Materials availability
New plasmids associated with this study were deposited to Addgene under Mugnier Lab and McCulloch Lab and are associated with this publication.
Statistics and reproducibility
Sample size was not predetermined and experiments were not randomized or blinded. In vitro experiments were derived from two independently generated biological replicates, arising from different parentals except for the truncation experiments (same parental, but biologically unique clones), and offered high congruence sufficient for analysis. Mouse experimental size was selected based upon previous experiments with sufficient effect size observed1,9. WT versus μMT experiments were performed three independent times with independent vials of T. brucei stock using five mice of each genotype at the start of the infection. ΔRAD51 and Tb427VSG-8 infections in μMT mice were performed once from the same vial of stock, infecting four and five mice, respectively. Flow cytometry experiments were performed twice for Fig. 5b, with one sample, mosaic 5, repeated across both experiments, and once for Fig. 5c. Data presented are technical replicates. Antisera collected from mice represent four biological replicates.
A Shapiro–Wilk normality test was performed for all statistics calculated from mouse experiments, followed by the specified Wilcoxon test. Significance from FACS experiments was determined from a one-way ANOVA with post hoc Tukey HSD. All graphs include mean if n = 2 or mean ± s.d. if n ≥ 3. Calculations were performed in R (v.4.0.2). Details with exact P values and statistical test results can be found in Supplementary Data 2.
Parasites
Pleiomorphic EATRO1125 AnTat1.1 90-13 T. brucei parasites were maintained in HMI-9 media with 10% heat-inactivated FBS (F0960, 500 ml, Sigma-Aldrich) and 10% Serum Plus (14008C, 500 ml, Sigma-Aldrich) or in HMI-11 with 10% FBS where specified60. These parasites originated from K. Matthews. Parasites were passaged when they reached approximately 5 × 105 cells per ml unless otherwise specified. Monomorphic Single Marker Lister427 VSG221 TetR T7RNAP bloodstream form (NR42011; LOT: 61775530)61 T. brucei were maintained in HMI-9 up to 1 × 106 parasites per ml. VSG221 has since been renamed to VSG-2. Monomorphic Single Marker 427 1339 Cas9 TetR T7RNAP (bloodstream form) (NR-56793; LOT: 70056027) T. brucei subsp. brucei were maintained in HMI-9 up to 1 × 106 parasites per ml. This was obtained through BEI Resources, National Institute of Allergy and Infectious Diseases (NIAID), National Institutes of Health (NIH). EATRO1125 AnTat1.1 J1339 pleiomorphic parasites were a gift from K. Matthews62. Parasites were verified for expected VSG expression via amplicon sequencing. Parasite cultures were not tested for mycoplasma contamination.
Plasmids
Plasmids were synthesized with Gibson Assembly with a custom master mix63 unless otherwise specified. Whole plasmid sequencing was performed by Plasmidsaurus using Oxford Nanopore Technology and their custom analysis and annotation pipelines. Detailed description of plasmid construction, using published publicly available plasmids64,65,66,67,68,69, can be found in the Supplementary Methods.
Transgenic parasites
To obtain transgenic parasites, 5–20 million parasites were electroporated with 5–10 μg of digested plasmid with an AMAXA Nucleofector II using X-001 in Human T-cell Nucleofector Solution (Lonza, VPA-1002) or using Z-001 in Tb transfection buffer70. All parasites were maintained in selection unless otherwise specified. Detailed descriptions64,71,72,73,74 of the generation of transgenic parasites can be found in the Supplementary Methods.
T7-guide synthesis and purification
Guide RNAs were designed with EuPaGDT75. DNA fragments containing a T7 promoter and guide RNA sequence were synthesized as described previously76. PCR products from 12 identical PCRs were pooled and purified via ethanol precipitation.
