Study sponsorship, authorizations and ethics
This study was an investigator initiated trial sponsored by Gustave Roussy and covered by a Biomedical Research Promoter Civil Liability insurance contract (contract 124.895) in accordance with the provisions of the French law (Decree 2006-477 of 26 April 2006 and Article L.1121-10 of the French Public Health Code). This study was approved by the French Health Agency (Agence Nationale de Sécurité du Médicament) on 29 March 2016 (ANSM 160104A-12) and by the national ethics committee (Comité de Protection des Personnes Ile-de-France VIII) on 16 February 2016 (CPP 160215). The study was registered on EUDRACT (2015-005429-37) on 30 November 2015 and on ClinicalTrials.gov (NCT02857569) on 19 July 2016. The study was conducted in accordance with the Declaration of Helsinki and international Conference on Harmonization Good Clinical Practice (GCP). All investigators were GCP certified. The study was approved by the Gustave Roussy institutional review board (CSET 2015/2334) on 24 November 2015. All patients provided written informed consent prior enrollment in the trial and before any study specific procedure for clinical data and ancillary analysis anonymous use. Ipilimumab and nivolumab were supplied by Bristol Myers Squibb.
Objectives
The study aimed to evaluate the 6-month treatment tolerance defined as the treatment-related grade 3–4 adverse event-free survival of the combination therapy intratumoural (IT) ipilimumab plus intravenous (IV) nivolumab. The IV ipilimumab plus IV nivolumab (same doses as in phase 1) arm will be used as an internal control to interpret the results obtained in the IT ipilimumab arm.
Secondary objectives were to explore the types of toxicities generated by the combination therapy in the two treatment arms and to evaluate the efficacy of IT ipilimumab in combination with IV nivolumab.
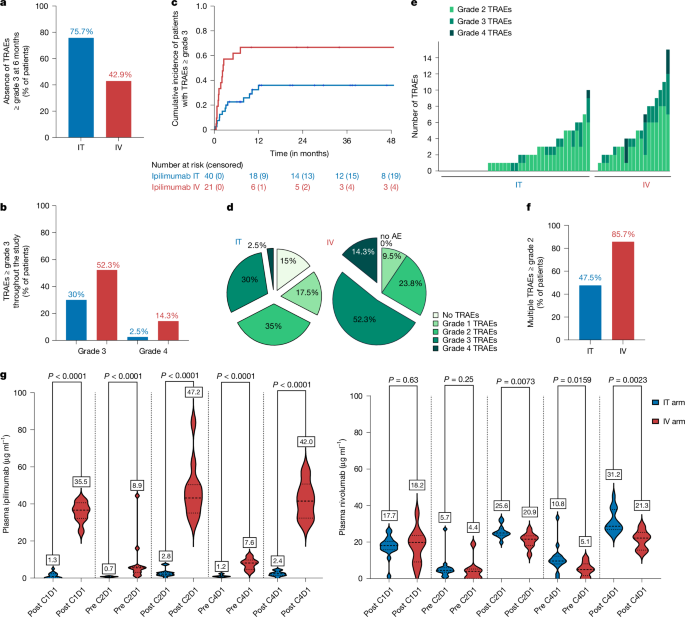
Translational objectives were to demonstrate the lower systemic exposure to ipilimumab in the IT arm and to identify predictive biomarkers of response.
To assess response to treatment using several end-points for efficacy based either on RECIST 1.1, or on immune-related response criteria.
Key eligibility criteria
Main inclusion and non-inclusion criteria are presented below (not exhaustive list, if needed cf. protocol).
Inclusion criteria
-
Men and women ≥18 years of age who signed a written informed consent before any study-related procedure.
-
Histologically confirmed and clinically or radiologically progressing unresectable stage III or stage IV melanoma, as per American Joint Committee on Cancer (AJCC) staging system.
-
Patients with at least two lesions: at least one injectable tumour lesion (≥1 cm3) and at least one target lesion (measurable lesion as per RECIST 1.1). Disease measurable by computed tomography or MRI (RECIST 1.1).
-
Good general status, Eastern Cooperative Oncology Group (ECOG) performance status of 0 or 1.
