Animal studies
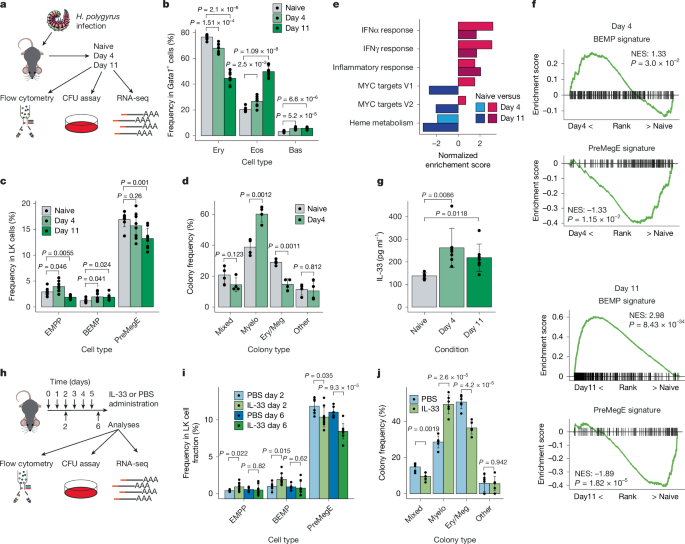
All animal studies were performed in accordance with UK Home Office regulations with approval by the University of Oxford Animal Welfare and Ethical Review Body (project license number 30/3359, PP2240412 and PP3246892) and the Francis Crick Institute Animal Welfare and Ethical Review Body (project license number PP8468807), as well as the Irish Health Products Regulatory Authority regulations License AE19136/P108 and P196, approved by Trinity College Dublin’s Animal Research Ethics Committee. NOD.Cg-KitW-41JTyr+PrkdcscidIl2rgtm1Wjl/ThomJ (NSGW41) mice45, Il1rl1-KO mice46, Gata2G320D knock-in mice30 and Gata1–eGFP BAC transgenic mice8 have been previously described. For IL-33 administration 9-week-old male C57BL/6J or Gata1–eGFP mice or short hairpin RNA (shRNA)-targeting Scramble or Lmo4, or control or St2−/− transplanted mice were intraperitoneally injected with 1 μg of recombinant mouse IL-33 (Peprotech) or PBS. For parasite infection, 9–12-week-old male C57BL/6J were used. Eight-nine-week-old Gata2D/D mice and wild-type littermates were infected with 200 H. polygyrus third-stage larvae, obtained from faecal cultures of H. polygyrus-infected mice, by oral gavage. Mice were maintained under specific pathogen-free conditions at 19–23 °C and 45–65% humidity on a 12-h light–12-h dark cycle. Sex and age-matched animals were randomly selected and allocated to experimental group, expect when assignment based on mouse genotype. Experiment and data collection were not performed blind to the condition of the experiment.
Plasmid construction
Flag–LMO2, MYC–LMO4, shRNA-targeting Scramble and shRNA-targeting Lmo4 constructs were obtained from Addgene (#64893, #22965, #59299 and #59292, respectively). The HA–FOG1 construct has been previously described17. Gata2 as well as LMO2/4 chimeras LMO-C1 and LMO-C2 cDNA were synthetized and subcloned into pLeGO-G (Addgene #27347) or pLeGO-C2 (Addgene #27339) under a mCherry-P2A reporter. For lentiviral transduction, Lmo4 cDNA was amplified and subcloned into pLeGO-C2 (Addgene #27339) or LT3-GEPIR (Addgene #111177) under a mCherry-P2A reporter.