Cas9 transient electroporations
Approximately 24 h before electroporation, parasites were seeded at a density of 83,000 parasites per ml and induced with 1 μg ml−1 doxycycline (Millipore Sigma, D9891-1G). The total culture volume was determined by the number of samples being electroporated; 12 ml was seeded per sample. Blasticidin selection (Cas9) and puromycin selection (silent VSG-228) were maintained, if applicable, and for Lister427 parasites expressing AnTat1.1, hygromycin (active VSG expression) was removed at this stage. Either 8 ml (Lister427) or 10 ml (EATRO1125) of the bulk parasite culture, approximately 5–10 million cells, was spun down for each sample. Medium was then removed, and parasites were resuspended in 100 μl of Human T-cell Nucleofector Solution. Each sample was electroporated using the X-001 program on the AMAXA Nucleofector II with approximately 1–1.5 μg of purified T7-guide in a volume less than 10 μl or a sample without any DNA as a negative control. Parasites were moved into 5 ml of HMI-9 in six-well plates to recover for 30 min then moved into 20-ml total in flasks to recover overnight. About 24 h after electroporation, parasites were counted and split. For EATRO1125, 12 million cells (or all cells if there were fewer than 12 million) were seeded into 60-ml total with blasticidin. For Lister427, 2 million cells were seeded into 20-ml total with blasticidin. At the 48-h time point, parasites were counted, collected and stored in TRIzol (Invitrogen, 15596026) for subsequent RNA extraction.
For the constitutively expressed Cas9 parasites, 1.7–2.3 μg of guide was electroporated into 5–10 million cells. Parasites were grown in 20–30 ml of HMI-11 media with 0.2 μg ml−1 puromycin for 24 h to recover, counted, then split into 60 ml of HMI-11 media with puromycin for the final 24 h. Twelve million cells (or all cells if there were fewer than 12 million) were seeded into 60-ml total. At the 48-h time point, parasites were counted, collected and stored in 15 μl of PBS with 150 μl of RNAlater (Invitrogen, AM7020) at −20 °C for subsequent RNA extraction.
Isolation of mosaic-expressing parasites
Parasites were isolated from inducible Cas9-expressing parasites. A plasmid expressing an sgRNA (either pT7-sgRNA or pLEW-T7-sgRNA) was inserted into each parental clone and then Cas9 was induced with doxycycline. Multiple dilutions of parasites were plated in 96-well plates and selected from plates with fewer than 30 surviving parasite clones. Upon isolation of a mosaic parasite clone, all drugs were removed.
EATRO1125 parasite clones expressing mosaic AnTat1.1 were isolated from parental lines with both pT7-sgRNA (guides 243 or 694; drugs: 5 μg ml−1 blasticidin and 1 μg ml−1 doxycycline) and pLEW-T7-sgRNA (guides 243, 694 or 1459; drugs: 5 μg ml−1 blasticidin, 0.1 μg ml−1 puromycin and 1 μg ml−1 doxycycline).
EATRO1125 parasite clones expressing mosaic Tb427VSG-8 were isolated from parental lines with pLEW-T7-sgRNA (guide 783; drugs: 5 μg ml−1 blasticidin, 0.1 μg ml−1 puromycin and 1 μg ml−1 doxycycline). EATRO1125 parasite clones expressing mosaic EATRO1125VSG-73 were isolated from parental lines with pLEW-T7-sgRNA (guides 194, 680 and 1436; drugs: 5 μg ml−1 blasticidin, 0.1 μg ml−1 puromycin and 1 μg ml−1 doxycycline).
VSG PCR and identification via sequencing
VSG sequences for clones were determined from extracted RNA. Complementary DNA was synthesized using the Superscript III Reverse Transcriptase (Invitrogen, 18080051) and a VSG-specific primer that binds to a conserved 14-bp sequence within the 3′ UTR (5′-GTGTTAAAATATATC-3′). Then, 2 μl of RNase-treated cDNA was amplified for 35 cycles with VSG-specific primers: a spliced-leader (5′-ACAGTTTCTGTACTATATTG-3′) and SP6-VSG 14-mer (5′-GATTTAGGTGACACTATAGTGTTAAAATATATC-3′) using Phusion polymerase (Thermo Fisher, F530L) (annealing temperature 55 °C, extension 45 s). PCR products were cleaned using the Monarch PCR & DNA cleanup kit (NEB, T1030L). VSG sequences were determined by amplicon nanopore sequencing performed by Plasmidsaurus using Oxford Nanopore Technology with their custom analysis and annotation in which fragmented VSG amplicons were sequenced and assembled into full-length sequences or targeted Sanger sequencing with the sequencing primer (5′-AGAGAATACTAAGCTAGTTGGC-3′) performed by Azenta Life Sciences.
