Mouse husbandry
All experiments were approved and conducted in accordance with the Cold Spring Harbor Laboratory Institutional Animal Care and Use Committee. All animals used were adult (>3 months) male and female mice. Laboratory mice were acquired from The Jackson Laboratory (C57BL/6J). Colonies of laboratory and singing mice were maintained at 20–22 °C, 30–70% humidity and a 12–12 h light–dark cycle.
Behavioural recordings and analysis
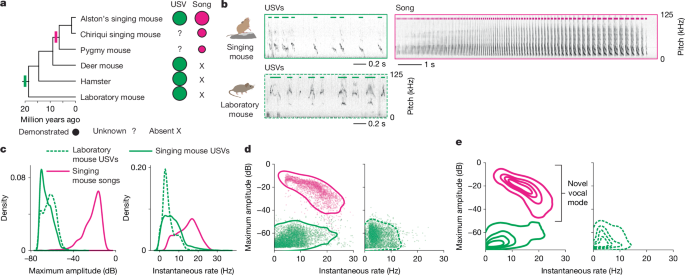
A female of each species was muted by synaptic silencing of the caudolateral PAG using tetanus-toxin light chain7,50 and allowed 2 weeks for protein expression. For five sessions per species, the mute female was placed with a conspecific male in a clean cage for interaction (Thoren Systems 8; 30.8 × 40.6 × 22.2 cm) lined with Alpha-pad cotton paper (Shepherd Specialty Papers). Audio of each dyad was recorded for 1–2 h using two Avisoft CM16/CMPA microphones positioned above the cage with high and low gain settings and sampling at 250 kHz (digitized with Avisoft UltraSoundGate 116H). Vocalizations were segmented from the audio using USVSEG69 software for MATLAB (v.09r2), and parameters were optimized to the sounds emitted by each species. For singing mice, songs and their individual notes were also detected using custom code in Python. Segmented sounds were manually curated in a customized spectrogram browser derived from the open-source MATLAB graphical user interface DeepSqueak70. Curation consisted of correcting any vocalization boundary errors and removal of abiotic false positives. Spectrogram inspection confirmed that stimulus females were mute, with no overlapping vocalizations observed; female singing mice also ceased singing during continuous post-surgery monitoring. Songs were operationally defined as sequences of loud notes with a minimum duration of 1.5 s and gaps of no greater than 0.5 s. Song notes were defined as part of the songs, whereas the remaining quiet notes were classified as USVs50. For each curated note, we calculated the maximum amplitude in a fixed frequency range (10–100 kHz). We calculated the instantaneous note rate as the inverse of the interval between two consecutive note start times. For note pitch, the minimum of the fundamental frequency of each note was calculated in Python, using USVSEG-based preprocessing to enhance vocal signals and to suppress background noise69. Kernel densities for note acoustic parameters were calculated using the kdeplot function in the Python package seaborn71, specifying either two or five levels.
Stereotaxic viral injections
Before surgery, subcutaneous meloxicam was delivered at a dose of 2 mg kg–1. Surgeries were performed on a stereotaxic apparatus under 1–2% isoflurane in oxygen. The OMC was localized on the basis of published data (laboratory mice, refs. 35,36; singing mice, ref. 32), which identifies the OMC as the area in the motor cortex that, when stimulated, results in orofacial muscle contraction (coordinates relative to bregma: anterior–posterior (AP), 2.25 mm; medial–lateral (ML), 2.25 mm). These coordinates were used for all OMC injections unless otherwise noted. To mute female stimulus mice for behavioural recordings, we bilaterally expressed tetanus-toxin light chain7,50 in the caudolateral PAG using AAVs. A female singing mouse and a laboratory mouse were bilaterally injected in the caudolateral PAG with 80 nl of a 1:1 mixture of AAV2/DJ-hSyn-flex-TeLC-eYFP (custom packaged by WZ Biosciences) and CamKII-Cre. The following coordinates were used: singing mouse, AP, –4.2 mm; ML: ±0.6 mm; dorsal–ventral (DV), –2.3 mm (ref. 50); laboratory mouse, AP, −4.7 mm; ML, ±0.7 mm; DV, −1.75 (ref. 7).
