Mice
All mice used for experiments were adult females from a mixed background, housed under standard laboratory conditions and receiving food and water ad libitum. All experiments were performed in accordance with the guidelines of the Animal Welfare Committees of the Netherlands Cancer Institute and KU Leuven. Sample size was determined using a resource equation approach, mice were randomly assigned to experimental groups and blinding was performed during data analysis. R26R-Confettihet (JAX stock no. 013731)61,62; R26-CreERT2het (JAX stock no. 008463)63 mice were injected intraperitoneally with tamoxifen (Sigma-Aldrich), diluted in sunflower oil, to activate Cre recombinase. To achieve clonal density labelling (fewer than one MaSC per duct on average), R26R-Confettihet;R26-CreERT2het mice were injected with 1âmg of tamoxifen per 25âg of body weight between 10 and 15âweeks of age. Ovariectomies were performed between 10 and 15âweeks of age, at least 7âdays before lineage tracing initiation. The third, fourth and fifth mammary glands of R26R-Confetti;Brca1fl/fl;Trp53fl/fl (refs. 31,64) or R26R-Confetti mice were intraductally injected with recombinant TAT-Cre protein (20âunits per gland diluted in 20âµl of PBS, Sigma-Aldrich) between 10 and 15âweeks of age. This TAT-Cre injection resulted in roughly one labelled cell for every 100â200 cells (Extended Data Fig. 4e). As the confetti construct comprises four distinct colours, there is, on average, one cell labelled with a confetti colour per 400 cells. Considering that a MaSC-progeny unit consists of roughly five to ten cells, a single confetti-labelled cell is induced in one out of 40â80âunits. Over time, many clones become extinct, leading to a dilution in the number of clones and making collisions even less likely. For each of the experiments, mice were analysed at different time points after lineage tracing initiation as indicated in Fig. 1b. The injected mammary glands of R26R-Confetti;Brca1fl/fl;Trp53fl/fl mice at the latest time point (225âdays) were analysed when one of the injected glands developed a palpable tumour of at least 5âÃâ5âmm, which was between 200 and 250âdays after recombination. Tumour sizes did not exceed 1,500âmm3 in accordance with the guidelines of the Animal Welfare Committee of the Royal Netherlands Academy of Arts and Sciences, the Netherlands Cancer Institute and KU Leuven. Samples were randomly allocated to the experimental groups, sample size was not determined a priori and investigators were not blinded to experimental conditions, except where indicated. For clonal analysis of the R26R-Confetti;Brca1fl/fl;Trp53fl/fl model, we analysed nâ=â4 mice (14âdays), nâ=â4 mice (64âdays), nâ=â5 mice (120âdays) and nâ=â6 mice (225âdays). For clonal analysis of the R26R-Confetti model, we analysed nâ=â3 mice (14âdays), nâ=â3 mice (64âdays), nâ=â5 mice (120âdays) and nâ=â6 mice (225âdays). Clonal analysis of the ovariectomized R26R-Confetti;Brca1fl/fl;Trp53fl/fl and R26R-Confettihet models was performed on at least nâ=â3 mice per time point. Adult CAG;;KikGR female mice65 (RIKEN no. CLSTCDB0201T-117830853340) were used to visualize the short-term dynamics of the mammary gland by repeated imaging through a mammary imaging window as described below. R26-mTmG female mice (JAX no. 007676)66 were used to visualize the long-term stability of the mammary gland through a repeated skin-flap procedure as described below.
