Experimental model and participant details
Human participants
All of the deidentified samples used in this study were obtained after informed consent of patients and/or their legal representatives who did not receive compensation. The study was approved by the institutional review board in agreement with local institutional ethics guidelines DFCI 10-417 (Boston Children’s Hospital and Dana-Farber Cancer Institute), S-531/2020 (Heidelberg University) and EK no. 1244/2016 (Medical University of Vienna). Cohort characteristics are provided in Supplementary Table 1.
ZFTA-RELA patient-derived cell lines
EP1NS, BT165 and VBT242 cell lines were generated as previously described47,48 and provided by collaboration partners. Cells were grown in complete neurobasal medium, comprising Neurobasal-A Medium (Thermo Fisher Scientific, 10888022) supplemented with 1× of antibiotic–antimycotic 100× (Thermo Fisher Scientific, 15240062), 1× GlutaMAX (Thermo Fisher Scientific, 35050061), 1 mg ml−1 heparin solution (StemCell Technologies, 7980), 1× B-27 (Thermo Fisher Scientific, 12587010), 20 ng ml−1 recombinant human FGF-basic (Shenandoah Biotech, 100-146) and 20 ng ml−1 recombinant human EGF (Shenandoah Biotech, 100-26). For further passaging, floating cells were centrifuged at 300g for 5 min, dissociated with Accutase (Innovative Cell Technologies, AT104-500) for 5 min at 37 °C and washed with PBS (Gibco, 10010023). All of the cell lines used were authenticated by single-nucleotide polymorphism analysis and cells were regularly tested for mycoplasma contamination.
ZFTA-RELA PDXs
EP1NS, BT165 and VBT242 cells (500,000 cells in 3 μl PBS per mouse) were injected stereotactically into the cortex of 6-week-old female NSG mice (NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ, The Jackson Laboratory, 005557), which were treated with 0.05 mg per kg buprenorphine and anaesthetized with 2% to 3% isoflurane. The skull of the mouse was exposed through a small skin incision, and a small burr hole was made using a 25-gauge needle at the selected stereotactic coordinates: 1.0 mm x, 1 mm y and −1.5 mm z. After injection, mice were checked daily for signs of distress, including seizures, weight loss or tremors. Tumour size was monitored monthly by small animal MRI and BLI starting 4 weeks after injection. Mice were euthanized as they developed neurologic symptoms, including head tilt, seizures, sudden weight loss, loss of balance and/or ataxia according to IACUC defined endpoints for central nervous system tumours. No mice exceeded these endpoints. Sample sizes were determined with reference to previous studies to provide sufficient statistical power for detecting biologically meaningful differences. Mice were randomized to experimental groups at the start of each experiment when possible. Blinding was not performed during data collection, but analyses were conducted according to predefined criteria to reduce bias. All animal studies were performed according to protocols approved by the Dana Farber Cancer Institute Institutional Animal Care and Use Committee (IACUC; 13-053).
Molecular classification of tumours
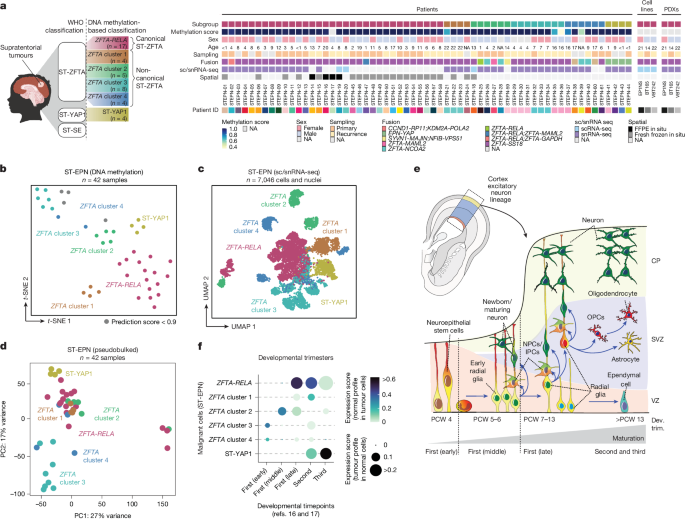
Molecular classification through DNA-methylation profiling was performed using the Heidelberg Brain Tumour Methylation Classifier v.12.4 (https://app.epignostix.com/) as previously described49. Genome-wide DNA methylation profiling was performed using the Infinium HumanMethylation 450k or EPIC Kit according to the manufacturer’s instructions (Illumina). Raw signal intensities from the resulting .idat files were calculated using the minfi Bioconductor package v.1.24.0. The normalization and batch effect correction were performed using the limma package v.3.34.5. Afterwards, β values were calculated before selecting CpG probes for downstream analysis. Unsupervised nonlinear dimensionality reduction was performed using the 1 − variance-weighted Pearson correlation between the samples to construct the distant matrix, which was subsequently used as an input for a t-distributed stochastic neighbour embedding visualization. Classification was performed using the established random-forest algorithm of the Heidelberg Brain Tumour Methylation Classifier v.12.4, which included all molecular EPN groups, including the newly established ST-EPN clusters derived in previous studies13,14.
Detection of fusion genes in tumours
Gene fusion status for most cases was derived from earlier studies13,14 using either RNA-seq or PCR with reverse transcription as described in the respective studies. The gene fusions and their respective isoforms were derived using two independent methods: InFusion and Arriba50,51.