-
First line treatment.
-
Recent (less than 3 month) tumour tissue provided for patient stratification and biomarker analyses.
-
Subjects with wild-type BRAF. Those with BRAF mutations can be included only if they have been treated with, or developed toxicity with or refused to be treated with, BRAF-and/or MEK-targeted therapy.
-
Usual laboratory test limitations (refer to protocol).
-
Usual limitations concerning women of childbearing potential and sexually active men.
Non-inclusion criteria
-
Active brain metastases or leptomeningeal metastases.
-
Ocular melanoma.
-
Any serious or uncontrolled medical disorder.
-
Prior malignancy active within the previous 3 years except for locally curable cancers.
-
Subjects with active, known or suspected autoimmune disease.
-
Subjects with a condition requiring systemic treatment with either corticosteroids (>10 mg daily prednisone equivalents) or other immunosuppressive medications within 14 days of study drug administration.
-
Prior treatment with an anti-PD1, anti-PD-L1, anti-PD-L2, anti-CTLA4, or any other antibody or drug specifically targeting T cell co-stimulation or immune checkpoint pathways.
-
Positive test for hepatitis B virus surface antigen (HBV sAg) or hepatitis C virus ribonucleic acid (HCV antibody) indicating acute or chronic infection, or HIV infection.
-
Patients presenting coagulation abnormalities and/or patients requiring concomitant treatment with therapeutic doses of anticoagulants. Prophylactic low dose of anticoagulants for thrombo-embolic events is allowed. Prophylactic anticoagulants shall be stopped during 24 h before and after deep lesion biopsies/injections. No stopping required for biopsies/injections of skin and sub-cutaneous lesions.
Treatment and follow-up scheduled
Experimental IT arm
-
A.
Ipilimumab: 0.3 mg kg−1 IT injection every 3 weeks until complete response, eradication of all injectable sites, disease progression or toxicity, for a maximum of 4 doses (control: IV standard of care and marketing authorization). Pure ipilimumab from commercial vials at 5 mg ml−1 was used for IT injections. For instance, a patient weighing 75 kg would receive 75 × 0.3 = 22.5 mg—that is, 4.5 ml of pure ipilimumab into the chosen tumour lesions. All injections were performed under image guidance by interventional radiologists using 22 gauge (22G) Chiba needles with a single endhole. The injected volume ranged between 3 and 6.5 ml (at 5 mg ml−1). For large lesions, a different part of the tumour was injected at each cycle in a clockwise manner. A very low injection rate was used (1 ml min−1) to avoid reflux and optimize distribution within the tumour. Local anaesthesia with lidocaine (2%) was used for all except some superficial skin lesions. Minimal size of injected lesions was 1 cm in diameter per protocol. The volume of ipilimumab per patient could be divided in order to inject several lesions but with the objective of adjusting the injected volume to the volume of the treated lesion as much as possible.
-
B.
Nivolumab: 1 mg kg−1, IV injection every 3 weeks during IT ipilimumab treatment period and 3 mg kg−1, IV injection every 2 weeks after IT ipilimumab treatment interruption. Treatment should be continued as long as clinical benefit is observed or until treatment is no longer tolerated by the patient, for a maximum of 12 months.
Standard arm
-
A.
Ipilimumab: 3 mg kg−1, IV injection every 3 weeks for a maximum of 4 doses as per standard of care and marketing authorization.
-
B.
Nivolumab: 1 mg kg−1, IV injection every 3 weeks during IV ipilimumab treatment period and 3 mg kg−1, IV injection every 2 weeks after IV ipilimumab treatment interruption. Treatment should be continued as long as clinical benefit is observed or until treatment is no longer tolerated by the patient, for a maximum of 12 months.
Statistical methods
Safety and efficacy analysis were based on patients who took at least 1 dose of the study drug.
Primary end-point was the 6-month treatment related grade 3–4 toxicity event-free survival (EFS), defined as the time from inclusion to first documentation of treatment-related grade 3–4 toxicity. The number of patients to be included in the experimental arm was calculated considering a Fleming’s two-stage design and assuming the following, where P is the probability of success (tolerance of 6-month treatment), and α type 1 error rate of 10% and power of 90%: P0 = 50%, the 6-month treatment related grade 3–4 EFS below which the combination will considered too toxic; P1 = 70%, the 6-month treatment related grade 3–4 EFS above which the combination will be considered safe.