Assessment of donor VSG following mosaic formation
Lister427 parasites containing constitutively expressed Cas9, active VSG AnTat1.1, a silent inserted VSG-228 donor and constitutively expressed guide targeting AnTat1.1 position 694 were expressing mosaic AnTat1.1 by the time they recovered from guide insertion, but only 1/12 positive colonies were clonal as determined by Sanger sequencing of the expressed VSG (5′-AGAGAATACTAAGCTAGTTGGC-3′) performed by Azenta Life Sciences. The VSG was amplified as detailed above in ‘VSG PCR and identification via sequencing’. The resulting sequencing traces were visually inspected to determine candidates likely to have a high concentration of cells with VSG-228 donation. These were subcloned and individual clones were obtained, with the analysis repeated to determine donor VSG. gDNA was isolated from cells with a VSG-228 donation. In brief, parasites were collected via centrifugation at 2,600g for 4 min. Parasites were washed once with around 500 μl of PBS, spun at 2,600g for 4 min and the PBS removed. DNA was extracted using the Monarch Spin gDNA Extraction Kit (NEB, T3010S). Approximately 40 ng of gDNA was used to amplify the donor VSG cassette, VSG-228, with one primer within the plasmid backbone and the other within the Blasticidin resistance gene (40 ng of input gDNA; Fwd: 5′-TTGACACCAGTGAAGATGCGG-3′; Rev: 5′-CGGCAGTTTACGAGAGAGATGA-3′; annealing temperature 60 °C, extension 90 s) for 35 cycles using Phusion DNA Polymerase (NEB, M0480L). The full sequence of the amplicon was determined by Plasmidsaurus using Oxford Nanopore Technology with their custom analysis and annotation. These were BLASTed (NCBI)77 against the original plasmid sequence.
Mouse infections
All experiments involving mice were performed in accordance with the protocol approved by the Institutional Animal Care and Use Committee at Johns Hopkins University. Mice were housed at 68–76 °C with 30–70% relative humidity (target 42%) under a 14.5 h:9.5 h light:dark photoperiod. The 8–12-week-old female C57BL/6 mice and µMT− (B6.129S2-Ighmtm1Cgn/J)35 mice (Jackson Labs) were infected with about five EATRO1125 parasites by intravenous injection in the tail vein. These parasites express either AnTat1.1 or VSG-421. Blood was collected 6 days after infection through a submandibular bleed. At 15 days after infection, mouse blood was collected by cardiac puncture.
The 8–12-week-old female µMT− mice were infected with about five Tb427VSG-8 EATRO1125 inducible Cas9 parasites. Blood was collected 6 days after infection through a submandibular bleed. At 13–15 days after infection, mouse blood was collected through a submandibular bleed. For one mouse, further blood was collected by cardiac puncture 18 days after infection.
The 8–12-week-old female µMT− mice were infected with about ten EATRO1125 ΔRAD51 parasites. Blood was collected 7–8 days after infection through a submandibular bleed. At 14 days after infection, mouse blood was collected by cardiac puncture.
Starting 4 days after infection, parasitaemia was monitored within mice every 2 days via tail bleed. After blood collection, extra gel packs and in-cage food pellets were provided to mice during recovery. Blood was stored in TRIzol LS for RNA extraction (Invitrogen, 10296028).
Tb427VSG-8/EATRO1125VSG-73 isolation
Tb427VSG-8-expressing EATRO1125 parasites were obtained by infecting a C57BL/6 mouse with five inducible Cas9 EATRO1125 AnTat1.1-expressing parasites intravenously. Starting 4 days after infection, parasitaemia was monitored within mice every 2 days via tail bleed. At 16 days after infection, the second peak of parasitaemia where AnTat1.1-expressing parasites are undetectable in the blood, mice were euthanized, and blood was collected via cardiac puncture. Parasites were cultured for 4 days in the presence of 5 μg ml−1 blasticidin and 1:100 Penicillin-Streptomycin (Gibco, 15140122) then frozen in HMI-9 with 10% glycerol. Parasites were thawed and subcloned into 96-well plates. Positive colonies were pulled from plates with fewer than ten colonies. Parasites were maintained in 5 μg ml−1 blasticidin. These parasites double roughly once per day.