Histology of these mice were also used in anterograde trans-synaptic tracing of the OMC to the PAG.
For mapping of bulk neuronal projections, neurons were targeted to express tdTomato using a 1:1 mixture of CaMKII-Cre and FLEX-tdTomato viruses (Supplementary Table 1). Next, 30 nl of this viral mixture was injected into the OMC at two depths: 500 and 750 µm ventral to the brain surface.
For MAPseq viral injections, Sindbis virus carrying the barcode library72 was diluted 1:3 in sterile saline. The OMC was injected with 50 nl of diluted Sindbis virus at 300, 600 and 900 µm ventral to the brain surface at two locations: AP, 2.0 and 2.5 mm; and ML, 2.25 mm.
For visualizing synaptic boutons, a 1:1 mixture of CaMKII-Cre and Flex-mGFP-2A-Synaptophysin-mRuby (Supplementary Table 1) was injected into the OMC using the coordinates listed above. Specifically, 60 nl of this viral mixture was injected at two depths: 500 and 750 µm ventral to the brain surface.
For anterograde trans-synaptic tracing73 of the OMC to the AudR, a 1:1 mixture of CaMKII-Cre and DIO-mCherry (Supplementary Table 1) was injected into the OMC of singing mice. Specifically, 80 nl was injected at depths of 500 and 750 µm ventral to the brain surface. DIO-eGFP virus (Supplementary Table 1) was injected into the AudR (AP, −1.2; ML, 4.25 mm). Three injections of 50 nl at 2-min intervals were delivered at a single depth of 2.3 mm ventral to the brain surface.
For retrograde tracing of the OMC, retrograde AAV carrying either GFP or GFP-NLS (Supplementary Table 1) was injected into the OMC of laboratory or singing mice. Next, 30 nl of virus was injected at two depths: 500 and 750 µm ventral to the brain surface.
Mice were transcardially perfused with 4 °C PBS and then with 4 °C 4% paraformaldehyde (PFA) 14 days after injection for OMC projection mapping, 44 h after injection for MAPseq experiment or 21 days after injection for all other viral tracing experiments.
Confocal imaging
After transcardial perfusion, brains were fixed in 4% PFA at 4 °C for 24 h. Brains were rinsed with PBS then embedded in 3% agarose. Brains were sectioned on a vibratome at 50 µm, stained with DAPI and mounted on glass slides with ProLong Gold Antifade mountant. Microscopy images were acquired in Airyscan SR-4Y multiplexing mode using a Zeiss LSM 980 Airyscan2 inverted confocal laser scanning microscope (Carl Zeiss Microscopy), equipped with a Plan-Apochromat ×10/0.45 NA M27 air-objective lens. DAPI was excited using a 405 nm diode laser, mCherry and tdTomato fluorophores were excited with a 561 nm diode laser, and GFP and YFP fluorophores were excited with a 488 nm diode laser. Emission signals for fluorophore signals were collected using a 32-channel gallium arsenide phosphide (GaAsP-PMT) Airyscan detector, operated in frame scanning mode with a pixel dwell time of 0.51 µs. Airyscan raw images were processed in Zeiss ZEN Blue (v.3.8.2) software using Fast Airyscan SheppardSum SR-4Y processing with an automated 2D Weiner filter (standard strength).
STPT imaging and image processing
STPT imaging was conducted using established protocols74,75. First, brains were embedded in 4% oxidized agarose followed by crosslinking in 0.2% sodium borohydrate solution. To image tdTomato projections from the OMC in the whole brain, entire coronal planes of the embedded brains were imaged every 50 µm on a TissueCyte 1000 STPT microscope (Tissuevision) by tiling and slicing the brain. A chameleon ultra (coherent) 150 fs pulsed laser at 930 nm, Olympus objective (×20, NA = 1.0) and 602/70 emission filter were used to image at 1 µm per pixel lateral resolution. Image correction and stitching was completed using custom-built software74,75.
