Fine-mapping
We obtained fine-mapping summary data, including PIPs for each variant and 95% CSs, for 48 traits in the UKBB and BBJ from previous studies31,66. Detailed methods are available from these studies, but we describe the general approach briefly here. The UKBB is a population-based cohort in the UK, of which 366,194 ‘white British’ individuals were selected for inclusion (https://github.com/Nealelab/UK_Biobank_GWAS). The BBJ is a large non-European hospital-based biobank from Japan, from which 178,726 individuals of Japanese descent were included in this study. We selected 48 traits across 12 domains (including an ‘other’ category) from the UKBB and BBJ (Supplementary Table 1). GWAS was performed on each using a generalized linear mixed model, as implemented in SAIGE67 (for binary traits) or BOLT-LMM68,69 (for quantitative traits), with the following covariates: age, sex, age2, age × sex, age2 × sex and the top 20 genetic principal components. Fine-mapping was then performed using SuSiE30 with the GWAS summary statistics and in-sample dosage linkage disequilibrium in merged 1.5-Mb windows.
Similarly, we obtained fine-mapping results across 49 tissues from the GTEx v.8 from previous studies34,66,70 and summarize the approach below. The GTEx project is a tissue-based biobank with genotypes (whole genome sequencing) and gene-expression data (RNA-seq) for 838 individuals across 49 tissues (Supplementary Table 1), which are grouped into 11 systems. We obtained cis-eQTL summary statistics, genotype principal components (PCs) and other associated data as input to fine-mapping, which was performed in a similar way to the fine-mapping of complex traits from the UKBB, with the main modification being that covariates (including genetic PCs) were projected out of the genotypes before the linkage disequilibrium calculation, because GTEx is a more ancestrally heterogeneous cohort39. When collapsing PIPs across multiple tests into a single estimate, the maximum PIP across traits and/or tissues is used. As well as individual trait CSs, merged CSs across traits or across tissues were obtained, as previously described31.
Variant selection and oligonucleotide design
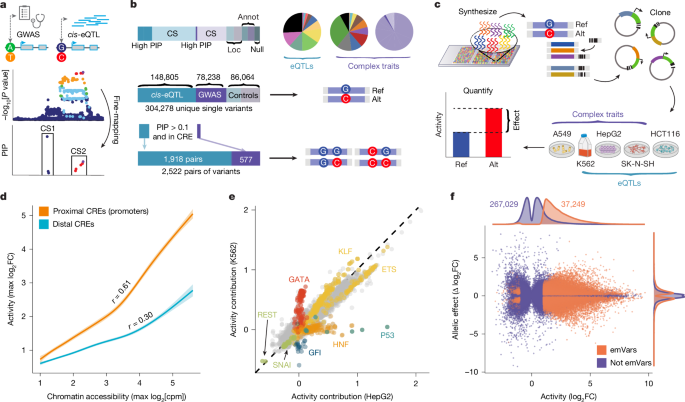
We designed 200-bp oligonucleotides centred around the reference and alternative alleles of fine-mapped eQTLs from GTEx v.8 and fine-mapped complex traits from the UKBB and BBJ cohorts71,72 (Fig. 1a). Variants were selected for inclusion if they demonstrated a PIP > 0.5 or greater in any tissue or a PIP > 0.1 in any trait, or if they fell within a 95% CS with fewer than 25 variants (eQTLs), 30 variants (quantitative traits) or 75 variants (binary traits). Variants with PIP > 0.1 in any of 3 GTEx tissues approximately corresponding to cell types for MPRA were also included. That is, we selected additional fine-mapped eQTLs from liver (matched to HepG2), transverse colon (matched to HCT116) and any brain tissue (matched to SK-N-SH). CSs were considered to be successfully assayed if variants containing at least 80% of the probability had MPRA measurements. Most (92.5%) of the selected variants were single nucleotide polymorphisms. The remaining 22,951 variants were small insertions or deletions (indels), of which 92.2% affected fewer than 10 bp, and only 11 involved more than 50 bp. In total, we included 148,805 eQTL test variants and 78,238 complex trait test variants (Fig. 1b and Supplementary Tables 1–3), with 5,631 variants shared across the two sets. All test and control variants (see below) were split into 8 libraries (Supplementary Table 29).
To test for potential non-additive effects between trait-associated variants in the same element, we selected 2,522 pairs of variants that were within 150 bp of each other, were fine-mapped with PIP > 0.1 for at least one trait, and overlapped with an endogenous CRE in at least one cell type (Supplementary Table 18). To maximize the power to detect non-additive effects, oligonucleotides were designed for up to six possible windows. Centring on each variant in the pair, the variant was placed at approximately 50 bp, 100 bp or 150 bp (the exact location could change for variants causing insertions or deletions), and windows in which the second variant was at least 10 bp from the edge of the oligonucleotide were included (Fig. 3a). For each window, we designed oligonucleotides for all four possible diplotypes (Ref/Ref, Ref/Alt, Alt/Ref and Alt/Alt).
As well as trait-associated test variants, we designed several comprehensive sets of controls. We selected 32,534 ‘location-matched’ controls that were more than 150 bp but less than 500 bp away from test variants and were not significantly associated with a complex trait (P > 0.0001) or change in gene expression (P > 0.001) in our summary data. Next, we selected 39,708 ‘annotation-matched’ controls that were not in 95% CSs or significantly associated with a trait that were matched for MAF, linkage disequilibrium scores, standard genomic annotations (promoter, 3′ UTR, coding, 5′ UTR, intron, CRE, evolutionary constraint) as well as CRE annotations from relevant ENCODE cell types and tissues. Matching was done by computing propensity scores and selecting the closest score match using the MatchIt R package. An example of this matching procedure is shown in Extended Data Fig. 1a. Finally, we selected 14,710 ‘null’ trait controls, for which we simulated a typical GWAS phenotype and performed fine-mapping and variant selection in the same manner as was done for complex traits for UKBB and BBJ. To simulate these phenotypes73, we drew causal effects from variants present in UKBB or BBJ, largely following a previous approach. That is, we assumed 20% trait heritability, either 1,000 (for binary traits) or 5,000 (for quantitative traits) total causal variants, and a MAF-dependent per-allele effect size consistent with previously reported complex-trait architecture73 (α = −0.38). Unlike annotation- and location-matched controls, null control variants were not expected to be enriched at CREs. Instead, this set of controls was designed to reflect the linkage disequilibrium patterns of real complex traits and provide a true null set from the perspective of genomic annotation. For all three classes of control, matching was done across PIP bins, resulting in 37,437 controls for eQTLs and 48,880 for complex traits (Fig. 1b).