Lentiviral production
HEK-293T (American Type Culture Collection CRL-11268) cells were plated in DMEM (Gibco) supplemented with 10% FBS (Gibco), 2 mM l-glutamine (Gibco) and 100 U ml−1 penicillin–streptomycin (Gibco), a day before co-transfection. Co-transfection was performed using PEIpro (Polyplus) according to the manufacturer’s recommendation, with psPAX2 (Addgene #12260), pMD2-G (Addgene #12259) and LeGO-C2-P2A-Lmo4/empty vectors. The viral supernatant was collected 48 h post-transfection, filtered using 0.45-µm cellulose acetate filter (Milipore) and concentrated using an Optima XPN-80 (Beckman Coulter) at 23.000 rpm for 1.5 h at 4 °C. HEK-293T cells were purchased from the supplier (Sigma) as a cell line authenticated by STR profiling, and was tested negative for mycoplasma.
Bone marrow transplantation
A total of 5 × 105 bone marrow cells were isolated from 12-week-old female Gata1–eGFP and Gata2+/+ or Gata2D/D mice and engrafted in lethally irradiated (10 Gy) 8–10-week-old female CD45.2 recipient mice by intravenous injection. To generate Il1rl1−/− chimeras, 5 × 105 total bone marrow cells from CD45.2 Il1rl1−/− or wild-type control mice were co-transplanted with an equal number of wild-type CD45.1 bone marrow cells into lethally irradiated (10 Gy) 8–10-week-old female CD45.1 recipient mice by intravenous injection.
Transplantation of genetically modified HSPCs
Bone marrow was isolated from 10–14-week-old female C57BL/6J or Gata1–eGFP mice8. Haematopoietic stem and progenitor cells (HSPCs) were enriched using CD117 MicroBeads and LS columns, according to the manufacturer’s recommendation (Miltenyi Biotec). Transduction of empty/Lmo4-expressing or shRNA-targeting Scramble/Lmo4 lentivirus was at a multiplicity of infection of 50 in StemSpan SFEM (STEMCELL Technologies) supplemented with 2 mM l-glutamine (Gibco), 100 U ml−1 penicillin–streptomycin (Gibco), 5 μg ml−1 polybrene (Merck), 100 ng ml−1 mouse SCF (Peprotech) and 100 ng ml−1 human TPO (Peprotech). Cells were spinfected at 33 °C and 700g for 1 h and incubated at 37 °C for 7 h or overnight. A total of 5 × 105 cells were injected intravenously into lethally irradiated (12 Gy; split dose) 8–10-week-old female CD45.2-recipient or CD45.1-recipient mice.
Colony-forming unit assay
Sorted cells (100–5,000) were plated in MethoCult GF M3434 (STEMCELL Technologies) supplemented with 25 ng ml−1 human TPO (Peprotech), 50 ng ml−1 mouse IL-5 (Peprotech), 50 ng ml−1 mouse IL-9 (Peprotech) and 100 U ml−1 penicillin–streptomycin (Gibco). Bulk or single-colony analysis was performed 10 days after cell plating. For single-colony readout, individual colonies were picked up in PBS (Gibco) supplemented with 5% FBS (Gibco) in 96-well plates and analysed by flow cytometry.
Human single-cell culture and xenotransplantation
Bone marrow from healthy volunteer was purchased from Lonza (Lonza Bioscience). CMP CD131+ was isolated by flow cytometry and cultured at the single-cell level in round-bottom 96-well plates in 50 μl of StemSpan (STEMCELL Technologies) with 20 ng ml−1 human SCF, 20 ng ml−1 human Flt3L, 20 ng ml−1 human IL-3, 50 ng ml−1 human IL-5, 20 ng ml−1 human IL-6, 50 ng ml−1 human GM-CSF, 20 ng ml−1 human G-CSF, 40 μg ml−1 human LDL (Sigma-Aldrich), 0.5 U ml−1 erythropoietin, 100 ng ml−1 thrombopoietin with 100 U ml−1 penicillin–streptomycin (Gibco), 0.1 mg ml−1 streptomycin and with or without 100 ng ml−1 human IL-33. Cytokines supplied by Peprotech. Lineage analysis was performed by flow cytometry, 20 days after cell plating. For xenotransplantation, 104 CD34+ cells isolated by flow cytometry were injected in NSGW41 mice by intrafemoral injection.