STPT data analysis
Alignment of image stacks to Allen CCF
Twenty-fold scaled down images were used for comprehensive and easy visualization between brains. Brains from a representative laboratory mouse and a singing mouse were chosen as a reference to align brains in the species. The Allen Brain atlas (2022 CCFv3)76 was also registered to each brain using the brainreg function with default parameters from the brainglobe suite (https://brainglobe.info/index.html)77. Annotated areas from the aligned Allen brain atlas were used to determine volumes of major brain regions. To determine the injection site of each animal, a threshold was determined using Otsu’s method. The intersection of the combined annotated primary and secondary motor cortical regions and the thresholded data were used to create a mask of the injection site for each brain. Injection sites of aligned brains were maximum projected in the DV axis to determine overlap between brains.
Quantification of fluorescent axonal signal
High-resolution images were visualized in Napari (v.0.5.6)77. Target regions of interest (OMCc, AudR, PAG and SNr) were then manually outlined using the Napari plugin brainglobe-segmentation78. Digital dissections were guided by anatomical landmarks to match MAPseq dissections as closely as possible. The injection site (OMCi) of each animal was outlined based on the presence of fluorescent cell bodies. Outlines were used to create a mask and applied to each relevant brain region. Images were then imported into LABKIT79, which was used to train a classifier to segment axonal fluorescent signals from background (Extended Data Fig. 6a,b). The classifier was trained on a representative image from every region in every mouse. After applying the trained classifier to all images, the segmented volume of each region was calculated and normalized to the injection site volume in each animal (Extended Data Fig. 6d). The fold change was calculated by taking the average normalized volume of each region among laboratory or singing mice (Extended Data Fig. 6e).
MAPseq tissue processing
After transcardial perfusion, brains were fixed in 4% PFA at 4 °C for 24 h. The brains were then rinsed with PBS and transferred to 300 mM glycine for 24 h at 4 °C. After incubation, brains were embedded in cryoembedding medium and frozen for sectioning. Brains were cut coronally at 200–250 µm sections for microdissection. Injection sites were identified through bright-field imaging of GFP. Target regions were chosen to include all OMC projection targets based on our STPT data, and regional boundaries were identified through visual landmarks determined from the Allen brain atlas as well as our STPT data (Supplementary Figs. 1 and 2). During dissection, samples were kept on wet ice during dissection. Throughout the procedure, tools and blades were changed between dissection targets and gloves were changed between samples to prevent area and sample cross-contamination. Samples were collected from the olfactory bulb and hippocampus as negative controls. Barcode RNA was extracted, reverse-transcribed and amplified using a published protocol43,80. An Illumina sequencing library was generated and sequenced on an Illumina NextSeq500 sequencer. Possible artefacts, including the effect of fibres of passage, viral toxicity, co-infections, double use of a single barcode sequence and various other sources of false negatives and false positives, have been previously characterized43,44. Notably, although MAPseq, like GFP tracing, does not distinguish fibres of passage, their contribution is minimized by avoiding large fibre bundles during the dissection of target areas. Also, by specifically injecting the barcodes into the OMC and subsequently only sequencing the barcode RNA, we only obtained signals from the targeted OMC neurons. Therefore, our results will not be affected by other non-OMC-projecting neurons to the target regions.
MAPseq data analysis
After sequencing, reads were demultiplexed, and the absolute counts of each barcode were determined on the basis of the unique molecular identifier sequence and error-corrected barcode sequences matched to the sequenced virus library. A matrix of size ((number of barcodes) × (number of dissected areas)) was constructed, whereby matrix entries corresponded to the absolute counts of individual barcodes in each area. Only barcodes with at least 30 barcode molecules at the injection site and 5 barcode molecules at the maximum target site were used. A barcode was determined to project to a target site if there were at least four barcode molecules at that target site. This threshold minimized barcode detection in negative control target regions. We did not find significant differences in OMC projections to the anterior or posterior PAG in either species (Wilcoxon signed-rank test, P > 0.05). Therefore, we decided to pool all barcodes to the PAG.
