Wiegand, T. et al. TnpB homologues exapted from transposons are RNA-guided transcription factors. Nature 631, 439–448 (2024).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Bikard, D. et al. Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system. Nucl. Acids Res. 41, 7429–7437 (2013).
Workman, R. E. et al. A natural single-guide RNA repurposes Cas9 to autoregulate CRISPR-Cas expression. Cell 184, 675–688.e619 (2021).
Wu, W. Y. et al. The miniature CRISPR-Cas12m effector binds DNA to block transcription. Mol. Cell 82, 4487–4502.e4487 (2022).
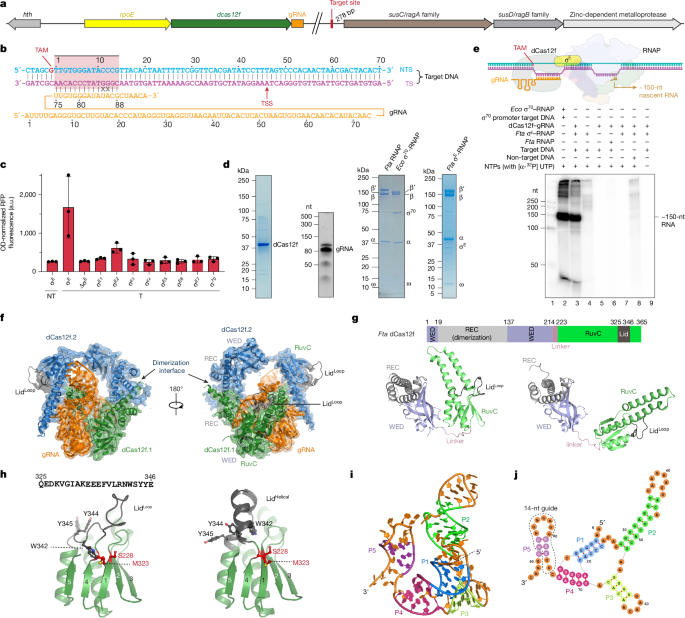
Hoffmann, F. T. et al. Exapted CRISPR–Cas12f homologues drive RNA-guided transcription. Nature https://doi.org/10.1038/s41586-026-10166-7 (2026).
deHaseth, P. L., Lohman, T. M., Burgess, R. R. & Record, M. T. Jr. Nonspecific interactions of Escherichia coli RNA polymerase with native and denatured DNA: differences in the binding behavior of core and holoenzyme. Biochemistry 17, 1612–1622 (1978).
Lee, D. J., Minchin, S. D. & Busby, S. J. Activating transcription in bacteria. Ann. Rev. Microbiol. 66, 125–152 (2012).
Feklistov, A., Sharon, B. D., Darst, S. A. & Gross, C. A. Bacterial sigma factors: a historical, structural, and genomic perspective. Ann. Rev. Microbiol. 68, 357–376 (2014).
Paget, M. S. & Helmann, J. D. The sigma70 family of sigma factors. Genome Biol. 4, 203 (2003).
Campbell, E. A. et al. Crystal structure of Escherichia coli sigmaE with the cytoplasmic domain of its anti-sigma RseA. Mol. Cell. 11, 1067–1078 (2003).
Zhang, Y. et al. Structural basis of transcription initiation. Science 338, 1076–1080 (2012).
Casas-Pastor, D. et al. Expansion and re-classification of the extracytoplasmic function (ECF) sigma factor family. Nucl. Acids Res. 49, 986–1005 (2021).
Mascher, T. Past, present, and future of extracytoplasmic function sigma factors: distribution and regulatory diversity of the third pillar of bacterial signal transduction. Ann. Rev. Microbiol. 77, 625–644 (2023).
Altae-Tran, H. et al. Diversity, evolution, and classification of the RNA-guided nucleases TnpB and Cas12. Proc. Natl Acad. Sci. USA 120, e2308224120 (2023).
Sampson, T. R., Saroj, S. D., Llewellyn, A. C., Tzeng, Y. L. & Weiss, D. S. A CRISPR/Cas system mediates bacterial innate immune evasion and virulence. Nature 497, 254–257 (2013).
Huang, C. J., Adler, B. A. & Doudna, J. A. A naturally DNase-free CRISPR-Cas12c enzyme silences gene expression. Mol. Cell 82, 2148–2160.e2144 (2022).