In vitro co-culture
As described previously15, primary cultures of rat cortical neurons and astrocytes were prepared from E19 embryos. They were plated at a density of 90,000 cells per cm2 on 12 mm diameter coverslips on 24-well plates that were previously coated with poly-l-lysine. The cells were maintained in neurobasal medium supplemented with B27 supplement (50×, 2% v/v) and l-glutamine (0.5 mM). After 7 days, Accutase-dissociated cells from patient-derived ZFTA-RELA cell lines were added and co-cultured (1,000 cells per well). For time-lapse live-cell imaging or immunohistochemistry, coverslips were used 4–14 days in vitro after seeding ZFTA-RELA cells. As a control, in vitro monocultures of ZFTA-RELA cells were established from patient-derived ZFTA-RELA cell lines as follows: 12-mm-diameter coverslips on 24-well plates were coated using poly-l-lysine and maintained in neurobasal medium supplemented with B27 supplement (50x, 2%, v/v) and l-glutamine (0.5 mM). After 7 days, Accutase-dissociated ZFTA-RELA cells were added and cultured (1,000–10,000 cells per well). For time-lapse live-cell imaging immunohistochemistry, coverslips were used 4–14 days in vitro after seeding ZFTA-RELA cells.
Generation of human iPS-cell-derived neurons
iPS cell lines were obtained from the Boston Children’s Hospital Human Neuron Core, with all appropriate permissions for use. Induced neurons were generated from the GON0515-03 iPS cell line according to established protocols52. In brief, iPS cells were plated at a density of 95,000 cells per cm2 on Geltrex-coated plates in mTeSR1 medium (StemCell Technologies, 85850) for viral transduction. The following lentiviral plasmids were obtained from Addgene: FUdeltaGW-rtTA (Addgene, 19780), a gift from K. Hochedlinger; pTet-O-Ngn2-puro (Addgene, 52047) and Tet-O-FUW-eGFP (Addgene, 30130), both gifts from M. Wernig. High-titre lentiviruses (~1 × 109 transducing units per ml) were generated by Alstem and used at the following concentrations: 0.13 µl per 50,000 cells for each virus. After transduction, cells were dissociated with Accutase (StemCell Technologies, AT104-500) and replated at a density of 200,000 cells per cm2 on Geltrex-coated plates in mTeSR1 medium supplemented with ROCK inhibitor (10 µM, StemCell Technologies, 72304) (day 0). On day 1, the medium was replaced with N2 medium supplemented with doxycycline (2 µg ml−1, Clontech, NC0424034), brain-derived neurotrophic factor (BDNF; 10 ng µl−1, Peprotech), neurotrophin-3 (NT3; 10 ng µl−1, Peprotech, 450-03) and laminin (0.2 µg µl−1, Life Technologies, 23017015). Doxycycline was maintained throughout the differentiation protocol. On day 2, medium was switched to N2B medium containing BDNF, NT3, laminin (same concentrations), Ara-C (2 µM, Sigma-Aldrich, 1768-100MG) and puromycin (1 µg ml−1, Gibco, A11138-03). From day 3 onward, cells were cultured in N2B medium supplemented with 1× B27 (Life Technologies, 17504044), doxycycline (2 µg ml−1), puromycin (5 µg ml−1), BDNF, NT3 and laminin, with half-medium changes performed every other day. At day 6, puromycin was withdrawn and cells were enzymatically dissociated using the Papain Dissociation System (Worthington Biochemical, LK003150) for downstream experimental setups. Induced neurons were plated on poly-d-lysine and laminin-coated six-well plates or 15 mm glass coverslips in co-culture with iPS-cell-derived human astrocytes (Ncardia, M0605), which were added at 10–15% of the total neuron cell density.
Culture of human iPS-cell-derived astrocytes
Human iPS-cell-derived Ncyte Astrocytes (Ncardia, M0605) were cultured in T75 flasks (Thermo Fisher Scientific) precoated with poly-d-lysine (R&D Systems, 3439-200-01). Cells were maintained in Astrocyte Medium (ScienCell, 1801), consisting of 500 ml of basal medium (1801), 10 ml of FBS (0010), 5 ml of astrocyte growth supplement (AGS, 1852) and 5 ml of penicillin–streptomycin solution (0503). Half-medium changes were performed every other day. Before co-culture, astrocytes were enzymatically dissociated using Accutase (StemCell Technologies) and subsequently plated together with day 6 iPS-cell-derived neurons. Astrocytes were added at a final density corresponding to 10–15% of the total neuronal cell count and maintained in neuronal medium.
In vitro coculture of ZFTA-RELA cells with human iPS-cell-derived neurons
On day 18 of human iPS-cell-derived neurons differentiation, ZFTA-RELA tumour cells were added to neuronal cultures at a ratio of 1:3 (tumour:neuron). Cocultures were monitored for 7 days and maintained with half-medium changes every other day with B27 medium, doxycycline (2 µg ml−1), puromycin (5 µg ml−1), BDNF (10 ng µl−1), NT3 (10 ng µl−1) and laminin (0.2 ng µl−1). For all scRNA-seq experiments, cocultures were dissociated using the Papain Dissociation System after the 7-day coculture period. To assess the role of neuronal activity on tumour cell behaviour, cocultures were treated with either tetrodotoxin (1 µM, Tocris, 1069) or vehicle control.