Heatmaps and cell-type labelling
Normalized barcode counts were calculated from barcode counts normalized to a RNA spike-in of known concentration and quantity in each sample. For heatmaps, 1,000 unique barcodes (that is, neurons) were randomly sampled from an individual laboratory or singing mouse, and the normalized barcode counts were plotted (Fig. 3b) and used to compare Sindbis infectivity between species (Extended Data Fig. 4a,b). Neurons were classified into three major projection, excitatory cell types: IT, CT and PT. PT cells were first defined as neurons having any projection to a midbrain or hindbrain region (HY, PG, PAG, SNr, SCm or HB). CT cells were defined as having no projections to midbrain or hindbrain regions, but having a projection to the thalamus. IT cells were defined as cells having no projections to the midbrain, hindbrain or thalamic regions, and only projections to the striatum and/or cortex.
Modelling extent of innervation
To eliminate animal-to-animal batch effects, we calculated a normalized barcode expression value. For each animal, we took the normalized barcode counts and divided these counts by the mean normalized barcode count at the injection site (OMCi). The normalized barcode expression was taken as a proxy for the amount of axonal material in each dissected area. The empirical cumulative distribution function of the normalized barcode expression for all neurons in a region was plotted to compare species differences. The median value of normalized expression in each region for an individual animal was compared between species, and a two-sided Mann–Whitney U-test was performed to determine species differences.
Modelling projection probabilities
To calculate the proportion of neurons projecting to different target regions, the data were binarized (threshold of four barcode counts). Proportions were calculated for each major cell type. This approach accounts for any potential variance in the infectivity across cortical layers, which could bias the relative proportion of major cell types in the dataset. For example, the proportion of OMC neurons projecting to the AudR regions were the number of unique barcodes that had above the threshold signal (at least four barcode counts) in the AudR region divided by the total number of unique barcodes classified as IT cells. AudR, OMCc and STR proportions were calculated over the total IT population. HY, PG, PAG, SNr, SCm and HB proportions were calculated over the PT population. A two-sided Mann–Whitney U-test was used to calculate significant differences between individuals of each species on a per area basis.
Degree and motif analyses
The node degree for a given neuron was defined as the number of projecting areas each neuron has. For instance, if a neuron has projections in the AudR and STR, this neuron would have a node degree of 2. For degree and motif analyses, only IT cells were analysed as there were many fewer PT cells per degree and motif. Motif proportions were calculated for IT cells of an individual mouse. The motif proportions were calculated on the basis of an estimation of the total IT cell population as previously derived44. In brief, the total IT cell population was adjusted to account for unobserved neurons with projections to uncollected regions or without projections. The total IT cell population can be estimated on the basis of finding the roots of a polynomial derived from observed numbers of neurons to collected regions. The total IT population was calculated for each individual mouse and used as the denominator for determining the observed proportion for each motif. The expected proportion for each motif was calculated by multiplying the probabilities of projecting to individual brain regions according to a binomial model. The difference between the experimentally observed versus expected proportions for each motif was aggregated for every individual animal. P values were calculated under the binomial model for each individual singing mouse, and the Fisher’s method was used to combine P values across individuals. The extent of deviation from the binomial model was computed for each species, and their distributions were compared using an F-test for equality of two variances.