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Takeda, S. N. et al. Structure of the miniature type V-F CRISPR-Cas effector enzyme. Mol. Cell 81, 558–570 e553 (2021).
Xiao, R., Li, Z., Wang, S., Han, R. & Chang, L. Structural basis for substrate recognition and cleavage by the dimerization-dependent CRISPR-Cas12f nuclease. Nucl. Acids Res. 49, 4120–4128 (2021).
Hino, T. et al. An AsCas12f-based compact genome-editing tool derived by deep mutational scanning and structural analysis. Cell 186, 4920–4935.e4923 (2023).
Wu, T. et al. An engineered hypercompact CRISPR-Cas12f system with boosted gene-editing activity. Nat. Chem. Biol. 19, 1384–1393 (2023).
Harrington, L. B. et al. Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science 362, 839–842 (2018).
Yang, H., Gao, P., Rajashankar, K. R. & Patel, D. J. PAM-dependent target dna recognition and cleavage by C2c1 CRISPR-Cas endonuclease. Cell 167, 1814–1828 (2016).
Zhang, H., Li, Z., Xiao, R. & Chang, L. Mechanisms for target recognition and cleavage by the Cas12i RNA-guided endonuclease. Nat. Struct. Mol. Biol. 27, 1069–1076 (2020).
Sternberg, S. H., Redding, S., Jinek, M., Greene, E. C. & Doudna, J. A. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature 507, 62–67 (2014).
Liu, B., Hong, C., Huang, R. K., Yu, Z. & Steitz, T. A. Structural basis of bacterial transcription activation. Science 358, 947–951 (2017).
Chen, J. et al. E. coli TraR allosterically regulates transcription initiation by altering RNA polymerase conformation. eLife https://doi.org/10.7554/eLife.49375 (2019).
Fang, C. et al. Structures and mechanism of transcription initiation by bacterial ECF factors. Nucl. Acids Res. 47, 7094–7104 (2019).
Lin, W. et al. Structural basis of ECF-sigma-factor-dependent transcription initiation. Nat. Commun. 10, 710 (2019).
Li, L., Fang, C., Zhuang, N., Wang, T. & Zhang, Y. Structural basis for transcription initiation by bacterial ECF sigma factors. Nat. Commun. 10, 1153 (2019).
Bae, B., Feklistov, A., Lass-Napiorkowska, A., Landick, R. & Darst, S. A. Structure of a bacterial RNA polymerase holoenzyme open promoter complex. eLife https://doi.org/10.7554/eLife.08504 (2015).
Lin, W. et al. Structural basis of mycobacterium tuberculosis transcription and transcription inhibition. Mol. Cell 66, 169–179.e168 (2017).
Marin-Gonzalez, A., Vilhena, J. G., Perez, R. & Moreno-Herrero, F. A molecular view of DNA flexibility. Q. Rev. Biophys. 54, e8 (2021).
Nudler, E. RNA polymerase active center: the molecular engine of transcription. Annu. Rev. Biochem. 78, 335–361 (2009).
Geszvain, K., Gruber, T. M., Mooney, R. A., Gross, C. A. & Landick, R. A hydrophobic patch on the flap-tip helix of E.coli RNA polymerase mediates sigma(70) region 4 function. J. Mol. Biol. 343, 569–587 (2004).
Minakhin, L. et al. Bacterial RNA polymerase subunit omega and eukaryotic RNA polymerase subunit RPB6 are sequence, structural, and functional homologs and promote RNA polymerase assembly. Proc. Natl Acad. Sci. USA 98, 892–897 (2001).
Ghosh, P., Ishihama, A. & Chatterji, D. Escherichia coli RNA polymerase subunit ω and its N-terminal domain bind full-length β′ to facilitate incorporation into the α2β subassembly. Eur. J. Biochem. 268, 4621–4627 (2001).
Bu, F. et al. Cryo-EM structure of Porphyromonas gingivalis RNA polymerase. J. Mol. Biol. 436, 168568 (2024).
Lane, W. J. & Darst, S. A. The structural basis for promoter −35 element recognition by the group IV sigma factors. PLoS Biol. 4, e269 (2006).