We also selected 128 elements containing variants from the original screen that overlap endogenous CREs on which to perform saturation mutagenesis. This set contained a total of 136 emVars, where 114 elements each contain 1 emVar, 8 elements contain 2 emVars, and 6 elements contain 1 emVar and 1 non-emVar (Supplementary Table 23). Three categories of variants were prioritized for saturation mutagenesis. First, we prioritized 83 emVars with ‘canonical regulatory mechanisms’, in which a fine-mapped emVar was predicted to disrupt the known motif at an occupied TF site (that is, the variant disrupts a TF motif and that variant resides within a ChIP–seq peak for that TF). Second, we prioritized 53 emVars with ‘non-canonical or unclear regulatory mechanisms’, in which a fine-mapped emVar did not disrupt the known motif of a TF with evidence of occupancy from ChIP–seq. Candidate variants were manually inspected and selected variants had at least one alternative measure of function, such as residing in a DHS footprint, lying in the flank (0–10 bp) of a motif at an occupied TF site, or having strong predicted allelic effects in sequence-based models of function, such as Enformer. Third, we prioritized 14 ‘same CRE variant pairs’, in which at least one variant in the pair was an emVar and the pair demonstrated a non-additive effect, or both variants were strong candidates for the first two categories. Unless otherwise indicated, the 22 emVars selected as variant pairs were also included in their respective canonical and non-canonical categories.
Overall, variants were chosen to survey a variety of complex and molecular traits, a range of PIPs to allow for both confirmation (PIP > 0.9) and exploration (PIP > 0.1 and PIP < 0.9), a variety of predicted effects on different TFs and in different cell types, and sufficient representation in all three categories. In total, we selected 83 variants from category 1 (canonical mechanism), 53 variants from category 2 (non-canonical or unclear regulatory mechanisms) and 14 variant pairs from category 3 (same CRE variant pairs). For each of the 114 single-variant elements, we designed 200-bp oligonucleotides using both the reference and the alternative background sequences, as well as all possible single-nucleotide substitutions for both backgrounds (3 alternative nucleotides × 200 positions × 2 backgrounds). For each of the 14 variant pairs, we designed mutagenesis oligonucleotides using each diplotype (Ref/Ref, Ref/Alt, Alt/Ref and Alt/Alt) as a background (3 alternative nucleotides × 200 positions × 4 backgrounds).
Finally, we also included technical controls to evaluate the quality of individual MPRA experiments. For each library, we included 91 activity controls, 96 emVar controls and 506 negative controls selected from previously published experiments16,74 (Supplementary Table 30). Activity controls were chosen for their ubiquitous activity across multiple cell types and were selected to represent a broad range of enhancer activities. For the saturation-mutagenesis library, we included an additional 2,126 negative controls, selecting 1,876 ORF controls and 250 shuffled sequences from previously published work75,76.
MPRA library construction
The MPRA libraries were constructed as previously described16 with several modifications to the original protocol. The 230-bp oligonucleotides were synthesized using Agilent Technologies HiFi libraries for complex traits (seven 60 K libraries) and eQTL (two 244 K libraries) or Twist Biosciences for saturation mutagenesis (one 300 K library) with the designed 200-bp oligonucleotide in the middle and 15-bp adaptor sequences on either end. Unique 20-bp barcodes were added by PCR using primers #82 and #202 (Supplementary Table 31). Each oligonucleotide amplification consisted of 16–32 50-μl reactions containing 25 μl Q5 2xMM Ultra II, 2.5 μl each of 10 μM primer and 0.5 μl oligonucleotide library using the following PCR conditions (98 °C for 30 s; 6× [98 °C for 10 s; 60 °C for 15 s; 65 °C for 45 s]; 72 °C for 5 min). The PCR products were purified using AMPure XP SPRI (Beckman Coulter, A63881) and incorporated into the SfiI digested pMPRAv3:∆luc:∆xbaI (Addgene #109035) backbone vector by Gibson assembly (50 μl 2 × NEB HiFi Assembly MM, 2.2 μg DNA oligonucleotide pool, 2.0 μg SfiI digested pMPRAv3:∆luc:∆xbaI in a 100-μl reaction incubated at 50 °C for 1 h). To achieve a library composition of 200–300 barcodes per oligonucleotide, a test transformation was performed using 1 μl of the 20 μl SPRI purified Gibson assembly mixture and 50 μl of 10-beta electrocompetent cells (NEB) to determine the optimal amount of Gibson assembly and 10-beta cells to achieve the desired colony-forming unit (CFU) count. Based on the results of the test transformation, a subset of a second identical transformation was taken directly after electroporation and split across ten 1-ml cultures with 10-beta recovery media (NEB), then incubated for 1 h at 37 °C. After 1 h, each culture was independently expanded in 20 ml of Luria broth (LB) with 100 μg ml−1 carbenicillin. In parallel, CFU colony counting plates were created from four of the ten cultures. After 6.5 h of growth at 37 °C, the cultures were individually pelleted and frozen, and the desired number of cultures from ten expanded cultures was selected, based on the CFU counting plates, to reach an average of 200–300 CFUs (barcodes) per oligonucleotide (15 million to 21 million CFUs for complex traits, 49 million CFUs for eQTLs and 86 million for saturation mutagenesis). Cultures were combined and purified using a Qiagen Plasmid Plus Maxi or Midi kit. For each library, 16 colonies from the CFU plates were checked by colony PCR to determine the oligonucleotide insertion rate for the test transformation (insertion-rate range: 93–100%). The expanded purified pMPRAv3:∆GFP plasmid library was sequenced using Illumina 2 x 150 bp chemistry to acquire oligonucleotide–barcode pairings. To construct the final MPRA libraries, 10 μg of the pMPRAv3:∆GFP library was digested with AsiSI, and a GFP amplicon with a minimal TATA promoter (amplified from pMPRAv3:minP-GFP, Addgene 109035) was inserted using Gibson assembly (125 μl 2 × NEB HiFi Assembly MM, 5.28 μg GFP amplicon, 1.6 μg pMPRAv3:∆GFP in a 250-μl reaction incubated at 50 °C for 1.5 h). The resulting pMPRAv3 library includes the 200-bp oligonucleotide sequence positioned directly upstream of the minimal promoter and the 20-bp barcode falling in the 3′ UTR of GFP. The Gibson reaction was purified using 1.5× SPRI and eluted in 40 μl water before electroporation of 2–16 μl of purified plasmid library into 100–400 μl 10-beta cells. The electroporation was split across six 2-ml cultures and recovered at 37 °C for 1 h followed by expansion of each 2-ml culture in 500 ml of TB media with 100 μg ml−1 carbenicillin. After 16 h of growth at 30 °C, plasmid was purified using Qiagen Plasmid Plus Giga kits. The CFU counting plates from 4 of the 500-ml cultures were made to monitor transformation efficiencies (23 million–185 million CFUs for complex traits, 285 million–1 billion CFUs for eQTLs, and 123 million–171 million CFUs for saturation mutagenesis). Then, 16 colonies from the CFU plates were checked by colony PCR to determine the GFP insertion rate for the test transformation, with all libraries having a minimum of 80% of plasmids containing a GFP insert.