A total of 38 evaluable patients were required in the experimental arm. If the number of patients without treatment-related grade 3–4 adverse events after 6 months of treatment was <23/38, the conclusion of the design would be unacceptable tolerance. If the number of patients without treatment-related grade 3–4 adverse events after 6 months of treatment was ≥23/38, then the conclusion of the design would be acceptable tolerance.
No formal comparison between the two treatment arms was performed. The standard arm only serves as an internal control of the hypothesis of P0 and at least 19 patients were to be included.
Median follow-up was estimated with inverse Kaplan–Meier method. Overall survival was defined as the time from inclusion to documentation of death due to any cause. PFS was defined as the time from inclusion to documentation of tumour progression or death, whichever occurs first. PFS was be assessed by tumour measurements using RECIST 1.1. Overall survival and PFS was estimated using the Kaplan–Meier method and presented with Rothman’s 95% confidence intervals. The software used for statistical method analysis was SAS software version 9.4.
Translational research
Nivolumab and Ipilimumab serum titrations
Nivolumab dosage method consists of one-day pre-analytical and analytical steps. For the pre-analytical step: Protein A purification was performed with Thermofisher scientific Protein A Spin plates; digestion was performed with Promega trypsin added to the samples and incubated overnight in a Thermomixer to thermostatically control the temperature under 300 rpm agitation; solid phase extraction was performed on Waters OASIS 96-well plates with washing steps with an aqueous solution and elution steps with an organic solvent; and sample concentration was performed with Speedvac vacuum concentrators with a temperature and vacuum level of 80 °C and 0.1 bar. For the analytical step: a Stepwave Xevo TQ-S UPLC-Mass Spectrometer (triple quadrupole, from Waters) with Masslynx software was used, over a range of 2.5–500 mg l−1. The nivolumab surrogate peptides used were ASGITFSNSGMHWVR (ASGI) and ASQSVSSYLAWYQQKPGQAPRLLIYDASNR (ASQS) with MRM transitions of 550.8 → 661.5 and 785.0 → 1,033.8, respectively. Bevacizumab was used as an internal standard with FTFSLDTSK (FTFS; MRM transition 523.4 → 797.4) and STAYLQMNSLR (STAY; MRM transition 642.3 → 861.3) as surrogate peptides. Separation was performed using a gradient of 0–1.0 min: 2.7% B, 1.0–6.0 min: 19% B, 6.0–6.10 min: 26.1% B, 6.1–8.0 min: 81% B, 8.10–10.0 min: 2.7% B.
Ipilimumab quantification was performed using an enzyme-linked immunosorbent assay (ELISA). Anti-ipilimumab Antibody Monovalent Fab, HCA330 (Bio-Rad AbD34433), was used as the capture antibody at a final concentration of 1 µg ml−1. The anti-ipilimumab antibody HCA329P (Bio-Rad AbD34429) was used as the detection antibody at 0.1 µg ml−1. Serum samples were diluted in PBS Tween 0.05% as follow: IV arm at baseline 1/200, IV arm after treatment at 1/40, IT arm at baseline 1/20, IT arm after treatment at 1/5. Enzymatic reaction was performed using QuantaBlu Fluorogenic Peroxidase Substrate Kit (Thermo Scientific). Fluorescence signal was quantified with Spectramax microplate reader (excitation 325 nm, emission 420 nm).
