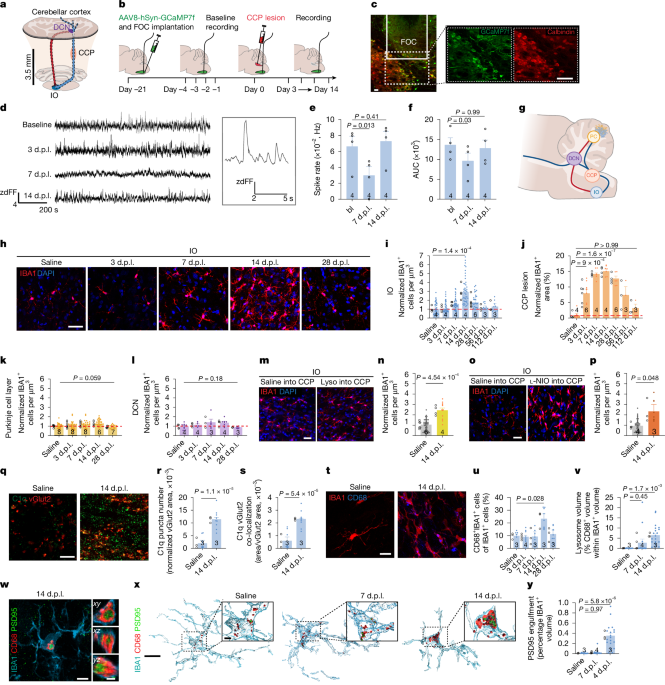
Animals
Experiments were performed in accordance with the EU guidelines for the care and use of laboratory animals, and with the guidelines of the UK Animals (Scientific Procedures) Act 1986 and subsequent amendments. Use of animals in this project was approved by the Animal Welfare and Ethical Review Body for the University of Cambridge and carried out under the terms of UK Home Office Licenses PP4353554, P9B1FBC4B and 70/7715, and the Animal Welfare and Ethical Review Body by the Icelandic Food and Veterinary Authority. All animals were maintained under a 12 h–12 h light–dark cycle with food and water supplied ad libitum, in individually ventilated cages at 20–24 °C and 45–65% relative humidity. All surgeries were performed aseptically under isoflurane anaesthesia, with standard pre- and post-operative analgesia and care. Animals were randomized into experimental groups. Experimenters were not blinded to experimental conditions during animal surgery and recovery, as lesioned animals, but not unlesioned controls, can develop side effects that require enhanced monitoring. However, subsequent tissue processing, imaging and quantification were performed blindly and validated by more than two experimenters. Experiments that did not require animal surgery were performed blindly at all stages of data collection and quantification.
Focal white matter demyelination in the CCP
EtBr model in rats
The CCP of 9–12-week-old female Sprague Dawley rats was bilaterally injected stereotaxically using a 10 μl Hamilton syringe. Four microlitres of EtBr (0.01% in saline) was injected at a rate of 1 μl min−1 to induce focal white matter demyelination. CCP coordinates were −7.3 mm (dorsal–ventral, DV); −2.3 mm (anterior–posterior, AP); ±2.6 mm (medium–lateral, ML) from lambda. In ageing experiments, CCP demyelinating lesions were induced in 18 month-old Sox10:dsRed rats.
EtBr model in mice
For mice, 3–5-month-old female C57Bl/6 mice were stereotaxically injected bilaterally with a mixture of EtBr and Dextran AlexaFluor 488 (Thermo Fisher Scientific; 1.50 μl, 0.01% and 2%, respectively, diluted in saline) at a rate of 0.25 μl per minute into the CCP; AP: −5.90 mm, ML: ±2.00 mm, DV −4.40 mm relative to bregma.
Lysolecithin model
Female Sprague Dawley rats (aged 9–12 weeks) were bilaterally injected with 2 µl of 1% lysolecithin (Sigma-Aldrich) in the CCP. Coordinates were AP: −2.3 mm; ML: ±2.6 mm, DV: −7.3 mm, relative to lambda.
Focal white matter hypoxic injury
Female Sprague Dawley rats (aged 9–12 weeks) were unilaterally injected in the CCP with 2 µl of 27 mg ml−1 L-NIO (N5-(1-iminoethyl)-L-ornithine) (Cayman Chemical), an irreversible inhibitor of nitric oxide synthase enzymes30. Two per cent Alexa-488 dextran was added to the injected mixture to retrogradely label IO neurons of which axons were exposed to L-NIO. Coordinates were as follows: AP, −2.3 mm; ML, ±2.6 mm; DV, −7.3 mm, relative to lambda. Rats were perfuse fixed at 14 d.p.l.
Focal white matter lesion in the corpus callosum
Female C57Bl/6 mice (aged 6 months) were stereotaxically injected with 1 µl of lysolecithin (1%) into the corpus callosum. Coordinates were as follows: AP, −2.5 mm; ML, −0.8 mm; DV, −1.2 mm, relative to bregma. Mice were perfuse fixed at 14 d.p.l.