MPRA library transfections
We used five cell types for MPRA transfections, representing five distinct tissue types. The complex trait libraries were tested in K562, SK-N-SH, HepG2 and A549 cells; the eQTL libraries were tested in K562, SK-N-SH, HepG2 and HCT116 cells; and the saturation-mutagenesis library was tested in K562 and HepG2 cells. All cell lines were acquired from ATCC and routinely tested for mycoplasma and other common contaminants by The Jackson Laboratory’s Molecular Diagnostic Laboratory. Each replicate experiment was expanded from a low-passage cryopreserved aliquot followed by consistent handling for each cell type across all libraries. Specifically, K562 cells were grown in RPMI (Life Technologies, 61870127) supplemented with 10% FBS (Life Technologies, 26140) maintaining a cell density of 0.5–1.0 million cells per ml; SK-N-SH cells were grown in DMEM (Life Technologies, 10566-024) supplemented with 10% FBS (Life Technologies, 26140); HepG2 cells were grown in DMEM (Life Technologies, 10566-024) supplemented with 10% FBS (Life Technologies, 26140); A549 cells were grown in Ham’s F-12K (Life Technologies, 21-127-030) supplemented with 10% FBS (Life Technologies, 26140); and HCT116 cells were grown in McCoy’s 5a (ThermoFisher, 16600108) supplemented with 10% FBS (Life Technologies, 26140). All transfections were done using a Neon transfection system (Life Technologies), transfecting 150 million (complex traits), 500 million (eQTL) or 700 million (saturation mutagenesis) cells for each replicate. Transfections were done using 100-μl tips with optimized settings for each cell type: K562, 10 million cells per 100 μl, 5 μg of plasmid, 3 pulses of 1,450 V for 10 ms; SK-N-SH, 10 million cells per 100 μl, 10 μg of plasmid, 3 pulses of 1,200 V for 20 ms; HepG2, 10 million cells per 100 μl, 5 μg of plasmid for complex traits and saturation mutagenesis or 10 μg of plasmid for eQTLs, 1 pulse of 1,200 V for 50 ms; A549, 10 million cells per 100 μl, 5 μg of plasmid, 2 pulses of 1,200 V for 30 ms; and HCT116, 10 million cells per 100 μl, 10 μg of plasmid, 2 pulses of 1,350 V for 20 ms. We did a minimum of 5 replicates for each library and collected each replicate 24 h after transfection by rinsing three times with PBS and collecting by centrifugation. Cell pellets were either frozen at −80 °C for later processing or homogenized immediately in RLT buffer (RNeasy Maxi kit) and dithiothreitol, before freezing at −80 °C. For each cell type and library, no more than two replicates were done on the same day and with parallel replicates processed from independently expanded batches of cells.
RNA isolation and MPRA RNA library generation
Total RNA was extracted from the cell homogenates using the Qiagen RNeasy Maxi kit followed by a 1-h Turbo DNase treatment (ThermoFisher) with 5 μl of DNase in 1,475 μl total volume for 1 h at 37 °C. To stop the DNase digestion, 15 µl of 10% SDS and 150 µl of 0.5 M EDTA were added, and the sample was incubated at 70 °C for 5 min. GFP mRNA was then pulled down using a mixture of three GFP-specific biotinylated primers at 0.5 nM (#120, #123 and #126; Supplementary Table 31) in 0.2× SSC and 33% formamide (MilliporeSigma 4650–500 ML) and incubated for 2.5 h at 65 °C. Biotin probes hybridized with GFP mRNA were then captured by adding 400 μl of pre-washed Sera Mag beads (Fisher Scientific) eluted in 500 μl of 20× SSX by agitation at room temperature for 15 min. The beads were captured on a magnet and washed once with 1× SSX and twice with 0.1× SSC. After removal of the last wash, 50 μl of water was added and the RNA was treated overnight with Turbo DNase at 37 °C. The beads were then collected by magnet and the supernatant removed and purified with 2× RNA SPRI. Complementary DNA was created using SuperScript III (Life Technologies) in a 100-μl reaction with a gene-specific primer for the GFP transcript (20 μM primer #19; Supplementary Table 30) and a modified elongation temperature (47 °C for 80 min). The GFP mRNA abundance was quantified by quantitative PCR to determine the cycle threshold for each replicate using a QuantStudio 5 qPCR instrument (Reaction mix: 5 μl Q5 Ultra II 2x, 0.5 μl of 10 μM primers #801 and #802, 1.66 μl 1:10,000 Sybrgreen 1, 1 μl cDNA or GFP mRNA in a 10 μl reaction, Cycle conditions: 98 °C for 30 s; 40× [98 °C for 10 s; 62 °C for 15 s; 72 °C for 30 s]; Supplementary Table 31). A standard curve with the final MPRA library (10 fg–1 ng) was run alongside 1 μl of cDNA and 1 μl of GFP mRNA for each replicate. Any samples showing amplification in the mRNA sample were discarded for having plasmid carry-over. Replicates within a cell type and library were diluted to approximately the same concentration, based on the quantitative PCR (qPCR) results, and first-round PCR (98 °C for 30 s; 6–13× [98 °C for 10 s; 62 °C for 15 s; 72 °C for 30 s]; 72 °C for 2 min) with primers #801 and #802 (Supplementary Table 31) were used to amplify barcodes associated with GFP cDNA sequences for each replicate. A second round of PCR (98 °C for 30 s; 6× [98 °C for 10 s; 68 °C for 15 s; 72 °C for 30 s]; 72 °C for 2 min) was used to add Illumina sequencing adaptors. The resulting MPRA barcode libraries were spiked with 0.01–1% PhiX and sequenced on an Illumina NextSeq 500 or NovaSeq SP.
MPRA data processing and main analysis
Data from MPRA were analysed as previously described16 using MPRAsuite, which includes MPRAmatch (barcode and oligonucleotide pairing), MPRAcount (tag assignment and counting) and MPRAmodel (activity and allelic effect estimates) (https://github.com/tewhey-lab/MPRASuite). In brief, oligonucleotide–barcode pairings were identified by merging paired-end reads into single amplicons using Flash (2.2.00)77. Genomic DNA sequences and barcodes were extracted and the genomic DNA element was mapped back to the oligonucleotide design file using minimap2 (v.2.17-r941)78. Alignments showing more than 5% error to the design files were discarded, leaving only high-quality alignments, which were compiled in a look-up table with their matching barcodes. After sequencing of the RNA and plasmid libraries, 20-bp barcode tags were assigned to oligonucleotides from the look-up table and oligonucleotide counts were aggregated by addition across all barcodes. Variants with fewer than 30 DNA counts or with exactly 0 RNA counts in oligonucleotides for either allele were excluded from all downstream analysis. In the diplotype analyses, a stricter filter was used, and variants with fewer than 100 DNA counts or fewer than 10 RNA counts across all 4 diplotypes were excluded. When variants were assayed in multiple libraries, the library with the highest sum of the square roots of plasmid counts in reference and alternative sequences was selected.