Fresh tumour processing
Live cells were isolated from immediately processed fresh tumour tissue acquired at the time of surgery. Fresh tumour tissue was mechanically dissociated and enzymatically digested using the Brain Tumour Dissociation Kit (Miltenyi Biotec, 130-095-942) and gentleMACS dissociator (Miltenyi Biotec, 130-096-427) for 30 min at 37 °C. Single-cell suspensions were filtered through a 70 µm strainer (Thermo Fisher Scientific, 03-421-228), centrifuged at 500g for 5 min and resuspended in a solution of 1% BSA (BioLegend, 644710) in PBS for fluorescence-activated cell sorting (FACS).
Frozen tumour processing
Nuclei for snRNA-seq were extracted from snap-frozen and optimal cutting temperature (OCT)-embedded tumour tissues. Snap-frozen tumour tissue was cut on ice, while OCT-embedded tissue was cut on a cryostat at −20 °C and washed with PBS. Frozen tumour tissue was suspended in cell lysis buffer (CST) and dissociated with sterile surgical scissors for 5 min until homogenous. Dissociated tissue was filtered through a 70 µm strainer, 1XST was added to the nucleus solution and once more transferred through a 70 µm strainer. The nucleus solution was centrifuged at 500g for 5 min. Single-nucleus suspensions were resuspended in PBS supplemented with 1% BSA (Smart-seq2) or 0.05% BSA (10x Genomics). All steps were performed at 4 °C.
Live-cell sorting
A single-cell suspension of 100 μl (kept in a FACS-tube in PBS/BSA at 4 °C) was used for all sorting procedures as an unstained control. For stained controls, 0.75 μl Calcein AM (Life technologies, C34851) and 0.5 μl TO-PRO3 Iodide (Life technologies, T3605) were added to resuspended cells in 100 μl PBS/1% BSA for 10 min at room temperature. Single-cell sorting was performed on the MA900 sorter (Sony). Single viable tumour cells were selected by positive staining for Calcein AM as well as negative staining for TO-PRO3 and sorted into pre-chilled (4 °C) 96-well plates containing TCL buffer (Qiagen, 1031576) + 1% β-mercaptoethanol (Thermo Fisher Scientific, 21985023). Sorted plates were centrifuged at 1,000g for 1 min at 4 °C and frozen on dry ice followed by transfer to a −80 °C freezer for long-term storage before whole-transcriptome amplification, library preparation and sequencing.
Nucleus sorting
Dissociated frozen tumour tissue derived single-nucleus suspensions were resuspended in PBS 1% BSA and stained with 2.5 mM Vybrant DyeCycle Ruby Stain (Life Technology, V10309). Nucleus sorting was performed on the MA900 sorter (Sony). Intact nuclei were sorted into prechilled (4 °C) 96-well plates containing TCL buffer + 1% β-mercaptoethanol. Sorted plates were centrifuged at 1,000g for 1 min at 4 °C and frozen on dry ice followed by transfer to a −80 °C freezer for long-term storage before whole-transcriptome amplification, library preparation and sequencing.
Sorting of cells grown in cocultures
Day 8 in vitro coverslips of mono- and co-culture were incubated with trypsin for 5 min. Next, 10% FBS was added, and cells were washed from coverslips. The samples were stained with a DAPI as a dead cell marker to identify the live-cell population. The single-cell suspension was sorted with a FACSAria Fusion 2 (BD Biosciences). Tumour cells were identified by their tdTomato expression.
Full-length sc/snRNA-seq
Smart-seq2 whole-transcriptome amplification, library preparation and sequencing of single cells/nuclei were performed according to the modified Smart-seq2 protocol as previously described1,53,54. RNA was purified with RNAClean XP beads (Beckman Coulter, A66514). Next Oligo-dT primed reverse transcription was performed using Maxima H Minus reverse transcriptase (Life Technologies, EP0753) and a template-switching oligonucleotide (TSO; Qiagen) followed by PCR amplification (20 cycles for scRNA-seq and 22 cycles for snRNA-seq) using the KAPA HiFi HotStart ReadyMix (KAPA Biosystems, 07958935001), and by AMPure XP bead (Beckman Coulter, A63882) purification. Libraries were generated using the Nextera XT Library Prep kit (Illumina, FC-131-1096). Libraries from 768 cells with unique barcodes were combined and sequenced using the NextSeq 500/550 High Output Kit v2.5 (Illumina, 20024906) on the NextSeq 500 sequencer (Illumina).
scRNA-seq data processing
Quality filtering
We aligned raw sequencing reads to hg19 genome by hisat2 (v.2.1.0) and quantified gene counts using RSEM (v.1.3.0) as raw counts. For Smart-seq2 data of frozen tumours, we excluded cells with genes <1,000 and an alignment rate of <0.1; for fresh tumours, the filtering threshold was <2,500 genes and an alignment rate of <0.2. We also removed genes with TPM >16 in <10 cells.
Normalization and scaling
For the remaining good-quality cells and genes, we computed the aggregate expression of each gene as Ea(i) = log2[average (TPMi,1, …, n) + 1] and defined relative expression as centred expression levels, Eri,j = Ei,j − average (Ei,1, …, n). In total, 7,873 high-quality cells were retained frozen tumours, and 1,093 for fresh tumours for downstream analysis. On average, we detected 3,660 unique genes per cell in frozen tumours, and 5,815 unique genes per cell in fresh tumours.