EATRO1125VSG-73-expressing parasites were obtained by inducing Cas9 with 1.0 μg ml−1 doxycycline in EATRO1125 AnTat1.1-expressing parasites for 24 h. Parasites were then electroporated with approximately 2 μg of guide targeting the sequence upstream of AnTat1.1 (AnTat1.1 upstream target: 5′-CAAAAAGGAGGAGAGGAAAT-3′). These were cultured for 7 days in the presence of 1:1,000 rabbit anti-AnTat1.1 antibody (J. Bangs), 5 μg ml−1 blasticidin and 1.0 μg ml−1 doxycycline and then subcloned into 96-well plates. The clone used was pulled from a plate with fewer than ten colonies.
After obtaining single colonies with a novel VSG, VSG sequences were determined as described in ‘VSG PCR and identification via sequencing’.
RNA preparation
Parasites were stored in TRIzol before RNA extraction. Blood with parasites was stored in TRIzol LS. RNA was extracted via phenol/chloroform extraction according to the manufacturer’s protocol. Purified RNA was DNase treated with Turbo DNase (Thermo Fisher, AM2239) and purified with 1.8X Mag-Bind TotalPure NGS Beads (Omega Bio-tek, M1378-01). Verification of effective DNase treatment was performed via PCR of hygromycin (EATRO1125 only) (Fwd: 5′-ACAGCGGTCATTGACTGGAG-3′; Rev: 5′-ATTTGTGTACGCCCGACAGT-3′, annealing temperature 52 °C, extension 30 s) or HSP70 (Lister427 & EATRO1125) (Fwd: 5′-AGAACACTATCAATGACCCCAAC-3′; Rev: 5′-CCATGCCCTGGTACATCT-3′, annealing temperature 50 °C, extension 15 s) genes for 30 cycles using OneTaq DNA Polymerase.
For the EATRO1125 RAD51 and BRCA2 knockout experiments, parasites were stored in RNAlater and shipped on dry ice. To extract RNA, parasites were spun at 2.6g for 4 min at 4 °C. RNAlater was removed and 1 ml of TRIzol was added. Some samples did not have a tight cell pellet and thus were spun multiple times and the RNAlater supernatant was carefully removed incrementally following each spin. For some samples, the RNAlater could not be removed entirely. These samples were extracted twice with TRIzol. The RNA pellet mixed with remaining RNAlater at the end of the isopropanol precipitation was resuspended in 1 ml of TRIzol and the extraction was repeated for clean RNA isolation.
VSG-seq
VSG-seq was performed as previously described1,9. Libraries were sequenced with 100-bp single-end reads on a NovaSeq6000. Analysis was performed using the VSGSeqPipeline found at github.com/mugnierlab.
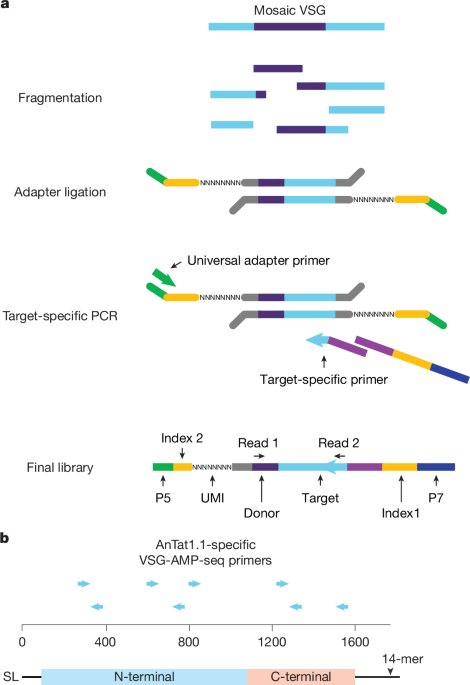
VSG-AMP-seq library preparation and analysis
VSG-AMP-seq was based upon anchored multiplex PCR (AMP)-seq78 and genome-wide, unbiased identification of DNA double-stranded breaks enabled by sequencing (GUIDE-seq)59. Detailed descriptions of the library prep and analysis pipeline, including all software dependencies79,80,81, can be found in the Supplementary Methods.