MPRAmodel uses DESeq2 (1.42.1) as the framework for normalization and statistical analysis. For each library, counts were normalized across samples using a ‘summit-shift’ approach, which centres the distribution of RNA/DNA ratios for each sample at 1. To determine significant differences between DNA plasmid count and RNA count, a negative binomial generalized linear model (GLM) was used in DESeq2 with independent dispersion estimates for each cell type and library. We estimated element activity in units of log2(fold change) for RNA counts compared with DNA counts by including a contrast in the design matrix of DESeq2 to compare treatment (RNA versus DNA) and used Wald’s test to evaluate the significance of this estimate, correcting for multiple hypothesis testing in each cell type and library using Bonferroni’s method. We estimated an allelic effect in units of ∆ log2(fold change) (or ∆log2 activity) for the difference in RNA counts relative to DNA counts at the Alt oligonucleotide compared with the Ref oligonucleotide by including a contrast in the design matrix of DESeq2 to compare alleles (Alt versus Ref) and treatment (RNA versus DNA) and used Wald’s test to evaluate significance followed by FDR estimation using Benjamini and Hochberg’s method. For simplicity, we refer to P values obtained after adjusting for the FDR as ‘FDR’ rather than ‘q-value’ or ‘adjusted P-value’ in all relevant instances. Elements passing a specific effect size and significance threshold for RNA versus DNA are defined as active, and variants in active elements passing a specific significance threshold are defined as expression-modulating variants or emVars. By performing a grid search as described in the following section, we define elements as active if |log2[fold change]| > 1 and Bonferroni-adjusted P value < 0.01, and variants as emVars if either the reference or alternative allele element is active and FDR < 10% for a non-zero allelic effect. For specific analyses requiring cell-type agnostic estimates, we performed fixed-effect meta-analyses of activity and allelic effects across cell types and used these estimates. In Supplementary Table 4, we report the best measurement across libraries (by plasmid coverage across variants) chosen per variant per cell type.
For saturation mutagenesis, we used the saturation-mutagenesis mode of MPRAmatch, which applies increased stringency requirements when pairing barcodes to oligonucleotide sequences requiring perfect alignment matches during barcode pairing. Because most of the saturation-mutagenesis library consisted of sequences with strong MPRA activity by design, we used a summit-shift normalization procedure where linear scaling factors were determined only from negative control sequences. Because the primary goal of the saturation-mutagenesis experiment was to compare the effects of substitutions across an element, rather than to detect individual emVars, we used an empirical Bayes approach to shrink the allelic effect sizes towards 0, based on how accurately they were measured (the number of RNA and DNA reads) (Supplementary Fig. 8c,d). First, we obtained an improved estimate of the baseline activity of the background sequence by pooling the log2(fold change) estimates of oligonucleotides between the observed 25th and 75th percentiles of substitutions across this element, assuming that approximately 50% of substitutions will have little effect and the mean activity of these substitutions is an unbiased estimate of true baseline activity. Taking the mean and variance of this set as a normal prior, we used DESeq2 estimates of activity (log2[fold change]) and their corresponding standard errors to define a normal likelihood, allowing for closed-form posterior inference, owing to conjugacy. We used this prior and likelihood to obtain a posterior distribution for baseline activity of the background sequence. Next, we used the mean and variance of activity estimates between the 5th and 95th percentiles of substitutions to form a less-stringent prior with higher variance, better reflecting the distribution of substitutions with true allelic effects. Using a similar Bayesian approach with this prior, we estimated the posterior activity distribution of each substitution. To obtain shrunken estimates of each allelic effect, we subtracted the posterior mean activity for each substitution from the posterior mean baseline activity for the background sequence.
Grid search for activity and allelic effect thresholds
To find the optimal thresholds for activity and for allelic effect, we did a grid search across Bonferroni-adjusted P value or FDR and effect magnitude, aiming to maximize precision and recall in a set with known positives and negatives. For activity, we used overlap with a CRE to approximate our positive set, and assigned all other oligonucleotides to the negative set, reasoning that sequences originating from accessible chromatin marked by H3K27ac should approximate active transcriptional regulatory elements. We selected sets of variants in CREs and not in CREs, and calculated the precision and recall across a grid of Bonferroni-adjusted P value and activity (log2[fold change]) thresholds (Supplementary Fig. 3a,b). For activity grid searches, to compare cut-offs using different annotations to separate positive and negative controls, we did not scale the sets 1:1. Setting |log2[fold change]| > 1 and Bonferroni-adjusted P value < 0.01 optimized the precision–recall trade-off and is used as the definition of active throughout the manuscript. For allelic effects, we repeated our grid search, using sets of high PIP (more than 0.9) and low PIP (less than 0.02) variants as positive and negative sets. We scaled the larger, negative sets down to the size of the positive set using the false-positive rate. First requiring all elements to be active, we calculated precision and recall across a grid of FDR thresholds and allelic effect (∆log2[fold change]) thresholds (Supplementary Fig. 4a,b). Based on these results, we defined emVars as variants residing in active elements with any magnitude of allelic effect and an FDR < 10% for a non-zero allelic effect.
Non-additive effect estimation and diplotype analyses
Because alternative and reference alleles for trait-associated variants are arbitrarily coded based on the reference genome, rather strictly by effect direction, disease risk or minor allele, we recoded the alleles to best aid the interpretation of MPRA effects. Using MPRA estimates of activity for each of the four diplotypes, we recoded alleles in order from lowest to highest activity, with lowest activity as the reference category. We set the reference category as the ‘decrease–decrease’ pair and the highest value as the ‘increase–increase’ pair, with ‘decrease-increase’ and ‘increase-decrease’ pairs in between. The additive allelic effect is defined as the sum of the individual allelic effect (decrease–increase + increase–decrease). The observed double-allele effect is the allelic effect for the increase–increase pair compared with the decrease–decrease pair. Interaction effects are the difference between the expected additive allelic effect and the double-allele effect, which are estimated in the DEseq2 model and recoded here. Interacting pairs of variants were detected as described below and classified into amplifying or dampening pairs (Fig. 3b,d, Extended Data Fig. 5i and Supplementary Tables 20 and 21).
To gain power to detect interaction effects between variant pairs, we performed several meta-analyses. First, we did fixed-effect meta-analyses to obtain both activity and allelic-effect estimates for each pair. Meta-analyses were done by aggregating across available windows (within a single cell type) or by aggregating across both windows and cell types. In these meta-analyses, activity and emVar definitions were the same as for the meta-analyses of single-variant elements but with the consideration of additional alleles; an element was considered active if any of the four elements (versus two elements for single-variant) were active. Interaction effects were also meta-analysed across windows within a single cell type and windows across all cell types using both a standard covariance matrix and an empirical covariance matrix.
For the meta-analyses with a standard covariance, the observed variance of each interaction-effect estimate was placed on the diagonal and zeros were placed elsewhere, encoding the hypothesis that windows are independent. We reasoned that windows might not be independent, and if we could capture this relationship in the covariance matrix, we would increase the power to detect significant interaction effects in meta-analyses. To estimate the relationship between windows, of which each variant pair could be tested in up to six (centring on variant 1 or variant 2; windows around the centred variant shifted left, middle or right), we calculated the expected correlation between these effects based on two factors that could influence these effects and how they covary: how much of the 200-bp oligonucleotide was shared, and the distance between the variants in the pair. Specifically, we split the haplotype-interaction effect measurements into a 7 × 9 grid of 7 bins of shared oligonucleotide percentage (0%, 25%, 50%, 60%, 70%, 80%, 95% and 100%) and 9 bins of variant distance in nucleotides (0, 10, 15, 20, 25, 50, 75, 100 and 200). In each bin, we then computed the correlation between observed interaction effects to obtain an empirical correlation matrix, and used a generalized additive model to smooth the correlation matrix across the grid (Extended Data Fig. 5g). The covariance matrix was obtained by scaling the correlation matrix by the variance vector, and fixed-effect meta-analysis was done using this covariance matrix.