Mammary imaging window implantation, repeated skin-flap procedure and intravital imaging
Mice were anaesthetized using isoflurane (Isovet) inhalation (1.5/2% isoflurane/air mixture). The fourth mammary gland of adult CAG;;KikGR female mice was imaged repeatedly through a mammary imaging window as previously described47,48. The fourth mammary gland of adult R26-mTmG mice was imaged repeatedly with a skin flap as previously described47. To visualize the mammary gland, mice were placed in a facemask within a custom designed imaging box. Isoflurane was introduced through the facemask and ventilated by an outlet on the other side of the box. The imaging box and microscope were kept at 34â°C by a climate chamber surrounding the entire stage of the microscope including the objectives. Imaging was performed on an inverted Leica SP8 Dive system (Leica Microsystems) equipped with four tuneable hybrid detectors, a MaiTai eHP DeepSee laser (Spectra-Physics) and an InSight X3 laser (Spectra-Physics) using the Leica Application Suite X (LAS X) software. All images were collected at 8âbit and acquired with a Ã25 water immersion objective with a free working distance of 2.40âmm (HC FLUOTAR L Ã25/0.95W VISIR 0.17). For the CAG;;KikGR model, Kikume Green was excited at 960ânm and detected at 490â550ânm. Each imaging session, all visible ducts through the imaging window were imaged using a tiled z scan with Ã1â2 zoom and a z-step size of 5â10âμm. For the skin-flap imaging, TdTomato was excited at 1,040ânm and detected at 540â730ânm. All visible ducts were imaged together in one tile z scan with a Ã0.75 zoom, a z-step size of 10â20âμm. These parameters allowed to scan large regions of up to 2âcm2 in less than 3âh. At the end of the first skin-flap imaging session, the skin was closed with a continuous, non-resorbable suture. After 3âmonths, the skin flap was re-opened for the second imaging session, and the same imaging fields were retraced using the nipple and collagen I structures of the first imaging session as landmarks.
Staging of the mice
To determine the oestrous cycle stage of the mice, a vaginal swab was collected as described67. In short, the vagina was flushed using a plastic pipette filled with 50âµl PBS, and the liquid was transferred to a dry glass slide. After air drying, the slide was stained with Crystal Violet and the cell cytology was examined using a light microscope.
Clone isolation and CNA sequencing
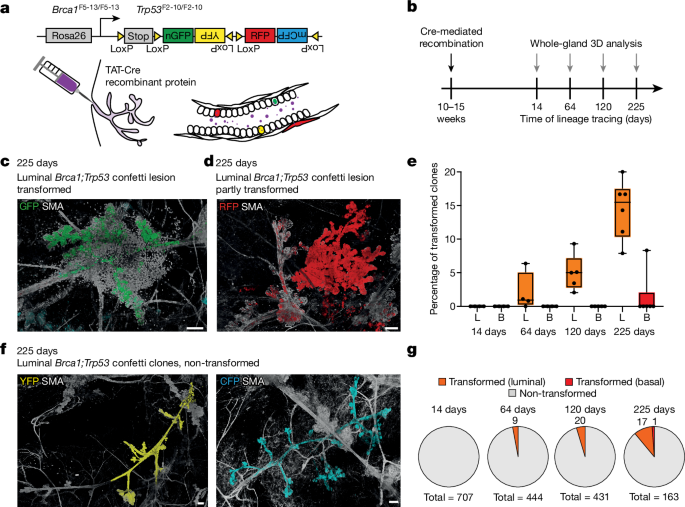
Here, 225âdays after Cre mediated recombination, fourth mammary glands were extracted, fixed overnight in 1% paraformaldehyde (PFA), incubated in sucrose overnight and stored in optimal cutting temperature (OCT) at â80â°C. For microdissection of individual clones, OCT blocks were thawed at room temperature in the dark for 30âmin and mammary glands removed from OCT and washed in 50âml of PBS on ice. Mammary glands were dissected under a benchtop fluorescent macroscope (Zeiss) using Dumont forceps and fine scissors, using the clone morphology to distinguish between transformed and untransformed clones. Each dissected clone was washed in 1âml of PBS on ice for 2â5âh (until the end of the dissection procedure). Per mammary gland, one piece of non-fluorescent tissue from the inguinal lymph node was dissected as internal sequencing control. After washing, pieces were lysed in 70âµl of Arcturus lysis buffer following the instructions of the ThermoFisher Scientific Arcturus PicoPure kit KIT0103. Lysis was carried out in a PCR cycler for 18âh at 65â°C followed by 30âmin at 75â°C and holding at 4â°C. Samples were then purified using the Roche FFPE DNA extraction kit 06650767001 50-588-384 following the manufacturerâs instructions with elution in 25âµl of PCR grade water. For DNA sequencing, library preparation was carried out with a KAPA Hyper kit (Roche; KK8504) according to the manufacturer protocol with four PCR cycles, before the samples were sequenced by low-coverage whole genome sequencing. The copy number alteration (CNA) analysis was conducted in R, using QDNAseq with 50âkb bins and the mm10 mouse reference genome. This methodology yielded copy number values from both normal and transformed clones, along with an internal control sample. Normalization was achieved by first converting copy number values to log2, then subtracting the internal control sampleâs values from those of the normal and transformed tumours. We averaged these adjusted copy number values for each replicate across both clone types (13 early normal clones and 13 early transformed clones). Data visualization was executed using the ggplot2 package in R, with specific emphasis on certain chromosomes. Regarding the late-stage tumours published in ref. 40, copy number profiling data corresponding to ten Wap-Cre;Brca1fl/fl;Trp53fl/fl (WB1P) female mice harbouring a WB1P late-stage mammary tumour, along with internal control samples (spleen) was used. The CNA sequence analysis included the use of cutadapt for adaptor sequence removal and BWA for sequence alignment (using bwa aln, bwa mem) to the mm10 mouse genome. This procedure mirrored the earlier steps up to plotting with ggplot2, repeated for ten WB1P replicates.