ELISA assay
Bone marrow supernatant was collected from tibias and femur by flushing. IL-33 was quantified with mouse/rat IL-33 Quantikine ELISA kit (R&D) according to the manufacturer’s recommendation, using CLARIOstar Plus (BMG labtech).
Flow cytometry analysis and cell sorting
Mouse bone marrow cells were isolated by crushing leg bones in FACS media that consisted of PBS (Gibco) supplemented with 2% FBS (Gibco). For HSPC hierarchy analysis, bone marrow cells were subjected to CD117 enrichment using CD117 MicroBeads and LS column, following the manufacturer’s recommendation (Miltenyi Biotec). Spleen cells were obtained by crushing the mouse spleen on a 0.70-μM filter using FACS media. Isolation of peritoneal cavity cells was performed by intraperitoneal injection of PBS (Gibco) in culled mice. After peritoneum massage, peritoneal suspension containing cells was collected, spun down at 500g for 5 min at 4 °C and cell pellet resuspended in FACS media. Peripheral blood was collected from the mouse tail vein into lithium heparin-coated microvettes (Sarstedt). An aliquot was collected for analysis of red blood cells and platelets. For mononuclear cell fraction analysis, samples were subjected to red blood cell lysis using ammonium chloride solution (STEMCELL Technologies) for 10 min in ice. After centrifugation at 500g for 5 min at 4 °C, the cell pellet was resuspended in FACS media. For xenografted human cell sorting, bone marrow cells isolated from NSGW41-transplanted mice were subjected to mCD45/Ter119 depletion using mCD45 and Ter119 MicroBeads and LS column, following the manufacturer’s recommendation (Miltenyi Biotec). Cells were incubated in FACS media containing Fc-block antibody (eBioscience) and subsequently stained with antibody cocktail for 20 min in ice. Samples were washed with FACS media, spun down at 500g for 5 min at 4 °C and resuspended in FACS media supplemented with living/dead cell dye: 1 μg ml−1 7AAD (Insight Biotechnology) or 0.1 μg ml−1 DAPI (Miltenyi Biotec). Intracellular LMO4 staining was conducted using the True-Nuclear Transcription Factor Buffer Set (BioLegend) according to the manufacturer’s protocol. Stained cells were analysed on LSR Fortessa or LSR X-20 flow cytometers (BD Biosciences). Cell sorting was carried out on FACSAria III, FACSAria Fusion or FACSymphony S6 (BD Biosciences) using FACS DIVA software (v9.0). Analysis was performed using FlowJo software (v10.8.1). The antibodies used are listed in Supplementary Table 13.
Co-IP and western blot
Transfection of HEK293T cells was performed using PEIpro (Polyplus) according to the manufacturer’s recommendation. Forty-eight hours post-transfection, cells were washed in PBS (Gibco) and isolated using trypsin–EDTA 0.05% (Gibco). Total cell lysis was performed using RIPA buffer (25 mM Tris-HCl, 150 mM NaCl, 1% NP-40, 1% sodium deoxycholate and 0.1% SDS) supplemented with cOmplete protease inhibitor cocktail (Roche) for 20 min on ice. Samples were sonicated for one cycle (30 s ON/OFF) using Bioruptor Pico (Diagenode) and spun down at 10,000g for 25 min at 4 °C. Proteins in supernatant were quantified using Qubit Protein broad range assay (Invitrogen). The following antibodies were used for immunoprecipitation: anti-GATA2 (R&D), anti-Flag (Sigma and Cell Signaling) or anti-MYC (Invitrogen and Cell Signaling). Immune complexes were purified using protein G magnetic beads (NEB) followed by serial washes in RIPA buffer supplemented with cOmplete protease inhibitor cocktail (Roche). Magnetic beads were resuspended in 2X Laemmli buffer (Bio-Rad) supplemented with 100 mM dithiothreitol (Merck) and incubated 15 min at 96 °C. Beads were separated on a magnetic rack and supernatant collected for western blotting, according to Pierce protocols.