Li, J. et al. Structure of the transcription open complex of distinct sigma(I) factors. Nat. Commun. 14, 6455 (2023).
Rohs, R. et al. The role of DNA shape in protein–DNA recognition. Nature 461, 1248–1253 (2009).
Feklistov, A. & Darst, S. A. Structural basis for promoter-10 element recognition by the bacterial RNA polymerase sigma subunit. Cell 147, 1257–1269 (2011).
Lim, H. M., Lee, H. J., Roy, S. & Adhya, S. A “master” in base unpairing during isomerization of a promoter upon RNA polymerase binding. Proc. Natl Acad. Sci. USA 98, 14849–14852 (2001).
Bayley, D. P., Rocha, E. R. & Smith, C. J. Analysis of cepA and other Bacteroides fragilis genes reveals a unique promoter structure. FEMS Microbiol. Lett. 193, 149–154 (2000).
Chen, S., Bagdasarian, M., Kaufman, M. G. & Walker, E. D. Characterization of strong promoters from an environmental Flavobacterium hibernum strain by using a green fluorescent protein-based reporter system. Appl. Environ. Microbiol. 73, 1089–1100 (2007).
Feng, Y., Zhang, Y. & Ebright, R. H. Structural basis of transcription activation. Science 352, 1330–1333 (2016).
Murakami, K. S., Masuda, S. & Darst, S. A. Structural basis of transcription initiation: RNA polymerase holoenzyme at 4 Å resolution. Science 296, 1280–1284 (2002).
Zuo, Y. & Steitz, T. A. Crystal structures of the E. coli transcription initiation complexes with a complete bubble. Mol. Cell 58, 534–540 (2015).
Murakami, K. S., Masuda, S., Campbell, E. A., Muzzin, O. & Darst, S. A. Structural basis of transcription initiation: an RNA polymerase holoenzyme-DNA complex. Science 296, 1285–1290 (2002).
Hubbard, B. P. et al. Continuous directed evolution of DNA-binding proteins to improve TALEN specificity. Nat. Methods 12, 939–942 (2015).
Feklistov, A. RNA polymerase: in search of promoters. Ann. N. Y. Acad. Sci. 1293, 25–32 (2013).
Velappan, N., Sblattero, D., Chasteen, L., Pavlik, P. & Bradbury, A. R. M. Plasmid incompatibility: more compatible than previously thought. Protein Eng. Des. Sel. 20, 309–313 (2007).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Tan, Y. Z. et al. Addressing preferred specimen orientation in single-particle cryo-EM through tilting. Nat. Methods 14, 793–796 (2017).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
McWilliam, H. et al. Analysis Tool Web Services from the EMBL-EBI. Nucleic Acids Res. 41, W597–600 (2013).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–324 (2014).
Kerpedjiev, P., Hammer, S. & Hofacker, I. L. Forna (force-directed RNA): simple and effective online RNA secondary structure diagrams. Bioinformatics 31, 3377–3379 (2015).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Grantz, J. M. et al. The platelet and plasma proteome and targeted lipidome in postpartum dairy cows with elevated systemic inflammation. Sci. Rep. 14, 31240 (2024).
Lee, Y. et al. Proteomics-based models of gene expression and cellular control of cotton fiber development. Preprint at bioRxiv https://doi.org/10.1101/2025.02.05.636703 (2025).
Demichev, V., Messner, C. B., Vernardis, S. I., Lilley, K. S. & Ralser, M. DIA-NN: neural networks and interference correction enable deep proteome coverage in high throughput. Nat. Methods 17, 41–44 (2020).
UniProt, C. UniProt: the Universal Protein Knowledgebase in 2025. Nucleic Acids Res. 53, D609–D617 (2025).
Katoh, K., Kuma, K., Toh, H. & Miyata, T. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucl. Acids Res. 33, 511–518 (2005).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinf. 10, 421 (2009).
Chamberlain, S. A. & Szocs, E. Taxize: taxonomic search and retrieval in R. F1000Res 2, 191 (2013).
Schoch, C. L. et al. NCBI Taxonomy: a comprehensive update on curation, resources and tools. Database https://doi.org/10.1093/database/baaa062 (2020).
Xu, S. et al. Ggtree: a serialized data object for visualization of a phylogenetic tree and annotation data. Imeta 1, e56 (2022).