Identification of malignant and non-malignant cells
To discriminate between malignant and non-malignant cells, we used (1) Seurat clustering; (2) unbiased cell type annotation using the automated annotation package SingleR (v.1.6.1) using the Human Primary Cell Atlas55 as a reference; and (3) inference of copy-number variation (CNV) using the InferCNV R package (v.1.8.0). We began by performing an initial annotation of cell lineages through dimensionality reduction using UMAP and unsupervised Louvain clustering. To refine our classifications, we analysed clustering patterns and the expression of established immune, endothelial and oligodendrocyte gene markers. We validated these initial lineage assignments using the automated annotation package SingleR, which assigns cell identities on the basis of their similarity to a reference set of bulk transcriptomes from healthy cells (Human Primary Cell Atlas data55). We then inferred genome-wide evidence for somatic chromosomal CNV using the InferCNV R package as previously described1,53,54. We spiked-in non-malignant immune and oligodendrocyte nuclei from previous publications1,53,54 as reference normal karyotype for CNV inference, and used hierarchical clustering of the single-cell copy-number profiles within each sample with the reference non-malignant copy-number profiles to classify the presence of CNVs. Owing to the presence of false-positive/false-negative rates of inferring CNVs with nuc-seq data from frozen samples, CNV results were used as secondary validations. CNVs should be detected in cell clusters classified as malignant cells by transcriptional clustering. By contrast, CNVs should be absent from cell clusters classified as a normal cell type. Reassuringly, we observed that nuclei lacking CNVs typically clustered together and were identified as macrophage, T cell or endothelial cell nuclei based on their high expression of myeloid, T cell or endothelial markers, respectively. All other clusters consistently exhibited CNVs and were classified as malignant tumour cells. Only cells identified as malignant were selected for subsequent NMF analysis.
Defining NMF programs and metaprograms
To identify heterogeneous transcriptional programs within each sample, we performed NMF analysis using frozen tumours profiled by snRNA-seq. As input for NMF, we selected the top 10,000 overdispersed genes using PAGODA2 (v.0.1.4) identified on the malignant cell compartment of each sample separately. We set the negative values to zero and ran NMF package (v.0.28) using rank = 6. To identify metaprograms (MPs), we first scored all malignant cells for the top 30 genes with the highest score from each NMF factor as explained in the ‘Scoring gene sets’ section. We next clustered the resulting scores across all NMF programs by hierarchical clustering as explained in the ‘Program-wise hierarchical clustering’ section and used the function cutree from the stats R package (v.4.4.2) to automatically extract n = 15–20 highly correlated MPs. We finally curated the resulting clusters manually, by (1) removing clusters that were associated with limited samples (and therefore considered to not to reflect relevant general biology of the tumours); (2) removing clusters that were not associated with any significant gene enrichment ontology term; (3) grouping together clusters that reflected similar underlying biological processes. This approach revealed eight highly correlated sets of programs or MPs within ST-EPN tumours.
Annotation of MPs
To annotate MPs, we used three approaches. First, we performed GO analysis to identify over-represented biological processes in each metaprogram as described in section: GO and gene set enrichment (GSEA) analysis. As an input, we used the top 30 marker genes from each MP. Second, we compared the MPs identified here de novo with MPs that we identified in previous work15. Third, we compared the gene signatures of each MP to a developmental reference scRNA-seq dataset of non-malignant cortical brain development31,32 as described in the ‘Generation of developmental signatures’ section. Assessment of significant changes in metaprogram proportion were made using the propeller test from the Speckle R package (v.1.8.0)56.
Generation of single-cell expression scores
Scoring gene sets
We first aggregated ST-EPN single cells/nuclei from all samples, and performed normalization and centring. We then scored these cells for NMF/developmental signatures as previously described1,57,58. We did this by calculating for each cell i a score SCj(i), quantifying the average relative expression (Er) of the top 30 genes within each MP (Gj) and comparing the score to the average relative expression of a control gene (Gjcont): SCj(i) = average[Er(Gj,i)] − average[Er(Gjcont,i)]. The control gene set was defined by binning all genes into 30 bins of aggregate expression levels (Ea) and randomly selecting 100 genes from the same expression bin for each gene in the gene-set Gj. ST-EPN cells/nuclei were annotated based on the maximum expression score for the respective MP signatures after excluding the one related to cell cycle (cycling).
Cycling scores
We used Seurat’s CellCycleScoring function to assign cell cycle scores. This function relies on gene signatures that have been previously shown to characterize S and G2/M cell cycle phases. We defined high-cycling cells as cells with S scores or G2/M scores > 0, and low-cycling cells as cells with S scores < 0 and G2/M scores < 0.
Generation of developmental signatures
To identify developmental signatures enriched in ST-EPN subtypes or cell states, we performed projection of tumour cells (pseudobulked by ST-EPN subtype or cell state) onto normal cell type signatures. We first assembled a single-cell reference atlas of the developing human cerebral cortex from two published scRNA-seq datasets31,32. We then subset the combined dataset to 500 cells (if available) per cell identity and used Seurat’s RunPrestoAll function (retaining only genes expressed in at least 25% of all cells) to derive the top differentially expressed genes (adjusted P < 0.05) across cell types, which are reflective of developmental-specific cell type signatures. We then scored all ST-EPN malignant cells with these marker genes and annotated ST-EPN cells based on the maximum expression score for the respective signatures.