Identification of mosaic VSG from ORFs
ORFs from ref. 9 were tiled into 20-bp k-mers overlapping by 1 bp each. All tiles were aligned to AnTat1.1 using bowtie82 with the following flags: -v 2. ORFs in which 15 or more tiles successfully aligned to the AnTat1.1 sequence were identified as possible mosaics (≥35-bp match, potentially discontinuous). AnTat1.1 sequences containing ORFs were assessed for sequence matching common donor VSGs: VSG-228, VSG-2986, VSG-3110 and VSG-7358. If a best match could not be determined or if recombination did not occur with one of the typical donor VSGs, VSG sequences were analysed by hand to identify remaining mosaic recombination events.
VSG clustering and family identification
VSGs were identified from published T. brucei genomes6,27,83,84 and expressed VSGs9. VSGs were clustered into family groups using either BLASTn77 or UCLUST algorithms85. Further details, including further software dependencies86,87, can be found in the Supplementary Methods.
Western blotting
EATRO1125 parasites with pLEW-FLAG-La-Cas9 and pLEW-T7sgRNA were grown for 24 h in the presence of 1 μg ml−1 doxycycline or DMSO vehicle control. After induction, 5 million parasites were collected and washed with 25 °C PBS. Then, pelleted parasites were resuspended in 50 μl of RIPA buffer (50 mM Tris, 150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS, pH 7.4) + 2 × Laemmli buffer. Lysed parasites were boiled at 95 °C for 5 min. Then, 5 μl of lysates were separated on a Tris-glycine polyacrylamide gel at 110 V for 100 min in Tris/glycine running buffer (25 mM Tris, 192 mM glycine, 0.1% SDS, pH 8.3). Proteins were then transferred onto a nitrocellulose membrane using transfer buffer (25 mM Tris, 192 mM glycine, 20% methanol) overnight at a 60-mA per transfer box at 25 °C. For FLAG (1:1,000) and EF1α (1:1,000), lysates were separated on 10% polyacrylamide gels and transferred onto 0.45-µm membranes. For γ-H2A (1:200), lysates were separated on a 15% polyacrylamide gel and transferred onto a 0.2-µm membrane. After transfer, blots were blocked with 0.5% BSA in TBS with 0.05% Tween 20 (TBST, 20 mM Tris, 150 mM NaCl, pH 7.6) for an hour at room temperature under constant agitation. Blots were incubated with primary antibody (see above dilutions) for 2 h at 25 °C. After five TBST washes, blots were stained for 1 h with goat anti-mouse (1:5,000, Cell Signaling, 7076S) or goat anti-rabbit-HRP-conjugated secondary (1:5,000, Cell Signaling, 7074S). After another five washes, blots were incubated with ECL (Cytiva, RPN2109) and film was exposed to blots in a dark room. Developed film was scanned and images were processed with FIJI (v.2.14.0/1.54.f)88. Primary antibodies used were mouse anti-FLAG (M2 clone) (Millipore Sigma, F3165-1MG), mouse anti-EF1α (CBP-KK1 clone) (Millipore Sigma, 05-235) and rabbit anti-γ-H2A, a kind gift from G. Hovel-Miner based upon ref. 89. Full uncropped images can be found in Supplementary Fig. 3.
Death assays and clonal outcome analysis
Multiple parasite clones detailed in ‘Isolation of mosaic-expressing parasites’ using the pLEW-T7-sgRNA with guides at cut sites 243, 694 or 1459 were replica plated in 96-well plates in 5 μg ml−1 blasticidin, 0.1 μg ml−1 puromycin and 1 μg ml−1 doxycycline or an equal volume of DMSO at multiple parasite dilutions. Uninduced replica plates had at most 76 colonies. Induced replica plates had at most 13 colonies. Individual clones were counted after 14 days. Upon isolation of a parasite clone, all drugs were removed. A subset of parasites were analysed to determine the VSG expressed.
VSG sequences for clones were determined as above in ‘VSG PCR and identification via sequencing’. Consensus.ab1 sequencing files were assessed for their clonality (Clonal: high agreement at all bases, versus Polyclonal Colony: disagreement at multiple base locations).