Interaction emVars were defined as pairs in which at least one diplotype was an emVar (versus the Ref/Ref diplotype) and the FDR was less than 0.05 and the absolute value of the interaction effect was greater than 0.25 for one of the 4 meta-analyses (standard versus empirical covariance, across windows in cell types versus across windows and cell types). In the saturation-mutagenesis dataset, interaction-effect estimates were similarly obtained by subtraction of the double allele and additive variant posterior activity estimates, and interaction emVars were defined as having an interaction effect greater than 0.415 (a 25% change in activity). Analyses of the distance between pairs of variants was restricted to pairs in which variants were tested in all three windows (left, middle and right) because the number of windows a variant pair is tested in is dependent on the distance between the variants (Fig. 3c).
Multivariate adaptive shrinkage
To account for differences in power when estimating whether element activity or allelic effects are shared across cell types, we used the Multivariate Adaptive Shrinkage in R (mashr) package79. Because MPRA was done in four cell types for complex trait variants a four cell types for molecular trait variants, we used only the canonical covariance matrices in the mixture model. We selected variants with PIP > 0.1 that are in CREs and fit the mash model separately for complex trait variants and eQTLs separately. Cell-type-specific effects were determined using the local false sign rate (lfsr) < 0.05 (Supplementary Table 7).
Coding and non-coding genomic annotations
Genomic annotations were obtained as described previously31,66. In brief, we ran the Ensembl Variant Effect Predictor v.85 (ref. 80), selecting the most severe consequence for each variant including coding (missense, loss of function and splice site) and UTR annotations. Non-coding variants were annotated as residing within a promoter (using annotations from the S-LDSC baseline model), a CRE or having evidence of neither. CREs were defined as accessible chromatin in at least one cell type and evidence of histone modification at H3K27ac in at least one cell type. We used the combined, normalized and quality-controlled chromatin accessibility and histone modification datasets described in ref. 66, including DNase-seq, ATAC-seq, single-cell ATAC-seq and H3K27ac measurements from seven different atlases9,40,41,81,82,83,84,85. This combined dataset consists of more than 1,000 cell types: ROADMAP Epigenomics DNase-seq across 39 cell types and H3K27ac across 98 cell types81; DNase-seq across 438 cell types9; single-cell ATAC-seq across 54 cell types82; ATAC-seq data for 18 haematopoietic cell types83; single-cell ATAC-seq across 24 brain cell types84; ATAC-seq for 25 immune cell types85; and ChIP-Atlas DNase-seq for 284 cell types and H3K27ac for 720 cell-types41 to identify CREs. Non-coding CSs were defined as 95% CSs where no variant was annotated as a missense, loss of function or splice-site variant.
For analyses comparing the strength of endogenous CRE activity with MPRA activity (Fig. 1d), only CREs from the dataset in ref. 9 are used and cell-type agnostic CRE activity is estimated as the maximum normalized DNase I hypersensitivity counts across cell types. To identify cell-type-specific CREs, we computed the euclidean norm of normalized counts from the dataset in ref. 9, selected the maximum value across biosamples representing similar cell types, and selected CREs with a euclidean norm greater than 0.2. Furthermore, we obtained representative tissue and organ system annotations provided in ref. 9. In CREs from this dataset9, variants were annotated for consensus DHS footprints across 243 cell types, which were downloaded from ref. 47 and lifted over to hg19.
Precision and recall of MPRA
We calculated the precision and recall of different annotations (such as emVars) for separating trait-associated variants from background variants. We approximated the true causal (positive) set of variants by using non-coding, trait-associated variants with PIP > 0.9. Background non-coding variant (negative) sets were defined using either location-matched controls, annotation-matched controls or trait-associated variants with PIP < 0.01. For the eQTL dataset, variants with PIPs up to 0.02 were included as low-PIP to have sufficient matches. We scaled the larger, negative sets down to the size of the positive set using the false positive rate to create a 1:1 set. Precision is defined as the number of true positives divided by the number of variants selected as positive by the annotation. Recall is defined as the number of true positives divided by all positives. All three control sets produced similar estimates, and results using low-PIP variants as controls have the lowest standard errors and are used in the main analysis shown in Fig. 2e. As well as evaluating MPRA measurements and genomic annotations, we estimated precision and recall across CS size (Extended Data Fig. 4e and Supplementary Table 11) and specific cell types used in MPRA or for defining CREs (Extended Data Fig. 4f,g and Supplementary Table 10). For cell-type analyses, meta points were estimated as averages of cell-type combination, and linear regression was used to predict the impact of each additional cell type on precision and recall.
Comparison with genome-wide reporter assays
Measurements of allelic effects using the Survey of Regulatory Elements (SuRE) reporter assay for 5.9 million variants in K562 and HepG2 cells were downloaded from ref. 44. To compare performance between the Survey of Regulatory Elements and MPRA, we again estimated precision and recall using PIP > 0.9 variants as positives and PIP < 0.05 variants as negatives, but now restricted to variants assayed by both approaches.
Regulatory quantitative trait loci
Chromatin accessibility quantitative trait loci and allele-specific TF occupancy quantitative trait loci were obtained from ENCODE9,86. Variants with an FDR < 0.25 for allele-specific differences were included in comparisons with MPRA allelic effects.
TF binding motifs and occupancy
Positional motif matches on oligonucleotides were identified using a modified version of FIMO (MEME Suite 5.5.5)87 implemented in motif liquidator88 for PWMs from the Homo Sapiens Comprehensive Model Collection (HOCOMOCO, H12CORE89) and JASPAR90 (2022 release). Motifs shorter than 8 bp or with an information content of less than 12 were excluded. Sequence matches were called when the absolute percentage (scaled by the difference between the best and worst possible sequence matches scores) and the relative percentage (scaled by the difference between the best and random sequence matches scores) of each sequence was greater than 12, and the significance for this match was less than P = 0.0001. Disruption of a motif required a difference of more than 10% in the absolute percentage of each sequence match to the PWM. Weak calls with disruptive alleles required only an information content score of more than 8, no relative percentage filter, P < 0.001 and an allelic difference of only 5% in the absolute percentage of each sequence. Motifs were defined as flanking variants if the motif started or ended within 10 bp of the variant (Fig. 2f).