Focal white matter demyelination in the cingulum
Female C57Bl/6 mice (aged 3–4 months) were stereotaxically injected with a mixture of EtBr and Dextran Alexa Fluor 488 (Thermo Fisher Scientific; 1 µl, 0.01% and 2% respectively, in saline) at a rate of 0.25 μl min−1, into the following cingulum coordinates: AP, +1.5 mm; ML, ±1.1 mm; DV, −2.1 mm, relative to bregma. Mice were perfuse fixed at 14 d.p.l.
Chronic model of demyelination
To generate a model of failed myelin regeneration we capitalized on the knowledge that OPCs are highly proliferating progenitors, sensitive to EdU51,52. Thus, following focal white matter demyelination with injection of EtBr into the CCP of 9–12-week-old female Sprague Dawley rats, a 30-gauge cannula was implanted 100 µm above the lesioned area and connected to an osmotic minipump for local delivery of 10 µg µl−1 EdU (Sigma-Aldrich), as described previously21.
Fibre photometry
Neuronal activity recordings
For neuronal activity recordings, the calcium indicator pGP-AAV-syn-jGCaMP7f-WPRE53 (1 μl, 5 × 1012 viral particles per ml; Addgene) or pGP-AAV-syn-jGCaMP8m-WPRE54 (1 μl, 5 × 1012 viral particles per ml; Addgene) were packaged into AAV8 particles and injected in the IO (DV, −10.6 mm; ML, ±0.55 mm, relative to lambda; and AP, −13 mm relative to bregma) to drive the expression of GCaMP7f or GCaMP8m in IO neurons. GCaMP8m has comparable temporal detection range as GCaMP7f but with greater signal to noise ratio54. During the same procedure, an FOC (200 µm in core diameter; numerical aperture, 0.37; Neurophotometrics) was implanted and secured on the skull surface using a pair of surgical screws (Plastic One) and dental cement (C&B Superbond). Fibre photometry was performed using a FP3002 console (Neurophotometrics). Ca2+-bound GCaMP7f/GCaMP8m was excited at 470 nm, (LED power set to 50 µW); fluorescence from Ca2+-unbound GCaMP7f/GCaMP8m was also recorded (excitation at the isosbestic point, 415 nm) to control for motion artefacts and correct for photobleaching.
Extracellular ATP measurements
For in vivo recordings of ATP release, 9–12-week-old female rats were unilaterally injected in the IO with 1 μl of AAV8 particles containing pAAV-hsyn-GRABATP1.0 (2.5 × 1012 viral particles per ml; Addgene) and pAAV-hSYn-mCherry (2.5 × 1012 viral particles per ml; Addgene). GRABATP1.0 is a modified human P2Y receptor with permutated enhanced GFP that allows for real-time sensing of extracellular ATP55. During the same surgery, an optical fibre cannula was implanted, as described for GCaMP recordings. Dual-colour fibre photometry was performed using the FP3002 console (Neurophotometrics), with GRABATP1.0 excited at 470 nm and mCherry excited at 560 nm. The mCherry fluorescence signal was used to correct for motion artefacts and photobleaching.
In recordings for both neuronal activity and ATP extracellular levels, the sessions were 10 min long, during which animals were allowed to freely explore a cage identical to their home cage. After data acquisition, baseline correction and signal standardization were performed and normalized dF/F was calculated in MATLAB (R2025a) using a published script56. In brief, fluorescence intensity values recorded from the 470 nm (Int470) and 415/560 nm (Int415/Int560) LEDs were subtracted from the respective intensity median and divided by the s.d. The standardized zInt415/560 signal was fitted to the zInt470 signal using non-negative robust linear regression. Parameters of the linear regression were then used to find fitted zInt415/560 values (fitzInt415) and zdF/F calculated using the equation zdF/F = zInt470 − fitzInt415/560 and plotted against recording time. GraphPad Prism v.10.5.0 was used for detecting peaks in activity traces (threshold set to 1–2 s.d.) and calculate the AUC. For ATP recordings, the average raw GRABATP1.0 fluorescence signal was plotted after normalization to the raw mCherry fluorescence signal.
Combined focal drug delivery and neuronal activity recordings
For simultaneous activity recording and microglial depletion, a custom-made device, conjugating optical fibre and drug-delivery cannula, was manufactured. Both components were positioned in the device such that the optic fibre could record data from the region of interest of drug delivery; the angle of offset between both the fibre and cannula being 15°. Devices were built using an Asiga MAX UV SLA 3D Printer with PrintoDent GR-10 UV curable (385 nm) biocompatible resin. Once printed, the devices were sonicated in isopropyl alcohol twice for 15 min, then exposed in a ultraviolet (UV) crosslinker for 15 min to ensure that the resin was fully cured. Devices were then subsequently assembled by inserting the optic fibre into place in the device and adhering it with Loctite 4305 LC, a UV curable cyanoacrylate resin. The cannula was not adhered before the experiment to enable correct placement during the surgical procedure.