In vitro signalling inhibitor treatment
Sorted EMPP isolated from Gata1–eGFP mice were plated in round-bottom 96-well plates in 50 µl of StemSpan SFEM (STEMCELL Technologies) supplemented with 100 U ml−1 penicillin and 0.1 mg ml−1 streptomycin, 2 mM l-glutamine, 20 ng ml−1 mouse SCF (PeproTech), 20 ng ml−1 human Flt3-ligand (PeproTech), 20 ng ml−1 human thrombopoietin (PeproTech) and PBS or recombinant IL-33 (PeproTech). Treatment was performed with SP600125 (1 μM; Medchem Express), U0125 (5 μM, Medchem Express), IKK-16 (0.5 μM, Medchem Express) or PBS for 16 h. Equal living cell numbers between treatment groups were sorted by flow cytometry directly in lysis buffer using CellsDirect One-Step RT–qPCR kit, immediately followed by cDNA synthesis and RT–qPCR using TaqMan Gene Expression Master Mix with QuantStudio 3 (Applied Biosystems), according to the manufacturer’s protocol.
Microfluidics single-cell gene expression analysis
Single cells were sorted in a 96-well PCR plate in 5 μl of CellsDirect One-Step RT–qPCR lysis buffer (Life Technologies) supplemented with 0.2X TaqMan probes for each targeted gene (Supplementary Table 14). cDNA synthesis and pre-amplification were conducted with the following program: 15 min at 50 °C, 2 min at 95 °C and 22 cycles of (15 s at 95 °C; 4 min at 60 °C). Samples were diluted in 5 volumes of TE buffer (Invitrogen) and gene expression quantified using the BioMark 192.24 Dynamic Array platform (Fluidigm) according to the manufacturer’s protocol.
Gene expression was measured according to ΔCt method using Hprt1 and B2m (for mouse) or HPRT and B2M (for human) as housekeeping genes with the geometric mean normalization method (Supplementary Tables 15–17). Low-quality cells marked by non-detectable housekeeping gene expression were excluded, as well as non-targeted cells characterized by non-detectable expression of Gata1 and expression of lymphoid genes (Dntt, Irf8, Pax5, Ebf1 or Tcf7). Principal component analysis, UMAP projection and clustering were performed using Seurat (v5)47. Single-cell GSEA was performed using ssGSEA (https://rpubs.com/pranali018/SSGSEA).
RNA-seq
Bulk RNA-seq was performed as previously described48. In brief, 100 cells were sorted in lysis buffer containing 0.18% Triton X-100 (Sigma-Aldrich), 4 U RNase inhibitor (Takara), 2.5 mM dNTP (Invitrogen) and 2.5 μM oligo-dT (Biomers.net). Reverse transcription was performed using 50 U SMARTScribe reserve transcriptase (Takara), and cDNA amplification was performed using 50 U SeqAmp Polymerase (Takara). cDNA was purified using SPRIselect beads (Beckman Coulter) and the library prepared using Nextera XT DNA Library Preparation Kit and Nextera XT Index kit (Illumina) according to the manufacturer’s recommendation. Libraries were sequenced on Illumina NovaSeq6000 or NovaSeq X Plus (150 bp; paired-end read).
ATAC-seq
Bulk ATAC-seq was performed based on the Omni-ATAC protocol49. In brief, 500–1,000 cells were sorted into tagmentation mix containing 1X TD buffer (Illumina), 0.01% digitonin (Promega) and 0.1% Tween-20 (Bio-Rad), and tagmentation was performed using TDE1 enzyme (Illumina) at 37 °C for 30 min. DNA was purified using the MinElute PCR Purification Kit (Qiagen) and amplified using NEBNext High-Fidelity 2×PCR master mix (NEB) and customized Nextera primers (IDT). PCR product was purified using SPRIselect beads (Beckman Coulter), quantified using the NEBNext Library Quant Kit (NEB) and sequenced on Illumina NovaSeq X Plus (150 bp; paired-end read).