TF occupancy for tested elements was determined by ChIP–seq support. We obtained ChIP–seq peaks from ChIP-Atlas, which uniformly reprocessed peaks from 25,823 experiments in 1,046 human cell types or tissues for 1,741 TFs40,41. TFs were retained in analyses if they met a minimum number of overlaps with variants or elements (as specified in the text). For each specific motif match or disruption, a corresponding TF was defined as occupying an element if that element overlapped with a TF peak for any TF that matched a highly similar motif (Pearson r > 0.9 for PWM similarity). For example, 1,513 TFs occupied at least 20 elements, and we observed that, of these, 1,355 were more likely to overlap active elements at a Bonferroni-adjusted P < 0.05 (Extended Data Fig. 2b). Furthermore, 54 ‘dark TFs’, or TFs that bind to closed chromatin, were obtained from ref. 91.
Motif contributions to the activity of elements derived from CREs containing trait-associated variants were estimated using linear regression, modelling the marginal effect of the number of motifs in each element on the activity in K562, HepG2 and SK-N-SH cells. Before fitting the linear regression, activity was quantile normalized across cell types and a generalized additive model using smoothing splines of 1-mer and 2-mer counts in each element to model activity (each cell-type measurement was an independent observation) was fit using the mgcv R package, and residual activity was used as the outcome for linear regression. P values for each motif were obtained from a Wald test and FDRs were estimated using Storey’s q value. Coefficients from these regressions for motifs present in more than 0.1% of elements are shown in Fig. 1e, although these estimates should not be considered causal effects. We also assessed whether changing the extent of PWM disruption for each motif was correlated with the extent of change in activity in each cell type. Restricting to variants that are emVars in at least one cell type and motifs represented by at least 10 variants, we computed Spearman’s correlations and estimated FDRs using Storey’s q value.
Regulatory sequence classes
We obtained regulatory sequence (Sei) classifications, including polycomb and heterochromatin annotations, and lifted the annotations over to hg19 (ref. 43). Variants were excluded if they fell into multiple Sei annotations after liftover (less than 1%). The proportion of emVars for each complex trait, stratified by domain, is estimated and shown in Supplementary Fig. 6a.
Enformer
To capture more regulatory mechanisms than those mediated through TF motifs catalogued in JASPAR and HOCOMOCO, we scored each variant (after lifting over to hg38) using Enformer48, a transformer-based neural network model trained to predict functional genomic data, including TF occupancy and chromatin accessibility. Allelic-effect predictions across 1,447 distinct tissue–TF pairs and 320 accessible chromatin predictions were z-score normalized to variants with non-emVars in CREs with PIP < 0.02. A combined score for each variant was computed by squaring its allelic effect for individual TF occupancy or accessible chromatin predictions and taking the sum. Thresholds were set separately for TF occupancy and accessible chromatin, each determined such that only 5% of non-emVars in CREs with PIP < 0.02 had higher scores. Thresholds for individual Enformer predictions for all variants were similarly obtained. A variant was considered disrupted by Enformer if it exceeded the threshold for either the TF occupancy or chromatin accessibility. Using these thresholds, 7.3% of non-emVars in CREs with PIP < 0.02 were classified as disrupted (Fig. 2f).
Comparison of eQTL and MPRA effect sizes
The MPRA allelic effects were compared with eQTL allelic effects by computing their Spearman’s correlation coefficients. Variants were restricted to emVars with PIP > 0.5 for at least one gene, and MPRA effect sizes were obtained from the meta-analyses across cell types. The eQTL effect sizes were obtained from the most significantly associated gene and tissue for each variant. To ensure a clear distinction between proximal and distal regulatory elements, distal CREs were further required to be more than 10 kb from the TSS of the associated gene, and proximal CREs were required to be less than 2 kb from the TSS of the associated gene. To compare agreement in sign, a binomial test that the proportion of variants agreed in sign more than expected (0.50) was performed (Pomnibus = 4.3 × 10−30). To compare agreement in magnitude, the absolute value of each effect-size measurement was taken before computing correlations (Pomnibus = 7.1 × 10−15). To control for background at distal CREs, correlations were computed for a set of variants that differed only in CRE status (not in a CRE), and the obtained correlation was subtracted off to obtain the adjusted estimate.
Multiple causal variants
We designed a hypothesis test to evaluate whether our data provided evidence of multiple causal variants in independent signals of genetic association (the same CS). First, we restricted our analysis to non-coding 95% CSs where we obtained sufficient MPRA measurements on variants harbouring most of the CS probability (more than 90%). Then, we removed CSs without evidence of the functional annotation being tested (CRE emVars, CRE or emVars). We counted the total excess number of variants in each remaining CS (containing at least one variant with a functional annotation) and compared this with the background rate estimated from controls (location matched, annotation matched or low PIP), computing both risk ratios and risk differences. Because the emVar call rate has both biological and technical variance across experiments, we designed each library such that individual traits were fully contained in a single library along with all three types of matching control. Thus, risk ratios and risk differences were computed in each experiment and meta-analysed using a random-effects model implemented in the R package metafor. Meta-analysis was performed separately for complex traits and eQTLs across multiple CS sizes (5, 10 and 15) and r2 thresholds (0.8, 0.9 and 0.99).
Single-TARV CS selection
Across all traits, we identified 95% CSs with only one TARV that lacked other ‘interesting’ variants (Supplementary Table 15). We reasoned that the TARV in these CSs were enriched for being the likely causal variant. To obtain this set, we first excluded CSs containing any coding variants, as well as those consisting of only a single variant. We then filtered to CSs containing a single TARV and removed CSs when any non-TARVs in that CS were not evolutionarily constrained, not in a 3′ or 5′ UTR. As an extra precaution, we also excluded CSs containing variants in promoters (not annotated as CREs) and variants that were emVars.
CRE-to-gene annotations
To better contextualize the variant effects identified in our MPRA, we annotated all autosomal TARVs using multiple CRE-to-gene prediction datasets:
-
PCHi-C in 27 diverse cell types92
-
PCHi-C in 17 primary haematopoietic cell types93
-
CRE–promoter correlation in diverse 127 cell types94
-
CRE–promoter correlation in diverse 808 cell types95
-
CRE–promoter correlation in 16 primary haematopoietic cell types28
-
E2G models trained on CRISPRi perturbations (E2G)96.
TARVs were overlapped with each dataset to generate a compendium of CRE–gene links, as and nearest gene annotations (Supplementary Table 16). These results include the following gene connections for TARVs highlighted in our study:
rs6536864738 (Fig. 4d,e): fine-mapped in UKBB for platelet count (PIP = 1.0). Target genes identified include the nearest gene KDM7A96, as well as UBE3C93, PARP12 (ref. 96), SSBP1 (ref. 93), MKRN1 (ref. 96), RAB19 (ref. 93,96), SLC37A3 (ref. 93,96) and KLRG2 (ref. 96).
rs11864973 (Fig. 4f,g): fine-mapped in BBJ for red blood cell count (PIP = 0.99). PCHi-C analysis identified 16 putative target genes, including the nearest gene NPRL3. Of these, several encode haemoglobin subunits: HBQ1 (refs. 94,95), HBA1 (ref. 28), HBA2 (ref. 28) and HBM28,94,95.
rs2529369 (Fig. 4i,j): fine-mapped in GTEx tibial nerve (PIP = 1.0) and associated with CDHR3 expression. This target is confirmed by PCHi-C93. Other targets include the nearest gene ATXN7L1 (ref. 93) and EFCAB10 (ref. 96).
rs191148279 (Fig. 5d–f): fine-mapped in UKBB for HbA1c levels (PIP = 0.919). PCHi-C identifies C2orf88 as a potential target28,96, which we confirmed through endogenous editing experiments. Other putative targets include SLC40A1 (ref. 28) and INPP1 (ref. 96).
rs7282770 and rs7282886 (Fig. 5h,i): located in the same CS in GTEx, associated with DOP1B expression in liver (PIP = 0.08), the loci shows target gene contacts to DOP1B in the PCHi-C and E2G datasets. Other targets include CBR3 (ref. 96) and CHAF1B92.
rs35081008 and rs34003091 (Extended Data Fig. 8d,e): located in the ZNF329 promoter and associated with ZNF329 expression in liver and LDL cholesterol levels. Both variants contact ZNF329 (refs. 94,96) and 83 other genes (Supplementary Table 16).