Tissue and blood sample analyses
Fresh whole blood was used to phenotype circulating immune cells by flow cytometry. Plasma was used to titre cytokines, chemokines and soluble factors. Only injected lesions were biopsied at baseline in the IT arm. Both injected and uninjected tumour lesions were biopsied at week 3 in patients from the IT arm. Up to six cores of tumour biopsies were collected from tumour lesions. WES and RNA-seq were performed on frozen tumour biopsies. IHC staining was performed on formalin-fixed paraffin-embedded (FFPE) tumour biopsies. Fresh tumour biopsies were also collected into media and sent immediately to a central laboratory for processing. Fresh tumour biopsies were subsequently mechanically and enzymatically dissociated and processed for flow cytometry. Cytokines, chemokines and soluble factors were titrated on supernatants (secretome) from fresh tumour biopsies. Tumour immunophenotyping and secretome analysis was performed for 22 and 17 patients, respectively. Sequential biopsies were performed at baseline (before treatment) and prior to cycle 2—that is, week 3 (on treatment)—in order to perform paired analysis whenever possible (n = 16). For patients in the IT arm, on treatment (week 3) biopsy samples were taken both from the injected and from an uninjected tumour (n = 12). Samples were collected for pathological analysis from 49 patients.
Immunohistochemistry
IHC was performed on 4-µm-thick, deparaffinized sections of FFPE tissue samples. The primary antibodies used in the study are listed in Extended Data Table 4; they were purchased from Abcam, Dako, DBioSystems, Dendritics, LS-Bio, MBL, R&D Systems, Roche Diagnostics and Spring Biosciences. Simple labelling immunoperoxidase techniques were used for CD26, CTLA4, HLA-I and HLA-II and mast cell tryptase. Three-plex chromogenic techniques were used for the co-detection of FOXP3/CD3/ICOS, CD8/PD-L1/Ki-67 and DC-Lamp/CD68/CD163. A four-plex chromogenic technique was used for the simultaneous detection of CD3/CD20/CD68/CD57. All techniques were performed on automated stainers (Benchmark ULTRA or Discovery, both from Ventana, or Bond RX, Leica Biosystems). The interpretation was performed by two experienced pathologists. The results were expressed according to the marker of interest, by H-scores, semi-quantitative evaluations or manual counting of labelled cells for the evaluation of cell densities (number of cells per mm²).
Flow cytometry, immune cell phenotyping and analysis from tumour biopsies
Core biopsy samples from tumour at baseline and prior to cycle 2, week3, (injected and uninjected) were immediately placed into 1 ml of NaCl 0.9% and sent to the laboratory (LRTI–U1015). After a minimum of 30 min of incubation, fine-needle biopsies were mechanically dissociated with the bottom of a 2 ml syringe in a wet 70-µm filter placed at the top of a 50 ml centrifuge tube. Isolated cells were then washed by centrifugation and the pellet was re-suspended in an appropriate volume of NaCl 0.9% for the cell surface staining protocol. Antibodies mix (Extended Data Table 5) was composed of immune markers CD45, CD3, CD4, CD8 and HLA-ABC, activation markers HLA-DR and CD25, Treg cell markers CD39 and CTLA4, immune checkpoint markers PD1, OX40 and TIGIT and a co-stimulator marker CD26. CTLA4 was first stained at 37 °C for 20 min before others surface antibodies were added and incubated at 4 °C for 15 min. Then cells were washed twice and acquisition were performed on an 18-colour flow cytometer BD Fortessa X20 (BD Biosciences). Data were processed in FCS 3.0 format and analysed with KALUZA software v2.1. From our population of interest, doublets were first excluded based on forward scatter height versus forward scatter area plot, and then viable cells were selected. Tumour-infiltrating T cells were then selected with a CD45+ followed by a CD3+ gate, and then divided into two sub-populations based on CD4 and CD8 expression (Extended Data Fig. 10).