Quantification of the distribution of proliferation
Oestrous-cycling mice and mice that had undergone ovariectomy with a 2-week recovery period (all above 8âweeks of age) received 0.5âmgâmlâ1 EdU in drinking water (refreshed every second day) for 1âweek. 3D imaging was performed on three cycling mice and five ovariectomized mice. Per mouse, one-quarter of the mammary gland was taken for subsequent analysis. Samples were fixed in 4% PFA overnight and stained using the FLASH protocol with FLASH Reagent 2 (ref. 68). Before adding the primary antibodies, samples were stained for EdU as follows. Tissues were incubated in 5âml of 3% bovine serum albumin for 1âh, followed by three washes in PBS for 20âmin each. EdU was detected with an Alexa-647 azide. The reaction cocktail for EdU fluorescent labelling was prepared according to the manufacturerâs guidelines using the Click-It EdU imaging kit (ThermoFisher Scientific). Per gland, 0.5âml of reaction cocktail was added for incubation for 4âh at room temperature with gentle agitation on a nutator. The cocktail was removed, samples washed once in 3% bovine serum albumin in PBS for 20âmin, followed by three washes in FLASH blocking buffer for 20âmin each. Subsequently, samples were stained with primary and secondary antibodies overnight each. Primary antibodies used were KRT8 (rat, Troma-I, Merck Millipore, 1:800) and αSMA (mouse IgG2a, clone 1A4, ThermoFisher Scientific, 1:600). Secondary antibodies used were donkey antirat Alexa-488 and donkey antimouse Alexa546 (ThermoFisher Scientific, catalogue nos. A21208 and A10036, respectively, 1:400), combined with Hoechst 33342 for nucleus detection. Samples were imaged on an Andor Dragonfly spinning disc system, installed on an inverted Leica DMI8 microscope with an Andor Zyla 4+sCMOS camera using a Ã10, 0.45 NA Fluo objective (Leica). Imaging was carried out with a 40âμm disc using 405ânm excitation, a 561ânm optically pumped semiconductor laser and 637ânm diode lasers. Images were visualized with Imaris Viewer using gamma correction, ortho slicers and cutting planes to depict deeper tissue layers. For each mammary gland, distribution of proliferation was quantified in five regions. Ripley analysis using QuPath69 was performed with a custom-made script (available at https://github.com/BioImaging-NKI/qupath_ripley). The image was opened in QuPath and a freehand line was drawn by hand to outline the duct for analysis. A multipoint annotation was drawn by hand to mark the positions of proliferating cells along the duct. The script calculated Ripleys K function and normalized it to an unclustered distribution resulting in Ripleyâs L function. Data were plotted in GraphPad Prism v.10. For simulations, we have generated clustered and unclustered data in Python.