ChIP–seq
ChIP was performed as previously described50. In brief, the HPC-7 cell line was transduced using doxycycline-inducible Lmo4/empty-expressing lentivirus and the transduced cell selected using puromycin. Doxycycline (1 μg ml−1) was added in culture media for 48 h and mCherry+ cells sorted by flow cytometry. Cells (1 × 106) were incubated with 2 mM disuccinimidyl glutarate and 1% paraformaldehyde, followed by addition of glycine to 125 mM and incubation for 5 min. Cells were pelleted and washed in PBS (Gibco) supplemented with cOmplete protease inhibitor cocktail (Roche). Pelleted cells were resuspended in lysis buffer A (50 mM HEPES pH 7.5, 140 mM NaCl, 1 mM EDTA, 10% glycerol, 0.5% NP-40 and 0.25% Triton X-100) supplemented with cOmplete protease inhibitor cocktail (Roche) and incubated in ice for 15 min. Nuclei were spun down and washed in lysis buffer B (200 mM NaCl, 1 mM EDTA, 0.5 mM EGTA and 10 mM Tris-HCl pH 7.5), then pelleted and resuspended in wash buffer (50 mM Tris-HCl pH 7.5 and 50 mM CaCl2) supplemented with cOmplete protease inhibitor cocktail (Roche). Nuclei were pelleted and resuspended in chromatin digestion mix (1X micrococcal nuclease buffer (NEB), 2,000 U micrococcal nuclease (NEB) and 0.1 mg ml−1 BSA) and incubated at 37 °C for 10 min. The reaction was stopped by adding Stop buffer (0.5% SDS, 5 mM EDTA and 5 mM EGTA final concentration) and tubes transferred in ice. Digested nuclei were sonicated for one cycle 30 s ON/OFF using Bioruptor Pico (Diagenode) and diluted in 4 volumes of ChIP dilution buffer (0.01% SDS, 1.1% Triton X-100, 1.2 mM EDTA, 16.7 mM Tris-HCl pH 8.1 and 167 mM NaCl). Relevant antibodies (Supplementary Table 13) were added and incubated at 4 °C overnight. Protein G magnetic beads (NEB) were added and incubated for 2 h at 4 °C. Bead-bound chromatin was washed twice in low-salt buffer (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris-HCl pH 8.1 and 150 mM NaCl), twice in high-salt buffer (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris-HCl pH 8.1 and 500 mM NaCl), twice in LiCl buffer (0.25 M LiCl, 1% NP-40, 1% deoxycholic acid, 1 mM EDTA and 10 mM Tris-HCl pH 8.1) and once in TE buffer (1 mM EDTA and 10 mM Tris-HCl pH 8.1). DNA was reverse crosslinked by adding 1.6 U proteinase K (NEB) and incubated at 55 °C for 3 h, then at 65 °C overnight. Finally, DNA was purified with a 1.8X ratio SPRI clean-up and quantified using the Qubit dsDNA HS Assay Kit (Thermo Fisher Scientific). Sequencing libraries were prepared using the NEBNext Ultra II DNA Library Prep Kit for Illumina (NEB) according to the manufacturer’s recommendation, and sequenced aiming 40 million reads per sample in a NextSeq550 or NovaSeq-X Plus (Illumina). The HPC-7 cell line was authenticated by differentiation into erythroid, megakaryocyte and myeloid lineages, and tested negative for mycoplasma.
RNA-seq analysis
Quality control and adaptor sequences were removed from fastq file using Trim Galore! (v0.6.5) and subsequent reads were quantified with Salmon (v1.2.0)51 using the mouse mm10 reference genome (https://www.gencodegenes.org/mouse/release_M10.html). Read count normalization and differential gene expression were performed using DESeq2 with default parameters (v1.42.0)52. GSEA was performed using the javaGSEA application (v4.2.3)53 and FGSEA R package (v1.34.2; http://bioconductor.org/packages/fgsea/).