Calculating total OMC neuron population
In our MAPseq experiment, the number of IT neurons was defined as the number of neurons that project exclusively to IT regions; that is, neurons that project to the AudR, OMCc and/or STR. However, the observed number of IT neurons in a MAPseq dataset is a biased sampling of OMC neurons. Owing to the nature of the technique, we did not account for neurons that could be part of the OMC IT population but project to regions outside the regions we sampled. Therefore, the number of OMC IT neurons we counted in our dataset is an underestimate of the total OMC IT population. If we wanted to make reasonable motif predictions under a binomial model, the total OMC IT population count must be adjusted. As previously described44, we can derive an estimate for the total OMC IT neurons (Ntotal) for each individual animal using our observed counts of how many neurons project to each target region. First, we observe that:
$${{Prob}}_{\mathrm{neuron\; projects\; to\; at\; least\; one\; region}}+{{Prob}}_{\mathrm{neuron\; projects\; to\; no\; regions}}=1.$$
(1)
We can also assume that the proportion of neurons that projects to a region is equivalent to the probability that a neuron projects to that region:
$$Pro{b}_{{\rm{r}}{\rm{e}}{\rm{g}}{\rm{i}}{\rm{o}}{\rm{n}}}=\frac{{N}_{{\rm{r}}{\rm{e}}{\rm{g}}{\rm{i}}{\rm{o}}{\rm{n}}}}{{N}_{{\rm{t}}{\rm{o}}{\rm{t}}{\rm{a}}{\rm{l}}}}$$
(2)
where Nregion is the number of neurons that project to that region, and Ntotal is the total number of OMC IT neurons. Considering we have three IT target regions (OMCc, AudR and STR), we can combine the above two equations to obtain the following equation:
$$\left(\frac{{N}_{\mathrm{obs}}}{{N}_{\mathrm{total}}}\right)+\left(\left(1-\frac{{N}_{\mathrm{OMCc}}}{{N}_{\mathrm{total}}}\right)\,\left(1-\frac{{N}_{\mathrm{AudR}}}{{N}_{\mathrm{total}}}\right)\left(1-\frac{{N}_{\mathrm{STR}}}{{N}_{\mathrm{total}}}\right)\right)=1$$
(3)
where Nobs is the number of unique barcodes (neurons) we recovered in our MAPseq experiment, NOMCc, NSTR and NAudR are the numbers of unique barcodes found to project to each individual area (OMCc, STR or AudR), and Ntotal is the total number of IT neurons that we are trying to account for. Expanding and rearranging the above equation, we can derive the following quadratic equation:
$$\begin{array}{l}({N}_{\mathrm{obs}}-{N}_{\mathrm{OMCc}}-{N}_{\mathrm{AudR}}-{N}_{\mathrm{STR}}){N}_{\mathrm{total}}^{2}+({N}_{\mathrm{OMCc}}{N}_{\mathrm{AudR}}\\ \,+{N}_{\mathrm{OMCc}}{N}_{\mathrm{STR}}+{N}_{\mathrm{AudR}}{N}_{\mathrm{STR}}){N}_{\mathrm{total}}-({N}_{\mathrm{OMCc}}{N}_{\mathrm{AudR}}{N}_{\mathrm{STR}})=0.\end{array}$$
(4)
We can then use a roots solver to solve for Ntotal. For each animal, we used the observed barcode counts for each region and calculated a quadratic equation using the above formula. We then used a roots solver to find the largest real root. We used this number as the adjusted OMC IT population size (Ntotal) and used this Ntotal to calculate the probability that a neuron projects to an individual region. We used these adjusted probabilities to estimate motif proportions expected under the binomial model to provide us with more accurate predictions.
Bootstrap resampling to match the number of neurons across species
Because more unique barcodes were recovered for laboratory mice, we verified our findings by matching the number of neurons in singing mice with a downsampled population of neurons in the laboratory mice. First, all laboratory mouse neurons of the same cell type (IT or PT) were pooled. The pooled neurons were randomly sampled, without replacement, seven times to exactly match the number of neurons recovered in each of the singing mice. Downstream target proportions and IT motif proportions were calculated for this downsampled population of laboratory mouse neurons. A two-sided Mann–Whitney U-test was used to calculate IT motif proportion differences between species.
Analysis of OMC cooling data
We analysed a previously published OMC cooling dataset32 to assess the causal contribution of the OMC on song rhythm. The instantaneous note rate was defined as the reciprocal of the time between successive note onsets. For each note, the instantaneous note rate was plotted against the time since the song start to construct song trajectories. For each animal, the difference between the instantaneous note rates for cooled and uncooled songs was calculated for all notes within each second (that is, a bin size of 1 s). Means and error bars (corresponding to s.e.m.) were calculated across all nine animals. Significance was evaluated with a two-sample t-test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.