Whole-mount immunofluorescence staining of mammary glands
The third, fourth and fifth mammary glands were dissected and incubated in a mixture of collagenase I (1âmgâmlâ1, Roche Diagnostics) and hyaluronidase (50âμgâmlâ1, Sigma-Aldrich) at 37â°C for optical clearance, fixed in periodateâlysineâPFA buffer (1% PFA; Electron Microscopy Science), 0.01âM sodium periodate, 0.075âM l-lysine and 0.0375âM P-buffer (0.081âM Na2HPO4 and 0.019âM NaH2PO4; pHâ7.4) for 2âh at room temperature, and incubated for at least 3âh in blocking buffer containing 1% bovine serum albumin (Roche Diagnostics), 5% normal goat serum (Monosan) and 0.8% Triton X-100 (Sigma-Aldrich) in PBS. Primary antibodies were diluted in blocking buffer and incubated overnight at room temperature. Secondary antibodies diluted in blocking buffer were incubated for at least 6âh. Nuclei were stained with 4,6-diamidino-2-phenylindole (DAPI) (0.1âμgâmlâ1; Sigma-Aldrich) in PBS. Glands were washed with PBS and mounted on a microscopy slide with Vectashield hard set (H-1400, Vector Laboratories). Primary antibodies used were: anti-KRT14 (rabbit, Covance, PRB155P, 1:700), anti-ECAD (rat, eBioscience, 14-3249-82, 1:700), anti-oestrogen receptor (rabbit, no. 13258, Cell Signaling, 1:100), anti-progesterone receptor (rabbit, Clone SP2, MA5-14505, ThermoFisher Scientific, 1:200) and anti-SMA (mouse IgG2a, clone 1A4, Sigma-Aldrich, 1:600). Alexa Fluor 647 and Alexa Fluor 488 Phalloidin were used 1:500 (A-22287 and A-12379, ThermoFisher Scientific) and incubated together with the secondary antibodies. Secondary antibodies used were: goat antirabbit, goat antirat or goat antimouse IgG2a, all conjugated to Alexa-647 (ThermoFisher Scientific, catalogue nos. A21245, A21247 and A21241, respectively, 1:400).
Whole-mount imaging of mammary glands
Imaging of whole-mount mammary glands was performed using an inverted Leica TCS SP8 confocal microscope, equipped with a 405ânm laser, an argon laser, a diode-pumped solid-state laser 561ânm laser and a HeNe 633ânm laser. Different fluorophores were excited as follows: DAPI at 405ânm, cyan fluorescent protein (CFP) at 458ânm, green fluorescent protein (GFP) at 488ânm, yellow fluorescent protein (YFP) at 514ânm, red fluorescent protein (RFP) at 561ânm and Alexa-647 at 633ânm. DAPI was collected at 440â470ânm, CFP at 470â485ânm, GFP at 495â510ânm, YFP at 540â570ânm, RFP at 610â640ânm and Alexa-647 at 650â700ânm. All images were acquired with a Ã20 (HCX IRAPO N.A. 0.70 WD 0.5âmm) dry objective using a Z-step size of 1â5âμm (total Z-stack around 200âμm). 3D overview tile scans of the mammary glands were acquired by scanning large tile-scan areas (xyz). Next, detailed images were obtained of the individual clones. All images were stitched and processed in the true 3D real-time Rendering LAS X 3D Visualization module (Leica Microsystems) and further processed using ImageJ software (https://imagej.nih.gov/ij/).
Clonal analysis on whole-mount glands
Three-dimensional tile-scan images of whole-mount and fully intact mammary glands were used to manually reconstruct the ductal network by outlining the ducts based on the labelling by ECAD, SMA or KRT14 (between 400 and 600 tiles, Ã10 objective, Z-step size of 5â10âµm). After localization of the confetti clones in these 3D overview scans, each clone was imaged in detail with a Ã25 water objective using confocal imaging by taking a Z-stack with step size between 1 and 3âµm. On the basis of the overlap with the luminal- or basal-cell-specific labelling and cellular morphology (that is, a cuboidal shape for luminal cells and an elongated shape for basal cell), the labelled confetti cells were identified and annotated in the schematic outline of the mammary tree, including information on their confetti colour (GFP, green; YFP, yellow; RFP, red and CFP, cyan) and their identity: that is, luminal or basal. Regions in which, for technical reasons, the gland could not be visualized well were omitted from analysis (in three out of 160 glands). Clone sizes, referring to the number of cells within each clone, were determined through manual visual inspection of tissue samples, with the quantification performed by eye using detailed Z-stack images and 3D rendering of each individual clone. Using custom-made.NET software (available on request from J.v.R.), the coordinates of the branch points, and the position of the labelled cells in ducts and in ductal ends were scored. To calculate the surviving clone fraction, the total number of clones was determined for each of the indicated lineage tracing time points by analysing the entire mammary gland in three dimensions using our whole-gland imaging approach (nâ=â3 glands per time point of two individual mice). Next, the average numbers of clones identified at 64, 120 and 225âdays after lineage tracing initiation were divided by the average number of clones identified 14âdays after lineage tracing initiation resulting in the surviving clone fraction as depicted in Figs. 2d and 5g.