Program-wise hierarchical clustering
To compare sets of NMF programs or MPs, we first scored all ST-EPN cells/nuclei for the program signatures as described in the ‘Scoring gene sets’ section. After scoring, we performed pairwise correlation for each possible pair of programs and show hierarchical clustering of the scores using a correlation-based distance metric (distance metric: 1 − Pearson correlation; linkage: Ward’s linkage).
Cell state plot
We separated ST-EPN cells into neuroepithelial-like/embryonic-like versus neuronal-like/ependymal-like as previously described58. In brief, we first calculated the y axis value as y = max(SCneuroepithelial-like,SCembryonic-like) − max(SCneuronal-like,SCependymal-like). For neuroepithelial-like/embryonic-like cells (y > 0), the x axis value was defined as x = log2(|SCependymal-like − SCneuronal-like| + 1); and, for neuronal-like/ependymal-like cells (y < 0), the x axis was defined as log2(|SCembryonic-like − SCneuroepithelial-like|).
Differential gene expression analysis
Single-cell analysis
To identify genes that are differentially expressed between two conditions (monoculture versus coculture) or cell states (neuroepithelial-like versus embryonic-like), we first used Seurat’s RunPrestoAll function (retaining only genes minimally detected in 15% cells). For downstream GO analysis we used the top 30 genes (ordered by average log2[FC]) with an average |log2[FC]| > 1 and adjusted P-value threshold as indicated in the corresponding figure legend; for GSEA, we used all genes with and an adjusted P-value threshold of 10−5.
Pseudobulk analysis
We performed pseudobulk differential expression analysis to identify transcriptional programs that distinguish ZFTA-RELA from ZFTA cluster 3 tumours. We first extracted raw count matrices from each tumour sample, and calculated pseudobulked gene expression counts using the SingleCellExperiment (v.1.28.1) aggregateAcrossCells function. We then conduceted differential gene expression analysis using edgeR (v.0.27). We filtered out genes with low expression counts using the filterByExpr function, based on our grouping variable (tumour subtype). We then normalized the counts using the calcNormFactors function. We constructed a design matrix to include tumour subtype, estimated negative binomial dispersions using the estimateDisp function and fitted a generalized linear model using glmFit, followed by hypothesis testing with glmLRT for differential expression. We considered significant genes based on a false-discovery rate (FDR) threshold of <0.001. For downstream GO analysis, we used the top 30 genes (ordered by log2[FC]) with FDR < 0.001 and |log2[FC]| > 1; for GSEA, we used all genes with FDR < 0.001. We visualized results using ggplot2 (v.3.5.0).
GO and GSEA analysis
To identify biological terms that are over-represented in a specific gene set, we performed GO and GSEA analysis using the enrichGO and gseGO functions, respectively, from the clusterProfiler package (v.4.6.2)59. As a database, we used the R package org.Hs.eg.db (v.3.18.0), which is an organism annotation package for Homo sapiens, relying on the Ensembl version released on 10 May 2023. Unless otherwise stated, we used the following parameters: ont = “ALL”, universe = all genes retained for differential gene expression analysis, minGSSize = 3, maxGSSize = 800, pvalueCutoff = 0.05, OrgDb = org.Hs.eg.db, pAdjustMethod = “BH”.
Validation of developmental signatures across bulk RNA-seq ST-EPNs
To validate the preferential mapping of ST-YAP1 versus ZFTA-RELA tumours onto ependymal and neuronal cells, respectively, we downloaded a microarray dataset2 available on the R2 platform (r2.aml.nl; R2 internal identifier: ps_avgpres_gse64415geo209_u133p2) containing normalized gene expression matrix (z score) data from n = 49 ZFTA-RELA and n = 11 ST-YAP1 patient tumour samples. After removing n = 10 ZFTA-RELA and n = 2 ST-YAP1 samples (as they were present in our discovery snRNA-seq cohort), we scored samples using Seurat’s AddModuleScore function, taking the NMF-derived gene signatures described above as the input.
Spatial transcriptomics with 10x Xenium
Gene panel design
For in situ 10x Xenium analysis, a custom gene panel of 100 genes was designed to include metaprogram marker genes identified across different types of high-grade gliomas, including: (1) ZFTA-RELA patient tumours analysed in this study (n = 35 genes); (2) diffuse midline gliomas35 (n = 10 genes); (3) glioblastoma37 (n = 27 genes); (4) genes that mark the same metaprogram in a combination of tumour types (intersection, n = 10 genes); and (5) normal cell types (n = 18) (Supplementary Table 6). For ZFTA-RELA tumours, those genes encompassed the following metaprograms: cycling (n = 4 genes), neuroepithelial-like (n = 8 genes), radial glial-like (n = 3 genes), NPC-like (n = 12 genes), ependymal-like (n = 8 genes) and mesenchymal-like (n = 4 genes); and three probes targeting ZFTA-RELA fusion transcripts. Moreover, we used the human brain predesigned panel containing probes for 272 genes encompassing additional non-malignant cell types (oligodendrocytes, immune cells, endothelial, astrocytes, neurons, microglia and VLMCs).
Fusion probe design
For detection of fusion transcripts, custom probes were designed for three different fusion sites between the exons of ZFTA and RELA (Supplementary Table 6). The 60 bp area surrounding the fusion site was used to derive junction-specific probes.