ATAC-seq analysis
Quality control and adaptor sequences were removed from Fastq files using Trim Galore! (v0.6.5). Reads were aligned to the mm10 reference genome using Bowtie2 (v2.4.1)54. Subsequent SAM files were converted to BAM files, and PCR duplicate reads were filtered out using SAMtools (v1.17)55. Peak calling was performed using MACS2 (v2.1.2)56 with the following parameters: -q 0.05-nomodel–shift -100-extsize 200. A normalized Bigwig file was generated using bamCoverage (deepTools v3.5.3)57 and visualized on Integrative Genomics Viewer (v2.19.1)58. Unique reads mapped to these regions were quantified using featuresCounts (v2.0.1)59. Differential open chromatin regions were measured using DESeq2 (v1.42.0)52. For GSEA, peaks were ranked by statistic from DESeq2 to generate a rank-ordered list that was imported into the javaGSEA application (v4.2.3)53 and FGSEA R package (v1.34.2; http://bioconductor.org/packages/fgsea/). Transcription factor footprint analysis was performed using TOBIAS (v0.12.1)25 with the JASPAR2020 vertebrate core database60. BAM files from replicates were aggregated into a single BAM file and used for transposition insertion bias correction using the ATACorrection function. The transcription factor footprint score within selected open chromatin regions was computed using corrected files with ScoreBigwig function. Finally, differential transcription factor occupancy was quantified using the BINDetect function.
ChIP–seq analysis
FASTQ files were processed to BAM files similarly as described in ATAC-seq analysis section, using mm10 (https://www.gencodegenes.org/mouse/release_M10.html) or hg19 (https://www.gencodegenes.org/human/release_19.html) reference genomes for experiment using HPC-7 or HEK293T cell lines, respectively. Peak calling was performed using MACS2 (v2.1.2)56 with the following parameters: -q 0.05-nomodel-extsize 200. Read counts mapped on these regions were quantified using featureCounts (v2.0.1)59 and a normalized Bigwig file was generated using bamCoverage (deepTools v3.5.3)57. Differential binding was measured using DESeq2 (v1.42.0)52 and visualized with deepTools (v3.5.3)57. GSEA was performed using FGSEA R package (v1.34.2).
Protein structural modelling
The structures and binding interfaces of the protein complexes were generated using AlphaFold3 (ref. 29). The interaction between LMO2 (residues 1–158), LDB1 LID (residues 337–375) and the GATA2 N-terminal zinc finger (residues 291–339) was modelled, and the binding interface between the GATA2 N-terminal zinc finger and FOG1 ZnF1 was based on the previously determined NMR structure (Protein Data Bank ID 1Y0J) of the GATA1 N-terminal zinc finger complexed with Drosophila melanogaster homologue, ush ZnF1 (dFOG; residues 202–235)61. Similarly, the complex containing full-length LMO4 (residues 1–165), LDB1 LID and GATA2 N-terminal zinc finger was also modelled using the same method. A third model was generated for the LMO-C2 construct, which included all residues from LMO2 aside from 112–151 of LMO4, in complex with LDB1 LID and GATA-2 N-terminal zinc finger. All models were visualized using the PyMOL Molecular Graphics System (v2.1.0; http://www.pymol.org/pymol), where proteins were rendered as cartoon diagrams.
Statistics and reproducibility
Statistical analysis and representation were performed using R software. For normally distributed data, two-tailed t-tests were applied with Welch’s correction, which did not assume equal variance. Normality was tested using the Shapiro–Wilk test. The statistical tests used have been described for each analysis in the corresponding figure legend. Sample sizes were chosen based on the variance previously observed in similar experiments, to allow detection of a 50% difference at P = 0.05 with 80% power. Block designs were used to allow variance estimates to be adjusted if necessary. Data collection and analysis were not performed blind to the conditions of the experiments. The boxplots display the median as the centre line of the box, with the box representing the distribution’s 25th (minima) and 75th (maxima) percentiles; the whiskers extend up to 1.5 times the interquartile range (Q3–Q1) from the minima and maxima.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.