Tb427VSG-8 mosaic detection in vivo
RNA from day 6, day 13, day 15 and day 18 mouse blood was extracted and DNase treated as above. cDNA was synthesized using the Superscript III Reverse Transcriptase and a VSG-specific primer that binds to a conserved 14-bp sequence within the 3′ UTR (5′-GTGTTAAAATATATC-3′). cDNA was treated with Rnase A and RNase H for 30 min then purified with 1.8X Mag-Bind Total NGS Beads. Purified cDNA was amplified for 25 cycles with VSG-specific primers: a spliced-leader (5′-ACAGTTTCTGTACTATATTG-3′) and a sample-specific barcoded SP6-VSG 14-mer (or one without the barcode, see VSG barcoding primers, Supplementary Table 1), using Phusion polymerase (annealing temperature 55 °C, extension 45 s). This PCR product was purified with a 1.8X bead cleanup.
Samples were pooled, prepared with Ligation sequencing DNA V14 kit (SQK-LSK114) then sequenced on an Oxford Nanopore Technology PromethION with a 10.4.1 flow cell yielding approximately 100.29 Gb of data with an average length of 1.67 kb.
Tb427VSG-8 and EATRO1125VSG-73 in vitro and in vivo mosaic identification and analysis
Identification of mosaic VSGs from mixed in vitro and in vivo samples, including software dependencies77,79,90,91,92,93, is detailed in the Supplementary Methods.
Mouse AnTat1.1 antiserum generation
Four 8–12-week-old female C57BL/6 mice were infected with about 100 Lister427 SM-AnTat1.1 transgenic parasites via intravenous injection in the tail vein. Parasites appeared in mice 4–5 days after infection. On day 5 after infection, mice were treated with 2 μg of Berenil (Cayman Chemical, 18678) in PBS via intraperitoneal injection to cure infection. A second dose of Berenil was repeated 24 h later. At 14 days after infection, mice were humanely euthanized and blood was collected via cardiac puncture. Serum was isolated using microtainer gel tubes (BD, 365967), spun at 6,000g for 3 min and stored at −80 °C until use.
Flow cytometry
In 96-well plates, 200,000 parasites were stained with 1:20,000 rabbit anti-AnTat1.1 primary antibody36 (J. Bangs) or 1:1,000 mouse anti-AnTat1.1 serum for 10 min at 4 °C while shaking in PBS + 10 mg ml−1 glucose. Parasites were washed once with 100 μl of PBS + glucose. Then, parasites were stained with Alexa Fluor 647-conjugated goat anti-rabbit IgG (H+L), F(ab’)2 Fragment (Cell Signaling, 4414S) or goat anti-mouse IgG (H+L), F(ab’)2 Fragment (Cell Signaling, 4410S), respectively, at 1:1,000 at 4 °C while shaking in PBS + glucose. After washing again with 100 μl of PBS + glucose, parasites were resuspended in PBS + glucose + 1:20 propidium iodide (BD Biosciences, 556463) and analysed on an Attune Nxt flow cytometer (Invitrogen). Data analysis was performed using FlowJo (v.10.8.1). A gating strategy is provided in Supplementary Fig. 2.
Analysis and modelling of VSG N-terminal domains
Full-length protein coding sequences from AnTat1.1 and its isolated mosaics were used for structural modelling. Only N-terminal domain sequences were used for protein structural prediction as this region of the VSG is the most well-defined experimentally. Signal peptides are cleaved from the mature VSG during processing, so we used SignalP 6.0 (ref. 94) to predict and remove the Sec/SPI sequence (–organism eukarya, –mode fast), resulting in a FASTA file of mature proteins. To determine the coordinate of the N-terminal domain, we used a Python analysis pipeline available at https://github.com/mugnierlab/find_VSG_Ndomains. The script identifies the boundaries of the VSG N-terminal domain using the HMMscan function under HMMer v.3.1b2 (ref. 95). Query sequences are searched against an HMM profile containing 735 known N-terminal domain sequences from ref. 6 and N-terminal domains defined by the largest envelope domain coordinate that meets the E value threshold (1 × 10−5, -domE 0.00001). The processed FASTA file containing only mature VSG N-terminal domain sequences was used as input for structural prediction of monomers using LocalColabFold (v.1.5.5)96 function colabfold_batch, run using the following arguments: –amber, –templates, –num_recycle 3. The best-ranked output model with the highest average predicted local distance test score (pLDDT), that is, the highest-confidence model, was used for downstream analyses. Model visualization and alignment were performed using UCSF ChimeraX v.1.7.1 (ref. 97), developed by the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco, with support from the NIH (grant no. R01-GM129325) and the Office of Cyber Infrastructure and Computational Biology, NIAID.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.