Sample preparation
For fresh-frozen samples, tissue sections were cut at a thickness of 10 µm using the Leica CM3050S cryostat and collected onto Fisherbrand Superfrost Plus microscope slides, followed by fixation and permeabilization. For FFPE blocks, 5-µm-thick tissue sections on a Xenium slide underwent deparaffinization and permeabilization. Hybridization of a mix of predesigned and custom gene expression probes was performed at 50 °C overnight. After multiple washes to remove unhybridized probes, ligation of probes and annealing of rolling circle amplification primer was performed at 37 °C for 2 h. Circularized probes were then enzymatically amplified at 30 °C for 2 h, after which background autofluorescence was quenched chemically. The slides were then loaded into the Xenium Analyzer after nucleus staining.
Xenium in situ imaging
Image acquisition and sample handling was automated within Xenium Analyzer for two slides per run. Fifteen rounds of fluorescence probe hybridization, imaging and probe removal occurred within the Analyzer. A fast area scan camera with ~200 nm per pixel resolution was used for image acquisition, and z stacks were taken with 0.75 μm step size across tissue thickness. All z-stack of images were then processed and stitched, using DAPI image as reference. Each fluorescently labelled oligonucleotide bound to amplified barcodes was detected and registered in each cycle. Unique optical signature from fluorescence intensity over the 15 rounds was used to identify a target gene. For cell segmentation, neural network was used to detect each nucleus from DAPI images.
Xenium preprocessing of data
The preprocessing of Xenium platform data was conducted using the Seurat pipeline. Initially, the per-transcript location data, cell × gene matrix, cell segmentation and cell centroid information provided in the Xenium outputs were imported using the LoadXenium function. Subsequently, SCTransform was used for normalization, followed by standard procedures for dimensionality reduction and clustering.
Xenium cell state/type cell assignment
To assign cell identities in Xenium data, we first performed unsupervised clustering using Seurat (v.5.0.2), performing RunUMAP on the least number of PCs capturing 80% or more variance, shared-nearest-neighbour graph using FindNeighbors and Louvain clustering (FindClusters) run at resolutions 0.1 to 1.0, in 0.1 increments. We then scored only for the normal cell types, by transferring labels from the single cell/nucleus reference object from corresponding tumour subgroup using FindTransferAnchors and TransferData (dims = 20), yielding a label-transfer score for every cell-to-cell type pair. Independent module scores were calculated using AddModuleScore_UCell in the R package UCell (v.2.8.0) using marker genes present in the Xenium panel. For each cluster, the average label-transfer score and module score were manually inspected and visualized through UMAP, and the clusters corresponding to normal cell types were identified in each sample. Cells belonging to the normal cell type annotation were removed, and the malignant subset of each sample was annotated in the same way as normal cell types. To mitigate systemic score inflation, scores for each malignant cell type were mean-centred across all cells: \(\rmc\rme\rmn\rmt\rmr\rme\rmd\,\rms\rmc\rmo\rmr\rme_ij=\rms\rmc\rmo\rmr\rme_ij-\bar\rms\rmc\rmo\rmr\rme_j\), where i corresponds to cells and j corresponds to cell types. Finally, each malignant cell was assigned with the cell type with highest mean centred score, and malignant and normal cells were combined to generate the final, fully annotated cells.
Spatial coherence score
We defined the spatial coherence score, a measure of how structure or disorganized a tumour is, as follows. After annotating each cell with a malignant metaprogram or non-malignant cell type, the data were divided into 200 bins per row and column, totalling 40,000 spots per tumour section, each of which was analysed to identify the cells that it contained. To achieve this, a spot annotation program was developed based on the program with the highest proportion within that spot. To determine whether a sample is organized or disorganized, a coherence score was calculated by examining each spot and its surrounding 3 × 3 square neighbours. If the neighbours were found to be like the spot annotation, a score of 1 was added; otherwise, a score of 0 was assigned. This process was carried out separately for each sample and metaprogram. The scores for each program were combined and divided by the number of spots associated with that program. The final coherence score for each sample was calculated by averaging all of the scores calculated for each program across that sample and scaled within 0–1.
Linear regression
We used simple and multiple linear regression to assess whether the abundance of cell state/type is associated with increased spatial coherence as well as to evaluate the concordance between cell state/type abundance identified by sn/scRNA-seq and spatial Xenium analyses. We performed logit transformation of cell state/type proportion using the logit function from R package car (v.3.1.3). To make the proportion values 0 and 1 not be infinite values, we applied a small adjustment of 0.025 (default for logit function of car) before value transformation. After data transformation, we used the lm function from the R stats package (v.4.4.2) to perform simple and multiple linear regression. We extracted the goodness of fit (R, R2 and P) using the summary() function. We adjusted P values for multiple comparisons using Bonferroni correction.
Spatial niche and metaniche identification
Spatial niche analysis was performed for each tumour section in R using the BuildNicheAssay function from Seurat (v.5.0.2), which demarcates regions of tissue defined by a similar composition of spatially adjacent cell types. This function, which requires a pre-established number of neighbours (neighbours.k set to 20) and expected niches (niches.k set to 6), considers the n = neighbours.k spatially closest neighbours to each cell, counts the occurrences of each cell state/type in this area and uses k-mean clustering to group cells with similar neighbourhoods together to resolve n = niches.k spatial niches for each tumour section. We applied BuildNicheAssay to our spatial cohort using niches.k = 4–6, and thereby resolved 4, 5 and 6, respectively, niches within each tumour section. To identify niches that were consistently detected across multiple tumour sections (termed metaniches), we first calculated the relative proportion of each cell state/type within a given niche section. We then computed correlation matrix between the different niches by hierarchical clustering as explained in the ‘Program-wise hierarchical clustering’ section, to understand how similar the distributions of cell states are across different spatial niches. Finally, we used the function cutree from the stats R package (v.4.4.2) to automatically extract n = 5–6 highly correlated metaniches.