Immunofluorescence
Animals were transcardially perfused with heparinized PBS, pH 7.4 (PBS) followed by 4% paraformaldehyde (PFA) in PBS. Brains were dissected and post fixed in 4% PFA for 1 h at 4 °C with gentle shaking. The brains were then either processed into 100-μm-thick vibratome sections or prepared for cryosectioning by incubation in 30% sucrose for cryoprotection, embedded in OCT, then cut into 10-μm-thick cryosections. Vibratome sections were blocked at room temperature with 10% goat or donkey serum and 0.5% Triton X-100 in PBS for 4 h followed by an overnight incubation with primary antibodies at room temperature. The primary antibodies were diluted in PBS. Sections were then washed with PBS three times, 20 min per wash, and incubated overnight at 4 °C with the appropriate secondary antibodies diluted in PBS. The sections were washed twice with PBS, incubated with DAPI (1 µg ml−1) for 10 min, washed with PBS and mounted in Fluoromount G. Coverslips of cultured cells and cryosections had minor changes to the protocol. Blocking was performed using 10% goat or donkey serum and 0.5% Triton X-100 in PBS for 1 h at room temperature; overnight primary incubation was performed at 4 °C; and secondary antibody incubation was performed for 1 h at room temperature. Images were collected using the Leica SP8 confocal microscope and Leica application suite X (LASX) software. Large brainstem overview z stacks were taken using an Evident VS200 slide-scanning system running SILA (speckle illumination acquisition), with a ×20 objective and 2.36 μm z spacing.
Spectral confocal reflectance microscopy was performed as described previously57. In brief, images of 100-μm-thick brain slices were taken using 488 nm, 561 nm and 633 nm laser light and reflectance was detected ±5 nm around the respective excitation wavelength.
The following primary antibodies were used in this study: rabbit CD68 (Abcam, ab125212, 1:100), mouse calbindin (Abcam, ab82812, 1:100), rabbit calbindin (Abcam, ab108404, 1:200), rabbit C1q (Abcam, ab182451, 1:100), mouse NG2 (Abcam, ab50009, 1:200), rabbit COX-IV (Abcam, ab16056, 1:100), chicken anti-m-cherry (Abcam, ab205402, 1:500), chicken anti-GFP (Abcam, ab13970, 1:1000), rabbit GABAARγ1 (Abcam, ab238130, 1:100), mouse P62 (Abcam, ab56416, 1:100), chicken GFAP (Antibodies.com, A85307, 1:300), goat IBA1 (Antibodies.com, a82670, 1:100), chicken MAP2 (Abcam, ab5392, 1:2000), rabbit IBA1 (Wako, 019-19741, 1:500), mouse PSD95 (Merck Millipore, MAB1596, 1:100), guinea pig vGLUT2 (Merck Millipore, AB2251-I, 1:500), mouse bassoon (Abcam, ab82958, 1:100), goat LDLR (R&D Systems, AF2255, 1:100), rabbit Olig2 (Merck, AB9610, 1:300). Secondary antibodies (all used at 1:500 dilution) include goat anti-rabbit Alexa Fluor 488 (Thermo Fisher Scientific, A32731), goat anti-rabbit Alexa Fluor 568 (Thermo Fisher Scientific, A11036), goat anti-rabbit Alexa Fluor 647 (Thermo Fisher Scientific, A21247), goat anti-mouse Alexa Fluor 750 (Thermo Fisher Scientific, A21037), goat anti-mouse Alexa Fluor 647 (Thermo Fisher Scientific, A21242), goat anti-mouse Alexa Fluor 555 (Thermo Fisher Scientific, A21424), goat anti-mouse Alexa Fluor 488 (Thermo Fisher Scientific, A11029), goat anti-mouse Alexa Fluor 568 (Thermo Fisher Scientific, A11031), donkey anti-goat Alexa Fluor 633 (Thermo Fisher Scientific, A21082), donkey anti-Rabbit Alexa Fluor 488 (Thermo Fisher Scientific, A-21206), goat anti-hamster Alexa Fluor 488 (Abcam, ab173003), goat anti-guinea pig Alexa Fluor 647 (Abcam, ab150187) and goat anti-chicken Alexa Fluor 568 (Abcam, ab175477). For microglial densities, CD68 expression, neuron counting, OPC counting and area measurement analysis, imaging used a ×20 objective with ×2 digital zoom, and z stacks were set up with 10 optical sections and 2.5 µm step size. Quantification of total number of IBA1+ microglia and the fraction of microglia expressing CD68 was performed in a blinded manner in ImageJ (v.1.54p). Areas containing the surgical needle track were excluded from the analysis. For analysis of synaptic densities, Zeiss LSM 980 with a Airyscan 2 module and Zen blue software (v.3.7.97.0.7000) were used. Imaging used the super-resolution mode (×2 sampling), with a ×63 objective and ×2 digital zoom, with z stacks set up to contain ten optical sections, with a 120 nm step size. Quantification of synaptic puncta density was performed in a blinded manner in ImageJ, using SynapseJ58. In all of the experiments, the laser power was adjusted to achieve a full dynamic range of intensity values.