Plasma and secretome cytokines measurement
Low-platelet plasma was generated on blood tubes collected in EDTA upon double high-speed centrifugation at 4 °C. Secretome experiments were standardized following the same procedure: one single 18 G fresh core needle biopsy was put into 1 ml saline an incubated at 4 °C for at least 30 min prior collection of the supernatants. To evaluate 51 soluble factors in patient plasma and/or 10 soluble factors in supernatant of biopsies, a Quickplex SQ120 platform enabling highly sensitive electro-chemo-luminescent detection (Meso Scale Discovery) was used following the manufacturer’s instructions. The analytes measured were GM-CSF; eotaxin and eotaxin-3; interleukins (IL-1α, IL-1β, IL-2, IL-4, IL-5, IL-6, IL-7, IL-8, IL-9, IL-10, IL-12p70, IL-12/IL-23p40, IL-13, IL-15, IL-16, IL-17A, IL-18, IL-18 BP, IL-21, IL-22, IL-27, IL-33, and receptor antagonist IL-1RA, CD25); interferons (IFNγ, IFNα2a, IFNλ (also known as IL-29)); IP-10; MIP1α and MIP1β; MDC; MCP1 and MCP4; TNFα and TNFβ; TARC; VEGF; fractalkine; calprotectin; immune checkpoint soluble proteins (PD1, PD-L1 and CTLA4); enzymatic proteins (GZMA and GZMB); BCA1; and TGFβ1, TGFβ2 and TGFβ3. Absolute concentrations of soluble analytes (in pg ml−1 or fg ml−1) in patient samples were calculated by use of a four-point-fit calibration curve of the standard dilutions (MSD Discovery Workbench analysis software) and were considered detectable if both runs of each sample had a signal greater than the analyte- and plate-specific lower limit of detection.
Flow cytometry immune cells phenotyping and analysis from blood samples
Immune cell phenotyping from blood samples was performed according to a previously described methodology49. The panels used are summarized in the Extended Data Table 6.
WES and RNA analysis
The exome libraries were prepared using the Agilent V6 XT kit and sequenced on a BGISEQ-500 instrument at BGI according to the manufacturer’s protocols.
Reads were mapped to the GRCh37 human reference genome using BWA-MEM (v0.7.12) software. A standard GATK best practice pipeline was used to process the samples and call somatic genetic variants using GATK Mutect250. Variants were annotated with oncotator software (v1.9.9.0) and oncoKB database. SCNA calling was done with FACETS software (v 0.5.14). All processing steps were implemented with a snakemake pipeline (v5.4.0)51. Quality control of FASTQ and BAM files was performed with FASTQC (v0.11.7) and samtools (v1.9) respectively52. Only somatic variants with PASS flag and supported by minimum three reads with at least one read from each strand were used for the further analysis. Known mutational signatures for WES mutations were deconvoluted using R package MutationalPatterns (fit_to_signatures_strict(), max_delta = 0.003) based on the mutational matrices from SigProfilerMatrixGenerator. Oncoplot was constructed using maftools R package53,54.
RNA samples were sequenced on BGISEQ-500 instrument. The reads were pseudoaligned on the human transcriptome database (hg38) with the Kallisto pipeline and final TPM values for each gene in each sample were received as described55. TPM values were used to perform absolute deconvolution of the tumour microenviroment using Kassandra algorithm (tumour mode)56. SCNA genomic instability score was established as follows: for each sample, the absolute copy number (ACN) profile generated by ASCAT through EaCoN was used. For each chromosome taken independently, the basal ACN level was identified as the one with the longest total width. Then, for each chromosome, ACN levels were converted into the absolute difference to this basal level [aDCN = (if the basis was 3 copies, a 1 copy segment value will be 2; 2 copies ≥ 1; 3 copies ≥ 0; 4 copies ≥ 1; etc.)]. The final SCNA score was computed as the width-ponderated sum of each of these converted copy number-to-basis values, divided by the total covered genome length.
Statistical methods
The data from IHC, flow cytometry (tumour biopsies and blood), secretome and plasma were processed with the dplyr package (1.1.4) for later use.
A paired two-sided Wilcoxon test was used to analyse statistical differences between baseline and week 3. A Wilcoxon test was used for statistical analysis between two conditions/groups (IT versus IV or no DCB versus DCB) at baseline. Statistical analysis was performed using the rstatix package (0.7.2) with the wilcox_test function, with the parameter paired = TRUE for paired analysis. Paired box plots and box plots were generated using the packages ggpubr (0.6.0) and ggplot2 (3.4.4). While the paired Wilcoxon analysis used only patients with a value between the two time points, the visual representation used all patients, even those not included in the Wilcoxon test. Centre values were defined as medians, consistent with the use of a non-parametric Wilcoxon test. All these analyses were performed using R software (v.4.3.3).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.