Spatial clusters identification by CellCharter
To simultaneously identify local spatial patterns across multiple samples and provide an orthogonal validation to our niche analysis, we used CellCharter47 (v.0.3.4) to identify spatial clusters. To enable joint analysis among different samples, CellCharter applied dimension reduction and batch correction through a variational autoencoder method, scVI. Next CellChater encoded spatial information into a spatial network based on spatial proximity among cells. To enable neighbourhood aggregation, CellCharter concatenated features from neighbour cells. We then performed GMM-based clustering on these concatenated and aggregated features to determine spatial clusters. We ran GMMs on a wide range of cluster numbers (N) to identify a stable cluster number for the final GMM and downstream analysis. Specifically, N clusters were deemed to be stable if cluster assignments are concordant across multiple runs of GMMs for N, N − 1 and N + 1 based on the Fowlkes–Mallows index.
Shannon entropy of sample proportions of spatial clusters
To evaluate the heterogeneity in abundances of spatial cluster across samples, we computed the Shannon entropy of sample proportions in each spatial cluster. We first calculated the proportions of cells from different samples within each spatial cluster, as pi. Next we calculated a Shannon entropy (H) using this formula \(H=-\sum _\i=1\^np_i\log _2p_i\), where n is the total number of samples present in this spatial cluster.
Experimental procedures
H&E staining
Tissue slides were immersed in haematoxylin for 20 min and washed in distilled water (four times for 1 min). After washing, the slides were immersed in 70% and 95% ethanol for 3 min, followed by eosin staining for 2 min. The slides were further dehydrated in serial ethanol incubation, including 95% ethanol (twice for 30 s) and 100% ethanol (twice for 30 s). Finally, the slides were incubated in xylene for 3 min, and mounted with coverslip to dry for 30 min before proceeding to imaging.
Immunofluorescence analysis of ZFTA-RELA monoculture and coculture cells
Cells were fixed with 4% paraformaldehyde (Thermo Fisher Scientific, J61899.AP) for 15 min at room temperature. Cells were washed with PBS and subsequently permeabilized and blocked with 5% normal goat serum (Thermo Fisher Scientific, 10000C) and 0.2% Triton X-100 (Fisher Scientific, AAA16046AE) in PBS (blocking solution) for 30 min at room temperature. The cells were incubated with primary antibodies overnight at 4 °C in blocking solution. The primary antibodies used in this study were as follows: anti-beta-III-tubulin (1:100; Novus Biologicals, NB100-1612, TUJ88977978), anti-S100B (1:500, Synaptic Systems, 287 044, 1-16), anti-Nestin (1:2500, Invitrogen, PA5-143578, ZD4294645), anti-vimentin (1:1000, CST, 5741T, 8), anti-DCX (1:50, CST, 40619S, 1), anti-CCDC40 (1:100, Thermo Fisher Scientific, PA5-54653, YI40449391B), anti-GFAP (1:500, Aves Lab, GFAP, GFAP857977) and anti-tdTomato (1:80, CST, 20163S, 1). The next day, the cells were washed with PBS three times, incubated with fluorophore-conjugated secondary antibodies at 1:1,000 dilutions (goat anti-rabbit IgG (H+L) cross-adsorbed secondary antibody, Alexa Fluor 488, A-11008; goat anti-mouse IgG (H+L) cross-adsorbed secondary antibody, Alexa Fluor 568; A-11004; goat anti-chicken IgY (H+L) secondary antibody, Alexa Fluor 647; A-21449; goat anti-guinea pig IgG H&L Alexa Fluor 750, ab175758) for 1 h at room temperature, and washed with PBS three more times. Cells were mounted onto SuperFrost Plus microscope slides using ProLong Diamond Antifade Mountant with DNA Stain DAPI (Thermo Fisher Scientific, P36962). Analysis of immunostaining of treated cells was performed using a ×63 oil-immersion objective on a laser-scanning confocal microscope (Zeiss LSM 980 with Airyscan 2) operated with the Zen Microscopy Software (Zeiss). All images were processed using ImageJ software (National Institutes of Health).
Quantification of immunofluorescence of ZFTA-RELA coculture cells
The tdTomato signal of each cell was segmented by setting a pre-set threshold to capture the morphology and area of the cell. Automated colocalization of fluorescence analysis was then performed using Coloc 2 to derive Pearson’s Correlation coefficient for colocalization of the tdTomato signal and the other channels. All of the images were processed using ImageJ software (National Institutes of Health).