Microglial morphology and engulfment of synaptic material
One-hundred-micrometre thick vibratome sections were immunostained for IBA1, CD68 and PSD95 or bassoon. For each animal, three to five regions of interest within the IO were imaged using the Leica SP8 confocal microscope and Leica application suite X (LASX) software. Images were obtained using a ×63 oil immersion objective (NA 1.4) with a ×2 optical zoom and 0.3 μm optical section thickness. The pixel resolution was 1,024 × 1,024 and the voxel size was 0.090 × 0.090 × 0.297 μm3. The field of view was 92.4 × 92.4 μm2 and each stack contained 33–50 planes (10–15 μm). Raw images were processed on Imaris (Bitplane) v.9.1 and v.10.2 for analysis. Imaris Filament Tracer (Autopath) was used to reconstruct cell morphology and quantify total cell process length, number of branch points and number of Sholl intersections. The estimated largest diameter of soma to calculate starting points was 6–8 μm. To calculate seed points, the thinnest and largest diameters of microglia processes were 0.2 μm and 2.00 μm, respectively. Starting point and seed point thresholds were adjusted accordingly. Cell body roundness was quantified in FIJI (v.1.54p). Microglia cells were chosen at random for analysis. Autothreshold of the IBA1 channel was performed to saturate the outline of the cell body. The long and short axes of the cell bodies were measured (in μm). For each cell, the short axis was divided by the long axis to obtain an index measure of roundness from 0–1; a score of one corresponds to perfect roundness, and a score of zero corresponds to the opposite. Roundness index = short axis (μm)/long axis (μm). Soma size was quantified in the commercially available cloud-based deep learning platform (Aiforia Create, Aiforia Technologies, https://www.aiforia.com/). An algorithm was trained using a supervised learning approach to recognize and trace the two-dimensional outline of microglial cell bodies. Six hundred diverse and representative microglial cell bodies across all groups were manually annotated and cycled through 19 iterative training sessions to create a generalizable model. The algorithm was validated by analysis of untrained images.
Microglial engulfment analysis
Imaging proceeded as for morphological analysis, with cells randomly selected for analysis. Imaris 3D surface rendering was used to analyse engulfment. First, a surface was created for the IBA1 channel. The CD68 channel was then masked to the IBA1 surface to obtain CD68 fully contained within IBA1 cells. A surface was then rendered for the masked CD68 channel. A mask was applied within CD68+ reconstructed lysosomes for PSD95 channel to obtain synaptic proteins inside the CD68+ lysosomes. Surfaces were created for the masked PSD95 channel. Data were calculated and presented as an engulfment index: (volume PSD95 within CD68+ lysosomes/IBA1 volume) × 100.
Analysis of microglial–neuron interactions
Vibratome sections (thickness, 100 µm) were stained for IBA1 and calbindin. Images of the IO were obtained using a ×20 objective with a ×2 optical zoom and 2.3 μm optical section thickness. Pixel resolution was 1,024 × 1,024 and voxel size 0.284 × 0.284 × 2 μm3. The field of view was 291 × 291 μm2 and each stack contained 5 planes (10 μm). Maximum-intensity projections of full z stacks were generated in ImageJ and all IBA1+ microglia in the field of view were categorized as interacting or non-interacting with calbindin+ neurons. Interacting microglia comprised microglia extending processes that contacted neurons, microglia with cell bodies in juxtaposition to neuronal cell bodies and microglia in which cell bodies and processes enwrapped neuronal cell bodies59. The density of interacting microglia was obtained by dividing absolute numbers of interacting microglia by the volume of the z stack.
Tissue culture
Cells were maintained at 37 °C and 5% CO2. Primary mixed glial cultures, OPCs, and microglia were prepared as described previously60. OPCs were resuspended in 50% DMEM, 50% Neurobasal, with modified SATO, 10 ng ml−1 biotin, 2% B27 and 1× GlutaMAX (Invitrogen), and 1 × 105 OPCs were plated per well onto 12-mm-diameter PDL-coated glass coverslips in a 24-well plate, with daily addition of growth factors (PDGF-aa at 10 ng ml−1 and FGF-b at 10 ng ml−1; Peprotech). Microglia were plated at 1 × 104 cells per well and cultured in DMEM containing 10% FCS (Life Technologies) and 1% penicillin—streptomycin; after 24 h, half of the medium was changed to the same culture medium as used for the OPCs. EdU (10 µM) was applied to the OPC and microglial cultures the day after plating; cells were fixed in 4% PFA 48 h after EdU application.
Fast Blue labelling and flow cytometry
The CCP was lesioned and, in the same procedure, injected with 0.5 µl of 0.5% Fast Blue for retrograde labelling of demyelinated neurons in the IO and associated microglia. Then, 3, 7, 14 and 28 days after lesion, rats were decapitated, and microdissected IO tissue was digested with papain solution in DMEM buffer for 30–45 min at 37 °C. Cells were triturated to single cells after blocking papain activity with ice-cold soybean trypsin inhibitor solution, then passed through a 40 µm strainer. Cells were separated from myelin debris using the myelin removal microbead kit from Miltenyi (130-096-733) according to the instructions. Dead and dying cells were labelled with propidium iodide for 5 min in PBS without Ca2+ and Mg2+ supplemented with 0.5% BSA fraction V before analysis. Cells from control cortical tissue were heat-shocked at 45 °C for 2 min to provide a positive control. The samples were analysed on the Beckton Dickinson FACS Aria2 system using FACSDiva (v.9.0.1) software. Single-cell gating and Fast-Blue-negative and propidium-iodide-negative populations were defined using cortical cells and heat-shocked cortical cells.