Mammary epithelial cell sorting and real-time qPCR
The third, fourth and fifth mammary glands of R26R-Confetti;Brca1fl/fl;Trp53fl/fl mice were intraductally injected with recombinant TAT-Cre protein (20âunits per gland diluted in 20âµl PBS, produced in-house) between 10 and 13âweeks of age. Then 120 to 180âdays after injection, mammary glands were harvested, minced and digested at 37â°C for 30âmin in a mixture of collagenase A (2âmgâmlâ1, Roche Diagnostics), hyaluronidase (300âμgâmlâ1, Sigma-Aldrich) and DNase (1âmgâmlâ1) in DMEM/F12 (Gibco). After 10âmin incubation with TripLE (Gibco) at 37â°C cells were strained through a 100âμm cell strainer (Fisher Scientific) to obtain single cells. Cells were spun down for 10âmin at 550âRCF (relative centrifugal force) at 4â°C followed by blocking for 15âmin on ice in 5âmM EDTA/PBS with 2% sterile filtered normal goat serum (Gibco). CD45-Alexa-647 (clone 30-F11, 03123, Biolegend, 1:200) and EpCAM-APC/Cy7 (clone G8.8, 118218, Biolegend, 1:200) were diluted in 5âmM EDTA/PBS with 2% normal goat serum and incubated for 30â45âmin on ice to label the immune population (CD45) and the epithelial population (epithelial cell adhesion molecule). Cells were centrifuged for 5âmin at 800âRCF at 4â°C and pushed through a 35âμm cell strainer. The FACS Aria III Special Ordered Research Product (BD Biosciences) was used to sort confetti+ and confetti cells, by applying a broad FSC/SSC gate, followed by gates excluding doublets (for the gating strategy, see Extended Data Fig. 1d). Afterwards, non-immune (AF647â; 670/30) confetti-positive (RFP+ (YG610/20), GFP+/YFP+ (BL530/30), CFP+ (V450/50)) and, separately, confetti-negative ((RFPâ (YG610/20), GFPâ/YFPâ (BL530/30), CFPâ (V450/50)) epithelial cells (APC/Cy7+; 780/60) were collected. Similarly, non-immune (AF647â; 670/30) epithelial cells (APC/Cy7+; 780/60) were collected from three R26R-Confetti;Brca1fl/fl;Trp53fl/fl mice that had not received TAT-Cre intraductally as a negative control. Data were analysed in FlowJo v.10 for the gating strategy (Extended Data Fig. 1d). Cells were spun down for 10âmin at 800âRCF at 4â°C and DNA was isolated using the PicoPure DNA extraction kit (Applied Biosciences; KIT0103) according to the manufacturerâs instructions. The same method was applied to isolate genomic DNA (gDNA) from K14-Cre;Brca1fl/fl;Trp53fl/fl mammary tumour organoids, representing fully recombined samples as a positive control. gDNA concentration was measured using the DeNovix DS-11 spectrophotometer. DNA was diluted to 50â75ângâmlâ1 and used for real-time qPCR using the SYBR Green Master Mix (ThermoFisher Scientific, catalogue no. 4309155) in a QuantStudio 6 Flex Real-Time PCR system using the primers listed in the table below. Reactions contained roughly 75âng of template gDNA and 1âµM of both forward and reverse primers in 20âµl reaction volume. Expression values were calculated by transforming deltaâdelta Ct values (2-ÎÎCt). Ribosomal Protein L38 (Rpl38) was used as a housekeeping gene. To confirm correct qPCR product amplification, 25âμl of qPCR product of the control samples (R26R-Confetti;Brca1fl/fl;Trp53fl/fl mice that had not received TAT-Cre intraductally) was loaded on an 2% agarose gel with loading buffer (Bioxline, catalogue no. BIO-37045) and a DNA ladder (Meridian Bioscience, catalogue no. BIO-33056) and run at 80âV for 1.5âh, after which the qPCR product was cut out of the gel and purified using the NucleoSpin Gel and PCR Clean-up kit (Macherey-Nagel, catalogue no. 740609.50) according to the manufacturerâs instructions, and was confirmed by sequencing using the qPCR primers.