Activity blocks
We developed a Gaussian filter approach to identify blocks of quasi-contiguous non-zero variant positions. Gaussian filters are widely used across fields such as image processing, in which they detect edges by reducing pixel noise; astronomy, in which they enhance features such as stars and galaxies against noisy backgrounds; and audio processing, in which they reduce high-frequency noise while preserving signal integrity. By smoothing data, Gaussian filters reduce noise while preserving localized features and highlighting important patterns in the signal. This approach emphasizes features by weighting nearby positions more heavily and provides a smooth transition between regions of high and low signal. Because of this, they are well suited to identify TF-binding sites and other regulatory sequence segments in which sets of contiguous variant positions all participate in transcriptional regulation.
To identify blocks, we used the crossover points between a threshold T ∈ ℝ and a smoothed signal S = [Sk ∈ ℝ] (at position k) of the ∆log2(fold change) [Δk] of an element as follows. For any given sequence element, each Δk was obtained by averaging all three posterior allelic substitution effects at each position k. Then, S was obtained by applying a one-dimensional Gaussian filter (scipy.ndimage.gaussian_filter1d) to [Δk] with a kernel s.d. of σ = 1.15. To identify blocks of negative allelic effects, which indicate positive contributions to activity by the reference sequence, we found the crossover indices ci ∈ ℕ between S and T = −0.2, and considered as salient those regions [ci, ci+1] such that:
-
i)
Sk ≤ T Sk ≤ T = −0.2 ∀ k ∈ [ci, ci+1]; that is, the signal S along the block is below the threshold;
-
ii)
ci+1 − ci + 1 ≥ 5; the length of the region is at least 5;
-
iii)
mean(Δk, k ∈ [ci, ci+1]) ≤ −0.15; the mean of the allelic effects has to be at most 0.15.
Similarly, we took T = 0.2 and mean(Δk, k ∈ [ci, ci+1]) ≥ 0.15 to identify blocks of positive allelic effects.
Motif matches in activity blocks
To find a TF motif match to a given activity block b = [bstart,bstop], we constructed a PWM from the allelic effects as follows. Let δak denote the three allelic effects at position k ∈ [bstart,bstop] between the reference allele r ∈ A,C,G,T and the alternative alleles A,C,G,T ∋ a ≠ r. We define a matrix Mtk, where t ∈ A,C,G,T and k ∈ [bstart,bstop], as Mtk = δak if t = a, Mtk = 0 if t = r. Then, we define PWM[bstart,bstop] = 20 × Mtk/Mmax for t ∈ A,C,G,T and k ∈ [bstart,bstop], as the PWM induced by the allelic effects in the activity block [bstart,bstop], where 20 is a chosen temperature parameter and Mmax = maxt(∑k|Mtk |). Using PWM[bstart,bstop], we construct the block position probability matrix, PPM[bstart,bstop], by applying the Softmax function to the values of PWM[bstart,bstop] at each position. We also define the block information content matrix, ICM[bstart,bstop], by mapping the values of PPM[bstart,bstop] at each position into information bits with respect to the uniform background pA = pC = pG = pT = 0.25. When we had obtained the matrices PWM[bstart,bstop] and ICM[bstart,bstop] from the allelic effects in an activity block b, we sought to find their best match to a human TF motif in JASPAR90 or HOCOMOCO89. Let \(\mathcalD\) represent the dataset of known human TF motifs in the union of JASPAR (2022 REDUNDANT) and HOCOMOCO (H12CORE). Let PWMm, ICMm and λm represent the position weight matrix, information content matrix and length, respectively, of a motif \(m\in \mathcalD\). We defined the best alignment offset o(b, m) ∈ ℤ between a block b = [bstart,bstop] and a motif \(m\in \mathcalD\) as the integer that maximizes the sum of the element-wise product between ICM[bstart,bstop] and PWMm for all possible alignments of the nucleotide positions between the two matrices (with the appropriate zero padding to match the dimensions of the two matrices). Next, we computed the (block) Pearson correlation between ICM[bstart,bstop] and ICMm (with the appropriate zero padding) in their best alignment given by o(b, m) for every m in \(\mathcalD\), and collected the motifs in the top 50 Pearson correlations. Then, for those top 50 motifs, we computed the (match) Pearson correlation between ICMm and ICM[bstart + o(b, m), bstart + o(b, m) + λm]; that is, the correlation to the ICM allelic effects given the alignment induced by the alignment to the activity block. Finally, we selected the motif with the highest match_Pearson + 0.2 × block_Pearson. If the overlap between the selected TF match and the activity block leaves five or more contiguous positions in the block unassigned, we repeat the process above to the unassigned sub-block.
Rare-variant analysis
Whole genome sequencing for 402,990 unrelated (max kinship coefficient < 0.884) individuals without filtering on genetic continental ancestry in the UK Biobank was analysed on DNAnexus. Carriers were defined as single-nucleotide substitutions with a higher MPRA allelic effect size than rs191148279 from the saturation-mutagenesis experiments in K562 cells. Residual HbA1c phenotypes were obtained as previously described31, and an odds ratio using Firth’s correction for more than 1 s.d. change in the phenotype was computed as well as Levene’s test, which was run to test whether the variances were different between carriers and non-carriers.
GWAS meta-analysis
We selected 121,247 individuals of African continental ancestries from UKBB (n = 5,803), All of Us (n = 7,442), Million Veterans Project (n = 98,427) and Alliance for Genomic Discovery (n = 9,575) with genotype calls at rs785206 and HbA1c level measurements. In brief, linear regression in UKBB and Alliance for Genomic Discovery was done as previously described31; linear regression in All of Us was done controlling for age, sex and three genetic PCs; and linear regression results from Million Veterans Project were obtained from ref. 32 (HARE, A1C_Min_INT). Fixed-effect meta-analysis was done using the R metafor package. Bonferroni-adjusted 95% confidence intervals were obtained using the critical value corresponding to P < 5.0 × 10−8 (5.33).
GTEx interaction analysis
Sex-biased eQTLs were mapped as described in ref. 97. In brief, a linear model with a genotype × biological sex interaction term was used to model normalized gene expression. Genotype PCs and expression PEER factors were included as covariates. The interaction P value was obtained from a Wald test.