Spatial DBiT-seq
Spatial transcriptomic sequencing was performed according to the previously published protocol and reagents61. In brief, 76 × 52 mm slides with four mounted lesion and IO sample sets (8 coronal sections; see the diagram in Fig. 2a) were thawed at room temperature for 10 min then fixed in 0.2% formaldehyde (−methanol) for 5 min at room temperature. The fixation was quenched with 1.25 M glycine for 5 min.
In situ reverse transcription was performed by adding reverse transcription mix in a humidifying chamber, the tissue was first incubated at room temperature for 30 min, then at 42 °C for 90 min. The samples were washed with 1× NEB Buffer 3.1 plus RNase Inhibitor and bright-field imaging was performed using the Zeiss Cell Observer system.
In situ spatial ligation barcoding was completed by first loading barcode A ligation reactions (100; Fig. 2a) into the vertical microfluidic PDMS chip followed by the second round of barcode B ligation reactions (100; Fig. 2a), which were run on a horizontal 5 pass PDMS chip. In situ barcoding with the ligation mix for each round of barcoding was prepared and a total of 4 μl of ligation mix was added to each 1 µl barcode + linker mix (A001-A100 or B001-B100) with additional loading of Alexa Fluor 488 dye and 647 dye in position 1 and 100 to demarcate the region of interest borders. The cassette, PDMS chip and slide were assembled so that the fluidic lanes were bypassing each region of interest, then 4.5 µl of the barcoded ligation mix was added to each microfluidic lane on the input side of the chip. A vacuum gasket was set over the outlet wells, and 200–300 p.s.i. of suction pressure was applied for 7 to 10 min. Lane-loaded barcoded ligations with the cassette intact were incubated for 30 min at 37 °C in a humidifying chamber. Cassette disassembly and sample washing was done before in situ B barcoding (following the same steps as for DAPI staining and bright-field and fluorescence imaging using the Zeiss Cell Observer system).
Reverse crosslinking of each region of interest was separately processed by first assembling a PDMS chip (with five regions of interest) cassette on each slide. Reverse crosslinking buffer was added to each well, then incubated at 58 °C for 2 h. Each lysate was transferred to an individual tube then further incubated overnight at 65 °C with agitation.
The cDNA library was constructed by first performing streptavidin-bead affinity pull-down then template switching using template-switch oligo mix that was added to each bead sample. The samples were incubated for 30 min at room temperature then for 90 min at 42 °C. The beads were washed then resuspended in PCR solution. Thermocycling was done (stage 1, 1 cycle, 95 °C for 3 min; stage 2, 5 cycles, 98 °C for 20 s, 65 °C for 45 s, 72 °C for 3 min; stage 3, 1 cycle, 72 °C for 1 min, 4 °C hold). The supernatant was removed from the streptavidin beads and a quantitative PCR (the same thermocycling programme except the 65 °C cycle was 20 s instead of 45 s) assessment to determine the final cycle number to achieve 1/3 of saturation signal was completed. Additional cycles were run on the remaining sample according to quantitative PCR results. Then SPRI bead cleanup was performed, and samples were eluted in EB buffer.
To prepare the final library for sequencing, the mRNA-derived cDNA was quantified using Qubit and BioAnalyzer assessment. Depending on results further purification was performed as needed. Nextera XT Library Prep Kit was used and the Tagment Mix was incubated at 55 °C for 5 min. NT buffer was added and the sample was then incubated at room temperature for 5 min. The final PCR mix was added and thermocycling was completed (stage 1, 1 cycle, 95 °C for 30 s; stage 2, 12 cycles, 95 °C for 10 s, 55 °C for 30 s, 72 °C for 30 s; Stage 3, 1 cycle, 72 °C for 4 min, 4 °C hold). The concentration and size distribution were determined using the Agilent Bioanalyzer High Sensitivity Chip. Next-generation sequencing was completed at NGI (National Genomics Infrastructure) using Illumina NovaSeq X Plus 25B in paired-end 150 bp mode with a total of 6 million reads for each sample and a 15% PhiX sequencing step.
Samples at 28 d.p.l. did not pass quality control and were excluded from all analysis.
DBiT-seq bioinformatics analysis
Tissue images acquired using the Zeiss Cell Observer microscope were imported into R (v.4.3.3) using the imager package (v.1.0.2)62. PDMS chip imprints were manually identified in each image and extracted. Extracted regions were converted to greyscale, and morphological operations (pixel erosion and dilation) were applied to reduce imaging artefacts while preserving tissue structure. To define the cassette pixel layout, the dimensions of the extracted tissue image were divided by the number of channels in the cassette, generating a grid corresponding to cassette channels positions. For each cassette pixel location, the number of underlying image pixels was aggregated to produce a pixel-intensity distribution mapped to the cassette structure. To correct for imaging artefacts—particularly in regions lacking tissue or along cassette borders—cassette pixel values were adjusted using a combination of internal pixel intensities, neighbouring pixel values and proximity to the chip edges. Regions with obvious absent tissue were identified and used as a distribution threshold to exclude off-tissue grid coordinates. The final set of valid cassette coordinates was exported as a tab-delimited file for downstream analyses.
