sigma )=\mathrmfp_2(x_\mathrmage)+x_\mathrmsex\beta _2,\mathrmsex+\gamma _2,S$$
(9)
$$g_3(\nu )=\alpha ,$$
(10)
where \(\gamma _1,S\) and \(\gamma _2,S\) are dataset-specific random effects for the ADNI dataset, and all other model parameters are fixed from the trained GAMLSS model fitted without ADNI data.
As all other effects are known from the reference model, alignment requires estimating only the unknown random effects \(\gamma _1,S\) and \(\gamma _2,S\). We initialized both to zero and restricted the estimation procedure to typically ageing individuals in the ADNI cohort. To minimize diagnostic confounding, we selected only participants with no history or future diagnosis of MCI or Alzheimer’s disease and used their earliest available scan. This cognitively normal subset is denoted \(S_\mathrmCN\).
For each cognitively normal participant \(j\) in \(S_\mathrmCN\), we computed the likelihood of their observed metric \(M_j\) under the GG distribution defined by equations (8)–(10). Each of the resulting probability densities \(P_j=GG(M_j|\mu ,\sigma ,\nu )\) were used to calculate the overall negative log-likelihood across all participants in \(S_\mathrmCN\):
$$\rmNegative\,\log \text-\rmlikelihood=-\mathop\sum \limits_j=1^S_\rmCN\log (P_j),$$
(11)
We iteratively re-estimated \(\gamma _1,S\) and \(\gamma _2,S\) by minimizing the negative log-likelihood until convergence. The final estimates, \(\hat\gamma _1,S\) and \(\hat\gamma _2,S\), were then applied to all participants in the ADNI dataset to compute adjusted centile scores aligned to the normative reference space.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.