|
Primer |
Sequence (5â²âââ3â²) |
|---|---|
|
BRCA1-ex10-FW |
TGTAACGACAGGCAGGTTCC |
|
BRCA1-ex10-RV |
ACAGAGTTTGCGGGTGAGTC |
|
P53-ex5-FW |
AAGACGTGCCCTGTGCAGTT |
|
P53-ex5-RV |
TCCGTCATGTGCTGTGACTTC |
|
RPL38_FW |
AGGATGCCAAGTCTGTCAAGA |
|
RPL38_RV |
TCCTTGTCTGTGATAACCAGGG |
Generation of Brca1;Trp53 mutant and WT organoids followed by Ki-67/CC3 staining
The fourth and fifth mammary glands of R26R-Confetti;Brca1fl/fl;Trp53fl/fl mice were intraductally injected with recombinant TAT-Cre protein (20âunits per gland diluted in 20âµl PBS, Sigma-Aldrich) between 10 and 15âweeks of age. Then, 64 or 225âdays post-induction, mammary glands were harvested and prepared for fluorescence-activated cell sorting (FACS) as described before. Both confetti-positive Brca1â/â;Trp53â/â and non-recombined control cells were seeded in a 24-well plate, 10,000 cells per drop of Cultrex Basement Membrane Extract (Type 2, 3532-010-02, R&D Systems) and cultured in the DMEM/F12 (Gibco) supplemented with iInsulin-transferrin-selenium (100Ã, catalogue no. 41400045, Gibco), B-27 Supplement (50Ã, catalogue no. 17504044, Gibco), NAC 1.25âmM (N-acetyl-l-cysteine, 0.125âM in PBS, catalogue no. 6169116, Biogems), mFGF2 2.5ânM (Fibroblast Growth Factor 2, catalogue no. 100-18B, PeproTech) and mEGF 2ânM (Epidermal Growth Factor, catalogue no. 3165-09, PeproTech). After 2âweeks of culture, organoids were fixed with 4% PFA (catalogue no. 47347, AlfaAesar) for 10â15âmin at room temperature inside the Basement Membrane Extract droplet on an orbital shaker at 25ârpm. Afterwards, organoids were washed three times for 10âmin with PBS, followed by incubation in permeabilization buffer (5% normal goat serum (catalogue no. 16210072, Gibco) and 0.5% Triton X-100 ((Sigma-Aldrich) in PBS) for 3âh. To stain for cell proliferation and cell death, primary antibodies Ki-67 (rat, clone SolA15, 14-5698-82, eBioscience, 1:100) and Cleaved Caspase-3 (rabbit, Asp175, no. 9661, Cell Signaling Technology, 1:400), respectively, were added in the blocking buffer (5% normal goat serum (Gibco) in PBS), and incubated overnight at 4â°C. Organoids were washed three times for 15âmin with PBS and secondary antibodies goat antirabbit IgG Antibody, Alexa Fluor 647 (catalogue no. A21244, Thermo Scientific, 1:400), goat antirat IgG Antibody, Alexa Fluor 647 (catalogue no. A21247, Thermo Scientific, 1:400) were added and incubated for more than 5âh at room temperature covered in aluminium foil on an orbital shaker at 25ârpm. Organoids were washed three times for 15âmin with PBS and stained organoids were mounted by adding 200âµl of Vectashield mounting medium (VECTASHIELD HardSet Antifade Mounting Medium, H-1400, Vector Laboratories). Organoid imaging was performed on an inverted Leica SP8 Dive system (Leica Microsystems), in which Alexa-647 secondary antibodies were excited at 635ânm and detected between 660 and 700ânm, and organoids were imaged using brightfield. The Ki-67/CC3 ratio was derived by first calculating the organoid area and Ki-67+ or CC3+ areas using ImageJ software (https://imagej.nih.gov/ij/), then calculating the percentage of Ki-67 or CC3 expressing cells per organoid, followed by calculation of the Ki-67/CC3 ratio for every organoid.
Statistics
P values and statistical tests performed are included in the figure legends or Supplementary Information 1. The longitudinal data of the clone fractions (Figs. 2d and 5g) was analysed using a regression model with a time effect, for which the interaction between time and group was tested. For full details, see Supplementary Information 2. Details on statistics concerning the mathematical modelling can be found in the Supplementary Information 4.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.