A total of 15 transcriptomes was sequenced at the in-house sequencing facility. Spatial barcode demultiplexing and FASTQ file preparation were performed using Snakemake (v.7.24.0)63 as the workflow management system. Sequencing reads from multiple lanes were merged into single FASTQ files per sample. Reads were filtered out using bbduk (v.39.01)64, if either PCR primers or spatial linkers were missing from the read construct. Spatial barcodes were extracted from sequencing reads using a custom Python script and written to separate FASTQ files. A predefined whitelist of spatial barcodes was supplied—annotation following the in situ spatial ligation barcoding—and cellranger count (v.7.1.0) was used to align reads to the Rattus norvegicus reference genome (mRatBN7.2, RefSeq annotation, NCBI). Tissue images, spatial barcode coordinates and gene count matrices were imported into R. Count matrices were loaded into a Seurat object (v.5.1.0)65, retaining only barcodes corresponding to tissue-covered pixels. All of the samples were subsequently merged into a single expression matrix for downstream analysis.
To account for potential lane clogging inherent to the DBiT-seq technology, which can result in abnormally low transcript counts in affected lanes, we implemented a filtering and imputation strategy. Vertical and horizontal lanes exhibiting total gene counts substantially below the distribution observed across the full dataset were identified as outliers and set to zero. Subsequently, gene expression values in these lanes were imputed by averaging the expression values of their immediately adjacent lanes. This correction was applied solely for purposes of clustering and visualization to improve spatial coherence. All DGE analyses were performed on the original, uncorrected expression matrix.
Gene count matrices were normalized by total library size, scaled by a factor of 10,000 and natural-log-transformed. Genes were then standardized by centring their mean expression and dividing by their s.d. Dimensionality reduction was performed by principal component analysis (PCA) on the 2,000 most variable genes, selected using the vst method. The first 30 principal components (PCs), determined by elbow plot inspection, were retained for downstream analysis. Samples were integrated using the IntegrateLayers function in Seurat, applying the reciprocal PCA (RPCA) integration method.
Neighbour graphs were constructed from the selected PCs using the FindNeighbors function with the ‘annoy’ approximate nearest neighbour algorithm, Euclidean distance metric and 50 trees. The number of nearest neighbours (k) was set to 20. Unsupervised clustering was performed using the FindClusters function with the Louvain algorithm and a resolution parameter of 1.2. For visualization, uniform manifold approximation and projection (UMAP) was computed from the first two dimensions of the neighbour graph.
IO gene expression markers were used to delineate an IO-specific pixel cluster from whole-tissue datasets. For each tissue, a k-nearest neighbour (k-NN) graph (k = 5) was constructed using the get.knn function from the FNN package (v.1.1.4.1)66 and subsequently converted into a graph object with graph_from_edgelist from the igraph package (v.2.1.4)67. Clusters within IO-associated pixels were identified using the components function. The largest cluster was defined as the principal IO cluster, with additional clusters of at least half the main cluster size to be aggregated to the main cluster. To exclude peripheral pixels from the main cluster, we applied a 0.5 quantile threshold on average graph edge distances per node. The coordinates of pixels within the main cluster were then dilated by a radius of two pixels in all directions to generate a contiguous IO region of interest.
Lesion sites were delineated manually, guided by staining of IBA1 immunohistochemistry on adjacent sections delineating the lesion location, along similar anatomical landmarks. Control regions were defined as 10 × 10 pixel squares on the ipsilateral side, spatially separated from the lesion.
To characterize calbindin-expressing neurons within the IO, all Calb1+ pixels were selected for subsequent analyses (total 3,442 pixels out of total 10,864 pixels allocated to the IO region). To characterize microglia within the lesion, a microglia score was computed using either three (P2y12r, Tmem119 and Cx3cr1 identified 1,289 pixels) or four marker genes (Aif1, Maf, Spi1 and Csf1r; 2,874 pixels identified) with the AddModuleScore function from Seurat, specifying ctrl = 100 and nbin = 24. Both approaches gave similar results. To characterize microglia within the IO, a microglial score was computed using four marker genes (Aif1, Maf, Spi1 and Csf1r) using the AddModuleScore function from Seurat, specifying ctrl = 100 and nbin = 24. Pixels with a microglia score greater than 0.05 were extracted from downstream analyses.
Gene count matrices were normalized to total library size, scaled by a factor of 10,000 and log-transformed (natural log). Gene expression values were then standardized by centring on the mean and scaling by the s.d. Dimensionality reduction was performed using PCA on the 1,000 most variable genes, identified using the vst method. The number of retained PCs—20 for IO, 20 for lesion, 20 for IO calbindin neurons and 15 for IO microglia—was determined by elbow plot inspection. Neighbour graphs were constructed from the selected PCs using FindNeighbors with the annoy approximate nearest-neighbour algorithm, Euclidean distance metric and 50 trees, with k set to 20. Unsupervised clustering was carried out using FindClusters (Louvain algorithm) at resolution parameters of 1 (IO), 0.5 (lesion), 0.8 (IO calbindin neurons) and 0.7 (IO microglia). For visualization, UMAP was computed from the first two dimensions of the neighbour graph. The number of pixels in the top levels of the chord/circos plots corresponding to each bottom-level category was quantified. To enable an unbiased visualization of proportional relationships, top-level categories were downsampled before plotting.