Constraint
Nucleotide-level PhyloP scores for constraint in 427 mammals were downloaded from Zoonomia4. Saturation-mutagenesis sequences were lifted over to hg38 and matched to PhyloP scores. Specific positions were defined as constrained if the PhyloP FDR was less than 0.05, corresponding to a PhyloP score of more than 2.27. Before estimating the correlation with PhyloP, ∆log2(fold changes) for all three substitutions at each position were averaged and inverted to obtain a contribution score, similar to the height of letters in the saturation-mutagenesis examples in Figs. 4 and 5. Correlations between either the signed contribution score or the |contribution score| and PhyloP using Pearson’s method and corresponding FDRs obtained using Storey’s q value are provided in Supplementary Table 26.
Endogenous base-editing
The K562 cells were cultured in RPMI 1640 medium, GlutaMAX Supplement (Life Technologies, 61870127) with 10% FBS (Life Technologies, 26140). For base-editing, 1 M cells were transfected in a 10-μl volume with 1 μg of a 3:1 mass ratio pool of base-editing plasmid, BE4 (Addgene, 100802)98 and sgRNA-expressing plasmid, BPK1520 (Addgene, 65777)99, using a Neon transfection system (Life Technologies) (3 pulses of 1,450 V for 10 ms). Two sgRNAs were used to target the centre GATA motif and transfected independently (AGTACTATCTTTGAGGATAG, AAGTACTATCTTTGAGGATA). Transfected cells were grown for 6 days followed by removal of 5 × 105 cells for gDNA extraction and PCR amplification of the target site (rs191148279_gDNA_F, rs191148279_gDNA_R; Supplementary Table 31) using Q5 Polymerase (NEB) to confirm presence of edits by Sanger sequencing. Cells were grown for a further 96 h, then individual cells were isolated by limiting dilution cloning, plating at 0.3 cells per well in 384-well plates to yield single-cell colonies in approximately one-third of wells. Single cells were expanded followed by gDNA extraction on an aliquot of cells using QuickExtract DNA Extraction Solution (Biosearch Technologies, QE09050). DNA was amplified using Q5 Polymerase (NEB) and the same primers for bulk verification followed by Sanger sequencing. Nine clones containing the rs191148279 G>A knockout mutation with a second silent G>A mutation outside GATA and 11 unedited clones with the wild-type allele containing the GATA site were selected for downstream analyses.
Expression profiling of base-edited clones
RNA was extracted from around 106 cells taken from K562 base-edited and unedited clones using Qiagen RNeasy Mini Columns (Qiagen, 74104). BRB-seq libraries were prepared as described previously100. Specifically, 100 ng (quantified on Qubit) of RNA was incubated with the BU3_index primer (Supplementary Table 31) for 10 min at 65 °C and placed on ice before performing the room-temperature reaction (14 μl RNA + primer mixture, 4 μl 5× First Strand Buffer, 2 μl 0.1 M DTT and 0.25 μl Superscript II Enzyme (ThermoFisher, 18064014)) (42 °C for 50 min, 70 °C for 15 min). Individual samples were then pooled and purified (Monarch Spin PCR & DNA Cleanup Kit, NEB T1130) before eluting in 20 μl water. Samples were then treated with exonuclease I (20 μl cDNA, 1 μl exonuclease I, 2 μl 10× reaction buffer (NEB, MO293), 30 °C for 30 min, 80 °C for 20 min) before second-strand synthesis. Second-strand synthesis was done by nick translation in the following reaction: 23 μl exonuclease-treated cDNA, 2 μl RNase H (NEB, M0297), 1 μl Escherichia coli DNA ligase (NEB, M0205), 5 μl E. coli DNA polymerase (NEB, M0209), 1 μl 0.2 mM dNTP (ThermoFisher, R0192), 10 μl 5× second strand buffer (100 mM Tris-HCl, pH 6.9, 25 mM MgCl2, 450 mM KCl, 0.8 mM β-NAD, 60 mM (NH4)2S04) and 11 μl water; overnight incubation at 16 °C. Tagmentation was then performed with 50 ng second strand synthesized sample in the following reaction: 10 μl 2× tagmentation buffer (Diagenode, C01019043), 9 μl DNA, 1 μl Tagmentase (Diagenode, C01070012-30), incubation at 55 °C for 7 min. Tagmented samples were purified (Monarch Spin PCR & DNA Cleanup Kit, NEB T1130) and resuspended in 20 μl water before final library amplification: 20 μl tagmented DNA, 2.5 μl 5 μM P5 BRB Primer, 2.5 μL 5 μM BRB Idx7N5 primer, 25 μl NEBNext High-Fidelity 2× PCR Master Mix (NEB, M0541) with the following thermocycler conditions: 3 min at 72 °C, 30 s at 98 °C, 15×(10 s 98 °C, 30 s 63 °C, 30 s 72 °C), 5 min at 72 °C. Amplified libraries were double SPRIselect purified (0.5×, 0.8×, Beckman, B23319) to select for 200–800-bp fragments before confirmation on an Agilent TapeStation with D1000 High Sensitivity ScreenTape (Agilent, 5067-5584).
Libraries were sequenced on a NextSeq 2000 sequencer with a P3-100 cycle kit (Illumina) with 21-bp read 1 using a custom primer (BRB_Ilmn_read_1; Supplementary Table 31), and 71 bp for read 2 with the standard primer. Sequenced reads were aligned with STAR (v.2.7.11b, –outFilterMultimapNmax 1)101 and demultiplexed with BRB-seq Tools (v.1.6.1)100. Count tables from BRB-seq Tools (CreateDGEMatrix) were used as input for differential expression analysis with DESeq2 (v.1.48.1)102 and significant genes (Wald test) were identified at an FDR threshold of 0.01 (Benjamini–Hochberg procedure). BRB-seq results for C2orf88 and HIBCH were confirmed using quantitative PCR. Total RNA (1–2.5 μg) from the same extractions was converted to cDNA using SuperScript III, and qPCR was done using Luna Universal Master Mix on a QuantStudio 5 with gene-specific primers for C2orf88 (C2orf88_F and C2orf88_R; Supplementary Table 31), HIBCH (HIBCH_F and HIBCH_R; Supplementary Table 31) and TBP as the normalizing control gene (TBP_F and TBP_R; Supplementary Table 31). Two independent experiments with four technical replicates each were done, and fold changes were calculated using the ΔΔCt method normalized to TBP and reference allele expression. Statistical differences were assessed using a two-sided Mann–Whitney U test (Extended Data Fig. 7c).
Data visualization and figure generation
All plots were generated in R using ggplot2 (v.3.5.0) and assembled with patchwork (v.1.2.0) and cowplot (v.1.1.3). Heatmaps were created using ComplexHeatmap (v.2.2.0). Colour palettes were from BuenColors (v.0.5.6) and PNWColors (v.0.1.0). Rasterization was done using ggrastr (v.1.0.2) and marginal distributions added using ggExtra (v.0.10.1).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.