10x Xenium analysis of ZFTA-RELA coculture cells
10x Xenium slides were precoated with poly-d-lysine and laminin to facilitate cell adhesion. Rat E19 cortical cells were dissociated and cultured in 10x Xenium slide cassettes (5 × 105 cells per cassette) for 5 days in neurobasal medium supplemented with B27 supplement (50×, 2%, v/v) and l-glutamine (0.5 mM). After 5 days, Accutase-dissociated cells from patient-derived ZFTA-RELA cell lines were added and co-cultured for another 4 days (1–3 × 105 cells per cassette). The slides were then processed using the same 10x Xenium protocol as for patient tissue and loaded into the Xenium Analyzer after nucleus staining. The same slides after processing on the Xenium Analyzer were stained for tdTomato using anti-tdTomato (1:80, CST, 20163S, 1) and fluorophore-conjugated secondary antibodies at 1:1,000 dilution (goat anti-Rabbit IgG (H+L) Alexa Fluor 647, Invitrogen A-21245). Analysis of immunostaining was performed using the Leica Thunder Imager Live Cell and 3D assay with a ×20 objective. All images were processed using ImageJ/Fiji (v.2.14.0)60 software. 10x Xenium explorer (v.3.0) was used to identify and quantify transcripts from Xenium colocalized with tdTomato signals.
In vivo imaging of ZFTA-RELA PDXs
For all in vivo imaging experiments, male NMRI nude mice were used. All animal procedures were performed in accordance with the European Directive on animal experimentation (2010/63/EU) and institutional laboratory animal research guidelines after approval of the Regierungspräsidium Karlsruhe, Germany. Efforts were made to minimize animal suffering and to reduce the number of animals used according to the 3R principles. Mice were clinically scored, and experiments were terminated if they showed marked neurological symptoms or weight loss exceeding 20%.
Surgical procedures
Before in vivo two-photon imaging, cranial window and tumour implantation was performed as described previously18,27,50,52.
Intravital two-photon microscopy
Tumour growth was monitored from 2 weeks after tumour implantation. A Zeiss LSM780 setup equipped with band-pass filter sets of 500–550 nm and 575–610 nm, using a ×20/1.0 NA apochromatic, 1.7-mm working distance, water-immersion objective (Zeiss) was used. A pulsed Ti:Sapphire laser (Chameleon Discovery NX; Coherent) was used at a wavelength of 960 nm.
In vivo 3D time-lapse imaging
Isoflurane gas was diluted in 100% O2 for intravital imaging. For the induction of anaesthesia, the mice were exposed to 4% isoflurane. Isoflurane was lowered to 0.5–2% for the rest of the experiment and was monitored throughout the experiment by checking the breathing rate. Eye cream was applied after anaesthesia induction. During imaging, the body temperature was monitored and kept at 37 °C using a temperature sensor and a heating plate. 100 μl fluorescein isothiocyanate–dextran (2 mg mol−1) was injected into the tail vein at a concentration of 10 mg ml−1 for an angiogram before in vivo imaging. A selected field of view was imaged every 5 min for approximately 4 h with a pixel size of 0.59 µm. The stacks for each time point were hyperstacked and registered in ImageJ/Fiji (v.2.14.0)60 through the vessel signal using the plugin Correct3D drift.
In vitro time-lapse live-cell imaging
Time-lapse imaging was performed using a widefield fluorescence microscope (Nikon Ti-HCS) with a ×10 objective (NA 0.3) and ×1.5 zoom; a confocal microscope (Zeiss Celldiscoverer7 with LSM 980) with a ×20 objective (NA 0.7) and ×0.5 zoom; and the Leica Thunder Imager live-cell imager with a ×20 objective. Coverslips were imaged every 15 min for 8–24 h at 37 °C with 5% CO2. Images were acquired with a pixel size of 486.86 nm (Ti-HCS) or 794.44 nm (CD7).
Cell division quantification
Cell division was quantified using live-cell time-lapse imaging in cocultures of neurons and ependymoma cells over 15–24 h. Cells were assigned their morphological cell state at the beginning of each time-lapse and cell division events were manually counted.
Cell migration speed measurement
Cell migration speed measurement was performed using ImageJ/Fiji(v.2.14.0)60. Images were registered using the HyperStackReg plugin in ImageJ/Fiji. Cell somata were outlined and the centre point of the cell was determined using the centroid function. The distance between the centre points of two timepoints was divided by the experimental observation time.
Measurement of TM dynamics
All TM measurements were performed in ImageJ/Fiji60. For both in vivo and in vitro experiments, TM protrusion was defined as TM outgrowth; retraction as decreased growth; and branching as TM dynamics whereby the secondary TM protrudes out of the pre-existing TM as previously described27,52. Net TM turnover was measured by measuring the length of all TMs at the starting time point and after 4 h.
Statistical analysis
Statistical analyses were conducted using R (v.4.3.3) and GraphPad Prism software (v.10). To determine statistical significance, we first assessed data normality using the Shapiro–Wilk test. If all groups followed a normal distribution (Shapiro–Wilk P > 0.05), we performed the Student’s t-test. If one or more groups did not follow a normal distribution (Shapiro–Wilk P < 0.05), we used the nonparametric Kruskal–Wallis test for comparisons across more than two groups. If the Kruskal–Wallis test was significant, we performed post hoc pairwise comparisons using the Wilcoxon rank-sum test with appropriate (for example, Bonferroni test) multiple-testing correction, or the Dunn’s multiple-comparison test. Only statistically significant P values are reported. Datapoints are presented as mean ± s.d. or mean ± s.e.m., depending on the experiment, as specified in the figure legends. The box plots follow the Tukey method, with bars extending from the 25th to 75th percentiles and the centre bar indicating the statistical median.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.