Bulk RNA-seq libraries
After the flow sorting above, Fast Blue+propidium iodide− cells were isolated directly using a 100 µm nozzle into 900 µl volume of RLT buffer of Qiagen RNeasy micro kit without β-mercaptoethanol. The isolated cell lysate was kept at 4 °C during the isolation process. Total RNA was isolated using the Qiagen RNeasy micro kit and kept at −80 °C until use. The RNA integrity number (RIN) was calculated for each sample for quality control, with a mean RIN of 9.57. A mRNA library was prepared using SMARTer Stranded Total RNA-Seq Kit v3 – Pico Input Mammalian (Takara Clontech) according to manufacturer’s instructions. Data were read on the HiSeq4000 system; a total of >350 million 150 bp strand-specific paired-end reads were generated.
Bulk RNA-seq deconvolution and bioinformatics analysis
R. norvegicus (Rn7) bulk RNA-seq raw count matrixes were converted to Mus musculus (Mm10) using the EnsEmbl Biomart database. Only orthologue genes with a conservation score of at least 80 between mm10 and Rnor7, ENSEMBL v.102, were selected for downstream analyses. The final number of genes was reduced from 33,295 genes to 15,678 orthologue genes.
Deconvolution was performed in R v.4.5.0 environment. For the single-cell RNA-seq reference, the medulla matrix from ref. 68 was downloaded (https://cells.ucsc.edu/?ds=mouse-nervous-system)69 and converted to SingleCellExperiment70 R object with SubClass and SampleID metadata labels. The raw count matrix with mouse annotations and ref. 68 medulla data were used as an input for MuSiC R package71,72. MuSiC was run with function music_prop for neurons and microglia (labelled as immune previously68) using the sample IDs from ref. 68. For each gene, the mean expression in each sample was first calculated. Then, based on the microglial and neurons deconvoluted proportions from MuSiC an inferred relative gene proportion was calculated. The individual gene inferred proportions were then rounded and converted to inferred counts. This measure is an approximation and is dependent on the proportions predicted from MuSiC.
The expression of the top 200 fold-change (versus 28 d.p.l.) genes is shown as heat maps in Fig. 3d. Heat maps were plotted using ComplexHeatmap R package73 (ComplexHeatmap (Bioconductor; http://bioconductor.org/packages/ComplexHeatmap/). Final visualization shows the scaled z scores of the top 200 fold-change (versus 28 d.p.l.) genest (scale function center = TRUE). GO term analysis was performed using EnrichR v.3.4 package74.
Analysis of mitochondrial density and morphology
Female rats (aged 9–12 weeks) were injected with 1 µl of pAAV2-CAG-MitoDsRed-IRES-GFP (a gift from J. Kittler) to genetically label mitochondria in the IO. The coordinates for stereotaxic injections were as follows: DV, ±10.6 mm; ML, ±0.55 mm relative to lambda; and AP, −13 mm relative to bregma. Then, 3 weeks after viral injection, the CCP was lesioned or injected with saline and rats perfused at 3 d.p.l. Then, 100-µm-thick parasagittal sections were produced with a vibratome and stained for IBA1 and calbindin.
For analysis of mitochondria in calbindin+ neuronal cell bodies, images were taken on a Leica SP8 confocal microscope with a ×63 objective, ×3 digital zoom and pinhole at 0.35 AU, and z stacks were taken with a 0.15-μm step size to image the entire cell. Images were deconvolved using Huygens Professional, following default settings and using 20–30 iterations. Deconvolved images were imported into FIJI and analysed in three dimensions using the Mitochondria Analyzer plugin, as previously described75. Thresholding and image processing were used according to the plugin workflow. Analyses were performed at both the per-cell and per-mitochondrion levels. Per-cell analysis was defined as the mean measurement of all mitochondria within a cell body; per-mitochondrion analysis represented measurements of each individual mitochondrion measured. The following parameters were extracted: mitochondrial density (mitochondrial count normalized to cell body volume in µm3), mitochondrial volume (summed volume of all mitochondria normalized to cell body volume) and volume-weighted mitochondrial sphericity (sphericity weighted by each mitochondrion’s contribution to total mitochondrial volume). Values closer to 1 indicate more spherical morphology. The formulas used for these calculations were adapted from the Mitochondria Analyzer documentation. If Si is the sphericity of mitochondrion i, Vi is its volume and SAi is its surface area:
$${S}_{i}=\frac{{{\rm{\pi }}}^{1/3}\times {(6\times {V}_{i})}^{2/3}}{{\mathrm{SA}}_{i}}$$
$${\text{Average Cell}\, S}_{\mathrm{weighted}}=\frac{{\sum }_{i=1}^{n}{V}_{i}\times {S}_{i}}{{\sum }_{i=1}^{n}{V}_{i}}$$
Cell body volume was estimated from the longest x and y axes of the soma together with the z-stack depth, using an ellipsoid volume approximation:
$${V}_{\mathrm{cell}}=\frac{4}{3}{\rm{\pi }}\times \frac{x}{2}\times \frac{y}{2}\times \frac{z}{2}$$
For analysis of mitochondria in neuron–microglia junctions, images were taken with a Leica Stellaris confocal microscope with ×63 objective, ×4 digital zoom and pinhole at 0.35 AU and z stacks were taken with 0.15 μm step size. Images were deconvolved using Huygens Professional, using the default settings and 20–30 iterations. Mitochondria in neuron–microglia junctions were analysed using Imaris 10.2 (Bitplane). In brief, surfaces were generated from both the IBA1 and the MitoDsRed channels, slightly exaggerated to highlight the overlap between volumes. These surfaces were then used to mask the individual channels, and the masked channel was used to create faithful surfaces for IBA1 and MitoDsRed. Segmented mitochondria statistics were exported from Imaris for downstream analysis.
Local microglial depletion
Female rats (aged 9–12 weeks) were stereotaxically injected with EtBr (4 µl, 0.01% in saline) to induce a demyelinating lesion in the CCP. During the same procedure, a 30-gauge cannula (Plastics One) was implanted into the IO and connected to an osmotic minipump (Alzet Micro-Osmotic Pumps, model 1004, DURECT), with a flow rate of 0.11 μl h−1. PLX5622 (10 μM; MedChemExpress), a potent Csf1r blocker, was continuously delivered to the IO to locally deplete microglia.
Histological analysis of remyelination
For analysis of remyelination levels in the lesion, rats were perfused with 4% glutaraldehyde in phosphate buffer, followed by 5 days of post-fixation. One-mm-thick sections containing the lesion were further fixed in 2% osmium tetroxide, overnight, at 4 °C; washed three times in PBS; dehydrated using 70%, 95% and 100% ethanol, followed by 100% propylene oxide and embedded in resin. Blocks of resin-embedded tissue were sectioned (1 μm), stained with 1% toluidine blue and imaged on a Zeiss Axioscan (×40 objective). The remyelinated area was quantified using Halo-AI v.3.6.4134 (Indica labs). Image analysis pipeline was generated by training an AI-based DenseNet network to automatically quantify the remyelinated area within focal white matter lesions at 28 d.p.l. The network was trained to segment the lesion area into remyelinated and non-remyelinated regions. More than 500 annotations on 10 representative lesions were produced and the network was trained for over 10,000 iterations with manual correction cycles until achieving excellent classification accuracy (cross-entropy < 0.001). The classifier was validated using correlative electron microscopy analysis of adjacent sections and conventional ranking analysis conducted by five independent assessors (Extended Data Fig. 9). Analysis of remyelination by the automated segmentation, ranking and electron microscopy was highly correlated. Remyelination by Schwann cells was excluded in the analysis. For each analysis automated segmentation was inspected, if needed classified tissue areas manually corrected and the network was further trained for a total of three cycles. Remyelination was further assessed by ranking, performed blindly and independently by two scorers and by electron microscopy.
Electron microscopy
Toluidine-Blue-stained semithin sections used for automated histological analysis of remyelination were glued to empty resin blocks and repeatedly dipped into liquid nitrogen until they detached from the glass slide. Ultrathin sections were cut at 75 nm (Leica EM UC7 ultramicrotome, DiATOME Ultra 35° knife), transferred to copper grids and post-stained for 1 min with UA-Zero and 30 s with lead citrate. The lesion was identified in 4 × 4 0.5kx overview images, then ten images were taken at random lesion positions per animal (Hitachi HT7800 TEM, 80 kV, 3kx magnification, EMSIS Xarosa camera, 5,120 × 3,840 px, 4.97 nm per px). Quantification of the percentage of remyelinated axons and g-ratios were performed in ImageJ, remyelination by Schwann cells was not included in analysis.
Quantification and statistical analysis
All statistical analyses were performed in GraphPad Prism v.10.2.3 and v.10.5.0 or in Python (v.3.11.13). Data are shown as mean ± s.e.m., with the number of animals (that is, the biological replicates) indicated in the figures and Supplementary Table 1. For non-hierarchical data (single measurement per animal), two-tailed Student’s t-tests (with Welch’s correction when variances were unequal) were used for two-group comparisons. For more than two groups, one-way ANOVA was used (Welch’s ANOVA when variances were unequal), followed in all cases by Bonferroni post hoc comparisons to correct for multiple comparisons. Consistent with statistical best practises in neuroscience76,77,78,79, for hierarchical data (for example, cells or images nested within animals), nested analyses were applied to preserve within-animal variability. Two-group comparisons used nested two-tailed t-tests, and multiple-group comparisons used nested one-way ANOVA, followed by Bonferroni post-hoc comparisons. Where assumptions of normality (Shapiro–Wilk test) or equal variance of nested data were not met, we used hierarchical blocked bootstrapping or permutation testing. For transcriptomic data, differential expression analysis was performed using the Wald test, with FDR controlled using the Benjamini–Hochberg correction. For comparing two cumulative distributions, Kolmogorov–Smirnov tests were used. For comparison of Gaussian fits, two-sided Wald z-tests, with Holm correction, were used.
Power calculation was performed to determine sample size. Given the technical complexity of some of the experiments, sample sizes are necessarily limited; nonetheless, the reproducibility of the effects across animals and modalities collectively support the conclusions.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.